

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144483.8 + phase: 0
(253 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
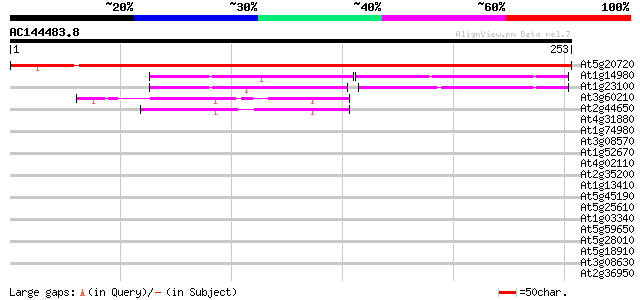
Score E
Sequences producing significant alignments: (bits) Value
At5g20720 chloroplast Cpn21 protein 363 e-101
At1g14980 mitochondrial chaperonin 10 59 2e-09
At1g23100 10kd chaperonin like protein 56 2e-08
At3g60210 unknown protein 54 6e-08
At2g44650 unknown protein 50 2e-06
At4g31880 unknown protein 31 0.73
At1g74980 putative ribosomal protein S9 31 0.73
At3g08570 hypothetical protein 30 1.6
At1g52670 unknown protein 30 1.6
At4g02110 predicted protein of unknown function 29 2.1
At2g35200 unknown protein 29 2.1
At1g13410 hypothetical protein 29 2.1
At5g45190 unknown protein 29 2.8
At5g25610 dehydration-induced protein RD22 29 2.8
At1g03340 unknown protein 29 2.8
At5g59650 serine/threonine-specific protein kinase - like 28 6.2
At5g28010 major latex protein homolog - like 28 6.2
At5g18910 protein kinase - like protein 28 6.2
At3g08630 unknown protein 28 6.2
At2g36950 putative farnesylated protein 28 6.2
>At5g20720 chloroplast Cpn21 protein
Length = 253
Score = 363 bits (931), Expect = e-101
Identities = 187/255 (73%), Positives = 221/255 (86%), Gaps = 4/255 (1%)
Query: 1 MATTQLTASSI--STRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPK 58
MA TQLTAS + S R+L+S + LR SS++F ++ + L Q +VVKAA+VVAPK
Sbjct: 1 MAATQLTASPVTMSARSLASLDGLRASSVKF--SSLKPGTLRQSQFRRLVVKAASVVAPK 58
Query: 59 HTTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVK 118
+T++KPLGDRVLVKIK+AEEKT GGILLPSTAQSKPQGGEVVA+G+G+T+GKNK+DI+V
Sbjct: 59 YTSIKPLGDRVLVKIKEAEEKTLGGILLPSTAQSKPQGGEVVAVGEGRTIGKNKIDITVP 118
Query: 119 AGAQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKT 178
GAQ++YSKYAGTEVEF+ KHLILK++DIVGILETE++KDLKPLNDRV IKVAEAEEKT
Sbjct: 119 TGAQIIYSKYAGTEVEFNDVKHLILKEDDIVGILETEDIKDLKPLNDRVFIKVAEAEEKT 178
Query: 179 AGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDGS 238
AGGLLLTE TK+KPSIGTVIAVGPG +D+EG PL + G+TVLYSKYAGNDFKGKDGS
Sbjct: 179 AGGLLLTETTKEKPSIGTVIAVGPGSLDEEGKITPLPVSTGSTVLYSKYAGNDFKGKDGS 238
Query: 239 DYIALRASDVMAILS 253
+YIALRASDVMAILS
Sbjct: 239 NYIALRASDVMAILS 253
>At1g14980 mitochondrial chaperonin 10
Length = 98
Score = 58.9 bits (141), Expect = 2e-09
Identities = 33/93 (35%), Positives = 55/93 (58%), Gaps = 2/93 (2%)
Query: 64 PLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNK-VDISVKAGAQ 122
P +R+LV+ KT+ GILLP + SK G+V+A+G G K + +SVK G
Sbjct: 7 PTFNRILVQRVIQPAKTESGILLPEKS-SKLNSGKVIAVGPGSRDKDGKLIPVSVKEGDT 65
Query: 123 VVYSKYAGTEVEFDGSKHLILKDEDIVGILETE 155
V+ +Y GT+V+ +++ + +DED++G L +
Sbjct: 66 VLLPEYGGTQVKLGENEYHLFRDEDVLGTLHED 98
Score = 56.2 bits (134), Expect = 2e-08
Identities = 36/96 (37%), Positives = 53/96 (54%), Gaps = 2/96 (2%)
Query: 157 VKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSI 216
+K L P +R+L++ KT G+LL E + K + G VIAVGPG D +G P+S+
Sbjct: 2 MKRLIPTFNRILVQRVIQPAKTESGILLPEKSS-KLNSGKVIAVGPGSRDKDGKLIPVSV 60
Query: 217 LPGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
G+TVL +Y G K + ++Y R DV+ L
Sbjct: 61 KEGDTVLLPEYGGTQVKLGE-NEYHLFRDEDVLGTL 95
>At1g23100 10kd chaperonin like protein
Length = 97
Score = 55.8 bits (133), Expect = 2e-08
Identities = 36/95 (37%), Positives = 51/95 (52%), Gaps = 2/95 (2%)
Query: 158 KDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSIL 217
K L P +RVL++ KT G+LL E + S G VIAVGPG D GN P+S+
Sbjct: 3 KRLIPTLNRVLVEKILPPSKTVSGILLPEKSSQLNS-GRVIAVGPGARDRAGNLIPVSVK 61
Query: 218 PGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
G+ VL ++ G K + +++ R D+MA L
Sbjct: 62 EGDNVLLPEFGGTQVKLGE-KEFLLYRDEDIMATL 95
Score = 53.1 bits (126), Expect = 1e-07
Identities = 33/90 (36%), Positives = 51/90 (56%), Gaps = 2/90 (2%)
Query: 64 PLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDG-KTVGKNKVDISVKAGAQ 122
P +RVLV+ KT GILLP + S+ G V+A+G G + N + +SVK G
Sbjct: 7 PTLNRVLVEKILPPSKTVSGILLPEKS-SQLNSGRVIAVGPGARDRAGNLIPVSVKEGDN 65
Query: 123 VVYSKYAGTEVEFDGSKHLILKDEDIVGIL 152
V+ ++ GT+V+ + L+ +DEDI+ L
Sbjct: 66 VLLPEFGGTQVKLGEKEFLLYRDEDIMATL 95
>At3g60210 unknown protein
Length = 138
Score = 54.3 bits (129), Expect = 6e-08
Identities = 43/128 (33%), Positives = 70/128 (54%), Gaps = 28/128 (21%)
Query: 31 RNNARI-AILTQWSLPSMVVKAATVVAPKHTTVKPLGDRVLVKIKDAEEKTQGGILLPST 89
RN+ RI A+ T+W P+ VV P DRVLV+++ EK+ GG+LLP +
Sbjct: 34 RNSFRINAVSTKWE-PAKVV--------------PQADRVLVRLEVLPEKSSGGVLLPKS 78
Query: 90 AQ--SKPQGGEVVAIGDGKTVGKNKVDISVKAGAQVVYSKYAGTEVEF--DGSKHLILKD 145
A + GEVV++ G VG+ V+ G +V++S + EV+F + +KH K+
Sbjct: 79 AVKFERYLTGEVVSV--GSEVGE------VEPGKKVLFSDMSAYEVDFGTEDAKHCFCKE 130
Query: 146 EDIVGILE 153
D++ I++
Sbjct: 131 SDLLAIVQ 138
Score = 34.7 bits (78), Expect = 0.050
Identities = 26/94 (27%), Positives = 49/94 (51%), Gaps = 12/94 (12%)
Query: 162 PLNDRVLIKVAEAEEKTAGGLLLTEATK--DKPSIGTVIAVGPGPVDDEGNRKPLSILPG 219
P DRVL+++ EK++GG+LL ++ ++ G V++VG + E PG
Sbjct: 53 PQADRVLVRLEVLPEKSSGGVLLPKSAVKFERYLTGEVVSVGSEVGEVE---------PG 103
Query: 220 NTVLYSKYAGNDFK-GKDGSDYIALRASDVMAIL 252
VL+S + + G + + + + SD++AI+
Sbjct: 104 KKVLFSDMSAYEVDFGTEDAKHCFCKESDLLAIV 137
>At2g44650 unknown protein
Length = 139
Score = 49.7 bits (117), Expect = 2e-06
Identities = 30/97 (30%), Positives = 54/97 (54%), Gaps = 10/97 (10%)
Query: 60 TTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQ--SKPQGGEVVAIGDGKTVGKNKVDISV 117
T V P DRVLV+++D K+ GG+LLP A + GE++++G ++V V
Sbjct: 50 TKVVPQADRVLVRLEDLPIKSSGGVLLPKAAVKFERYLTGEIISVG-------SEVGQQV 102
Query: 118 KAGAQVVYSKYAGTEVEF-DGSKHLILKDEDIVGILE 153
G +V++S + EV+ ++H K+ D++ ++E
Sbjct: 103 GPGKRVLFSDVSAYEVDLGTDARHCFCKESDLLALVE 139
Score = 32.7 bits (73), Expect = 0.19
Identities = 24/93 (25%), Positives = 46/93 (48%), Gaps = 10/93 (10%)
Query: 162 PLNDRVLIKVAEAEEKTAGGLLLTEATK--DKPSIGTVIAVGPGPVDDEGNRKPLSILPG 219
P DRVL+++ + K++GG+LL +A ++ G +I+VG G PG
Sbjct: 54 PQADRVLVRLEDLPIKSSGGVLLPKAAVKFERYLTGEIISVGSEVGQQVG--------PG 105
Query: 220 NTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
VL+S + + + + + SD++A++
Sbjct: 106 KRVLFSDVSAYEVDLGTDARHCFCKESDLLALV 138
>At4g31880 unknown protein
Length = 873
Score = 30.8 bits (68), Expect = 0.73
Identities = 22/80 (27%), Positives = 40/80 (49%), Gaps = 3/80 (3%)
Query: 101 AIGDGKTVGKNKVDISVKAGAQVVYSKYAGTEVEFDGS--KHLILKDEDIVGILETEEVK 158
++G GK G++ V +K + + Y G +D + KHL++ D+ IL + K
Sbjct: 597 SLGQGKASGESLVGSRIKVWWPMDQAYYKGVVESYDAAKKKHLVIYDDGDQEILYLKNQK 656
Query: 159 DLKPLNDRVLIKVAEAEEKT 178
PL++ L + EA ++T
Sbjct: 657 -WSPLDESELSQDEEAADQT 675
>At1g74980 putative ribosomal protein S9
Length = 132
Score = 30.8 bits (68), Expect = 0.73
Identities = 20/57 (35%), Positives = 32/57 (56%)
Query: 1 MATTQLTASSISTRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAP 57
MA+ ASS+S+ + SS R ++I FPR N+ A+ + + + + ATV AP
Sbjct: 1 MASITNLASSLSSLSFSSQVSQRPNTISFPRANSVFALPAKSARRASLSITATVSAP 57
>At3g08570 hypothetical protein
Length = 612
Score = 29.6 bits (65), Expect = 1.6
Identities = 21/61 (34%), Positives = 30/61 (48%), Gaps = 6/61 (9%)
Query: 165 DRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDE---GNRKPLSILPGNT 221
+++L+K E+ GG LLT +K IG + G GP + + NRK S L T
Sbjct: 547 EQILMKQGMMEKSGHGGTLLTSLSK---GIGRISIFGGGPTEGKLRNANRKSKSRLERKT 603
Query: 222 V 222
V
Sbjct: 604 V 604
>At1g52670 unknown protein
Length = 274
Score = 29.6 bits (65), Expect = 1.6
Identities = 24/109 (22%), Positives = 46/109 (42%), Gaps = 4/109 (3%)
Query: 17 SSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPKHTTVKPLGDRVLVKIKDA 76
++F RLR ++ P N + S+ +V A + +TV P + +KD+
Sbjct: 16 TNFSRLRCGNLLIPNNQRLFVDQSPMKYLSLRTTLRSVKAIQLSTVPPAETEAIADVKDS 75
Query: 77 EEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAGAQVVY 125
+E T+ ++ P+ EV A+ T + + +K G +Y
Sbjct: 76 DE-TKSTVV---NTHLMPKSSEVEALISEITDSSSIAEFELKLGGFRLY 120
>At4g02110 predicted protein of unknown function
Length = 1293
Score = 29.3 bits (64), Expect = 2.1
Identities = 33/144 (22%), Positives = 52/144 (35%), Gaps = 11/144 (7%)
Query: 109 GKNKVDISVKAGAQVVYSKYAGTEVEFDGSKHLILKDEDIVGI-------LETEE----V 157
GK ++ + + + S TEV D +KH+ + E + GI L EE V
Sbjct: 618 GKVSAPVTGNSNQKEISSPVLNTEVVQDMAKHIDTETEALQGIDSVDNKSLAPEEKDHLV 677
Query: 158 KDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSIL 217
DL D++ K EA + +L D P+ VD +++
Sbjct: 678 LDLMVNQDKLQAKTPEAADAEVEITVLERELNDVPTEDPSDGALQSEVDKNTSKRKREAG 737
Query: 218 PGNTVLYSKYAGNDFKGKDGSDYI 241
G L G+ F K G +
Sbjct: 738 VGKNSLQRGKKGSSFTAKVGKSRV 761
>At2g35200 unknown protein
Length = 201
Score = 29.3 bits (64), Expect = 2.1
Identities = 19/60 (31%), Positives = 27/60 (44%), Gaps = 2/60 (3%)
Query: 151 ILETEEVKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGN 210
ILETE KDLKP R L ++ + + + K S+ + P PV +GN
Sbjct: 114 ILETE--KDLKPGRGRWLWRLFRGNREEETKIKAAKVMKKSNSVAGEVLFSPTPVTSKGN 171
>At1g13410 hypothetical protein
Length = 511
Score = 29.3 bits (64), Expect = 2.1
Identities = 14/41 (34%), Positives = 23/41 (55%)
Query: 105 GKTVGKNKVDISVKAGAQVVYSKYAGTEVEFDGSKHLILKD 145
G+T+G NKVD++VK +Y K E+ + K + +D
Sbjct: 251 GETLGYNKVDVNVKNALIDMYGKCGAIEIAMEVFKGIKRRD 291
>At5g45190 unknown protein
Length = 579
Score = 28.9 bits (63), Expect = 2.8
Identities = 22/66 (33%), Positives = 36/66 (54%), Gaps = 7/66 (10%)
Query: 126 SKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTAGGLLLT 185
S+ G E++ DG+ H ++ D+ L ++ KDLK L D+V K +A++ LL
Sbjct: 394 SEREGGELQDDGAVHKS-RNVDVGDALISQSPKDLKLLRDKVKAKREKAKK------LLG 446
Query: 186 EATKDK 191
E T+ K
Sbjct: 447 ERTRKK 452
>At5g25610 dehydration-induced protein RD22
Length = 392
Score = 28.9 bits (63), Expect = 2.8
Identities = 17/63 (26%), Positives = 30/63 (46%), Gaps = 11/63 (17%)
Query: 81 QGGILLPSTAQSKPQGGEVVAIGDGKTVG----------KNKVDISVKAGAQVVYSKYAG 130
+GG+ + T + KP GG V++G GK G + D+ V G V++++ G
Sbjct: 84 KGGVRV-DTGKGKPGGGTHVSVGSGKGHGGGVAVHTGKPGKRTDVGVGKGGVTVHTRHKG 142
Query: 131 TEV 133
+
Sbjct: 143 RPI 145
>At1g03340 unknown protein
Length = 385
Score = 28.9 bits (63), Expect = 2.8
Identities = 21/64 (32%), Positives = 31/64 (47%), Gaps = 1/64 (1%)
Query: 14 RNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPKHTTVKPLGDRVLVKI 73
RN + + +L QF R ++ + I T LP V A +PK T +KP + KI
Sbjct: 182 RNSTRWRKLSRLVPQFQRFDSEVPIDTL-QLPEDAVTLAPPKSPKKTRLKPSPKKQNPKI 240
Query: 74 KDAE 77
+D E
Sbjct: 241 RDKE 244
>At5g59650 serine/threonine-specific protein kinase - like
Length = 892
Score = 27.7 bits (60), Expect = 6.2
Identities = 16/41 (39%), Positives = 21/41 (51%)
Query: 27 IQFPRNNARIAILTQWSLPSMVVKAATVVAPKHTTVKPLGD 67
IQFPRN + ILT S S+V A K + ++ GD
Sbjct: 355 IQFPRNQLHLLILTSLSSTSVVAVKNIEAAYKLSRIRWQGD 395
>At5g28010 major latex protein homolog - like
Length = 166
Score = 27.7 bits (60), Expect = 6.2
Identities = 12/33 (36%), Positives = 19/33 (57%)
Query: 98 EVVAIGDGKTVGKNKVDISVKAGAQVVYSKYAG 130
EV + +GK +V++ +KA A + Y YAG
Sbjct: 6 EVTTMPKSSLLGKLEVEVEIKAPAAIFYHIYAG 38
>At5g18910 protein kinase - like protein
Length = 511
Score = 27.7 bits (60), Expect = 6.2
Identities = 21/88 (23%), Positives = 39/88 (43%), Gaps = 4/88 (4%)
Query: 58 KHTTVKPLGDRVLVKIKDAEEKTQ----GGILLPSTAQSKPQGGEVVAIGDGKTVGKNKV 113
K +K L D +L D EE + + + T+ ++PQ +VV I G +K+
Sbjct: 408 KENKIKQLVDPILEDDYDVEELDRLVFIASLCIHQTSMNRPQMSQVVEILRGDKCSLDKL 467
Query: 114 DISVKAGAQVVYSKYAGTEVEFDGSKHL 141
+ Q YS+ E++ +++L
Sbjct: 468 RERENSKLQRTYSEELLDNEEYNSTRYL 495
>At3g08630 unknown protein
Length = 339
Score = 27.7 bits (60), Expect = 6.2
Identities = 14/50 (28%), Positives = 28/50 (56%)
Query: 3 TTQLTASSISTRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAA 52
+T+ S++ L + R T+ + FPRN + ++ T +S P+++V A
Sbjct: 11 STKSDQSNVRLPRLINLSRDPTARVLFPRNGSVSSLHTNFSSPNIMVPCA 60
>At2g36950 putative farnesylated protein
Length = 386
Score = 27.7 bits (60), Expect = 6.2
Identities = 39/170 (22%), Positives = 69/170 (39%), Gaps = 31/170 (18%)
Query: 71 VKIKDA-EEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNK----------------V 113
VK+++ EEKT+ ++L + K +G A+G+ K G +K V
Sbjct: 98 VKLQEKLEEKTKRKVVL-ANPPPKVEGPVAAAVGEKKADGGDKEAAPPAPAPAAPKESVV 156
Query: 114 DISVKAGAQVVYSKYAGTEVEFDGSKHLILKD-EDIVGILETEEVKDLKPLNDRVLIKVA 172
+ ++ + K ++ G + + + +D+V + T +VK+L PL + L +
Sbjct: 157 PLKIRLHCEGCIQKIKKIILKIKGVETVAIDGAKDVVTVKGTIDVKELVPLLTKKLKRTV 216
Query: 173 E----------AEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRK 212
E A E +A K+ PS G A G D G +K
Sbjct: 217 EPLVPAKKDDGAAENKKTEAAAPDAKKEAPSAGVNEAKKEG--SDGGEKK 264
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.311 0.131 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,525,638
Number of Sequences: 26719
Number of extensions: 247216
Number of successful extensions: 483
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 12
Number of HSP's successfully gapped in prelim test: 12
Number of HSP's that attempted gapping in prelim test: 460
Number of HSP's gapped (non-prelim): 29
length of query: 253
length of database: 11,318,596
effective HSP length: 97
effective length of query: 156
effective length of database: 8,726,853
effective search space: 1361389068
effective search space used: 1361389068
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144483.8


