

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144723.3 + phase: 0
(127 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
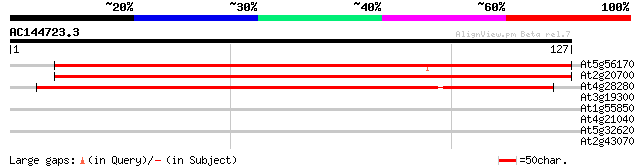
Score E
Sequences producing significant alignments: (bits) Value
At5g56170 predicted GPI-anchored protein 169 3e-43
At2g20700 predicted GPI-anchored protein 145 5e-36
At4g28280 putative GPI-anchored protein 142 6e-35
At3g19300 unknown protein 35 0.013
At1g55850 putative cellulose synthase catalytic subunit (At1g55850) 26 4.7
At4g21040 putative protein 26 6.2
At5g32620 putative protein 25 8.1
At2g43070 unknown protein 25 8.1
>At5g56170 predicted GPI-anchored protein
Length = 168
Score = 169 bits (428), Expect = 3e-43
Identities = 77/120 (64%), Positives = 93/120 (77%), Gaps = 3/120 (2%)
Query: 11 CSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINLY 70
C VNFEF+NYTIITSKCKGPKYPPKECCG+FK+FACPY D +NDL++DCA+TMFSYINLY
Sbjct: 49 CPVNFEFMNYTIITSKCKGPKYPPKECCGAFKDFACPYTDQLNDLSSDCATTMFSYINLY 108
Query: 71 GRYPPGLFASECREGKEGLACDA---LPPSVSADDTANQIVHTPSLVLVLTACIFLILLF 127
G+YPPGLFA++C+EGKEGL C A LPP SA+ A + + V A + + LF
Sbjct: 109 GKYPPGLFANQCKEGKEGLECPAGSQLPPETSAEVNAATTSSSRLWLTVSAALLVFVKLF 168
>At2g20700 predicted GPI-anchored protein
Length = 181
Score = 145 bits (366), Expect = 5e-36
Identities = 67/117 (57%), Positives = 83/117 (70%)
Query: 11 CSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSYINLY 70
C +F NYTIITS+CKGP YP CC +FK+FACP+A+V+ND NDCASTMFSYINLY
Sbjct: 62 CKEDFANKNYTIITSRCKGPNYPANVCCSAFKDFACPFAEVLNDEKNDCASTMFSYINLY 121
Query: 71 GRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILLF 127
GRYPPG+FA+ C+EGKEGL C + S SA + T + + VL+ + L LLF
Sbjct: 122 GRYPPGIFANMCKEGKEGLDCTDVTQSASATSDSIPRASTTASLAVLSTFLVLCLLF 178
>At4g28280 putative GPI-anchored protein
Length = 160
Score = 142 bits (357), Expect = 6e-35
Identities = 66/117 (56%), Positives = 84/117 (71%), Gaps = 1/117 (0%)
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A A C +F NYTIITSKCKGP YP K CC +FK+FACP+A+V+ND DCASTMFSY
Sbjct: 44 AKATCKEDFAAKNYTIITSKCKGPNYPAKVCCSAFKDFACPFAEVLNDEKTDCASTMFSY 103
Query: 67 INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFL 123
INLYGRYPPG+FA+ C+EGKEGL C + P+ S+ + +V T L++ ++ L
Sbjct: 104 INLYGRYPPGIFANMCKEGKEGLDCTDVTPT-SSSHASIPLVSTHVLLITVSILFHL 159
>At3g19300 unknown protein
Length = 663
Score = 34.7 bits (78), Expect = 0.013
Identities = 23/76 (30%), Positives = 34/76 (44%), Gaps = 11/76 (14%)
Query: 6 IAPAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEF----ACPYAD------VINDL 55
+ AGC ++F N+T++ S C K CC F YA+ V +DL
Sbjct: 25 LTEAGCPLDFTSSNFTLVASVCSNNTERAK-CCRYMNAFVAISVARYANYTADLGVTSDL 83
Query: 56 TNDCASTMFSYINLYG 71
T C +T+ + LYG
Sbjct: 84 TEICITTISRTMELYG 99
>At1g55850 putative cellulose synthase catalytic subunit (At1g55850)
Length = 729
Score = 26.2 bits (56), Expect = 4.7
Identities = 10/26 (38%), Positives = 14/26 (53%)
Query: 43 EFACPYADVINDLTNDCASTMFSYIN 68
++ CP DVI LT C +Y+N
Sbjct: 436 KYGCPVEDVITGLTIQCRGWKSAYLN 461
>At4g21040 putative protein
Length = 232
Score = 25.8 bits (55), Expect = 6.2
Identities = 10/26 (38%), Positives = 12/26 (45%)
Query: 58 DCASTMFSYINLYGRYPPGLFASECR 83
D +T F Y N Y + P F CR
Sbjct: 31 DSDNTKFCYYNNYSEFQPRYFCKNCR 56
>At5g32620 putative protein
Length = 301
Score = 25.4 bits (54), Expect = 8.1
Identities = 10/40 (25%), Positives = 20/40 (50%)
Query: 74 PPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSL 113
PPG+ A++ + K + + P+ DD+ + H +L
Sbjct: 215 PPGVKAAKAKGRKSAIVKEGKKPATVKDDSGQSVEHFQNL 254
>At2g43070 unknown protein
Length = 543
Score = 25.4 bits (54), Expect = 8.1
Identities = 16/50 (32%), Positives = 24/50 (48%), Gaps = 3/50 (6%)
Query: 81 ECREGKEGLACDALPPSVSADDTANQIVH---TPSLVLVLTACIFLILLF 127
+ E LA D DD +I+ T ++ ++TA IFL+LLF
Sbjct: 222 QANESYSILAKDVSSAGTRKDDPEKEILDISVTGAVFFIVTASIFLLLLF 271
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.325 0.141 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,099,506
Number of Sequences: 26719
Number of extensions: 121944
Number of successful extensions: 326
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 317
Number of HSP's gapped (non-prelim): 8
length of query: 127
length of database: 11,318,596
effective HSP length: 87
effective length of query: 40
effective length of database: 8,994,043
effective search space: 359761720
effective search space used: 359761720
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144723.3


