

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144405.9 - phase: 0
(166 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
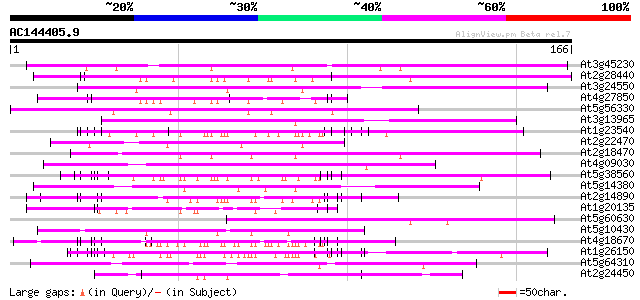
Score E
Sequences producing significant alignments: (bits) Value
At3g45230 serine/proline-rich protein 115 1e-26
At2g28440 En/Spm-like transposon protein 99 1e-21
At3g24550 protein kinase, putative 83 8e-17
At4g27850 putative proline-rich protein 74 4e-14
At5g56330 putative protein 70 4e-13
At3g13965 unknown protein 70 5e-13
At1g23540 putative serine/threonine protein kinase 70 7e-13
At2g22470 arabinogalactan-protein AGP2 69 2e-12
At2g18470 putative protein kinase 69 2e-12
At4g09030 arabinogalactan-protein AGP10 68 3e-12
At5g38560 putative protein 67 6e-12
At5g14380 agp6 67 6e-12
At2g14890 arabinogalactan-protein AGP9 66 1e-11
At1g20135 anter-specific proline-rich protein APG precursor 65 1e-11
At5g60630 unknown protein 64 5e-11
At5g10430 AtAGP4 63 6e-11
At4g18670 extensin-like protein 63 8e-11
At1g26150 Pto kinase interactor, putative 63 8e-11
At5g64310 arabinogalactan-protein AGP1 (gb|AAC77823.1) 62 1e-10
At2g24450 predicted GPI-anchored protein 62 1e-10
>At3g45230 serine/proline-rich protein
Length = 175
Score = 115 bits (287), Expect = 1e-26
Identities = 80/175 (45%), Positives = 105/175 (59%), Gaps = 19/175 (10%)
Query: 6 FSLVFLLLSFLVNIAS--SADSPAPTP-ATNSSLNSPSPTPIPTPSPANSPPAPTP---T 59
F +V ++LS ++ + SPAP+P +S L SP P+ +++ PA +P +
Sbjct: 5 FIIVAMMLSLVLVSGEILTKSSPAPSPDLADSPLIHASP---PSKLGSHNSPAESPIEYS 61
Query: 60 PTPSPHSDSPPAPSPDNSPSSSP-------SPSPSSSPAPSPDEAADNNAISHTGI-GED 111
P P ++ P+PSP NSPS SP SPS S+SP+PSP EA+D N TGI GE
Sbjct: 62 SPPEPETEHSPSPSPANSPSVSPPLPNDSQSPSSSASPSPSP-EASDVNHSDITGIEGEK 120
Query: 112 GKS-SGGGMSSGKKAGIAVGVIAAVGVVALGAMVVKKRRQNIQRSEYGYTARREL 165
S SGGGMS GKK G+A G IAAV VV + V KKR++NI+RS YGY AR L
Sbjct: 121 LPSGSGGGMSGGKKVGVAFGAIAAVCVVGVAGFVYKKRQENIRRSRYGYAAREIL 175
>At2g28440 En/Spm-like transposon protein
Length = 268
Score = 99.0 bits (245), Expect = 1e-21
Identities = 67/157 (42%), Positives = 85/157 (53%), Gaps = 14/157 (8%)
Query: 23 ADSPAPTPATNSSLNSP-SPTPIPTP-SPANSPPAPTPTPTP-SPHSDSPPAPSPDNSPS 79
ADSP P P+++ NSP SP P P S A+SP P P P P SP S S P P+P +PS
Sbjct: 113 ADSPLP-PSSSPEANSPQSPASSPKPESLADSPSPPPPPPQPESPSSPSYPEPAPVPAPS 171
Query: 80 SSPS---PSPSS----SPAPSPDEAADNNAISHTGIGEDGKSSGG---GMSSGKKAGIAV 129
S P P + SPAPSP+ + + GE+ G GMS +KAGIA+
Sbjct: 172 DDDSDDDPEPETEYFPSPAPSPELGMAQDIKASDAAGEELNDERGEDYGMSGLEKAGIAI 231
Query: 130 GVIAAVGVVALGAMVVKKRRQNIQRSEYGYTARRELL 166
G I VG + +GA+V KKRR N+ R+ Y Y E L
Sbjct: 232 GTILGVGAIVIGALVYKKRRDNMTRARYTYFTEGEFL 268
Score = 47.0 bits (110), Expect = 5e-06
Identities = 34/95 (35%), Positives = 51/95 (52%), Gaps = 9/95 (9%)
Query: 8 LVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTP-TPTPSPHS 66
+V L + L+ + A +P+PA S PS T P SP++SP +P +P+ SP
Sbjct: 8 IVMLSICLLIFDFAGAQEESPSPAAVSPGREPS-TDSPL-SPSSSPEEDSPLSPSSSPEE 65
Query: 67 DSP--PAPSPDN----SPSSSPSPSPSSSPAPSPD 95
DSP P+ SP+ +PSSSP +P+ SP+
Sbjct: 66 DSPLPPSSSPEEDSPLAPSSSPEVDSPLAPSSSPE 100
Score = 45.4 bits (106), Expect = 1e-05
Identities = 32/82 (39%), Positives = 45/82 (54%), Gaps = 8/82 (9%)
Query: 22 SADSPAPTPATNSSLN-SPSPTPIPTPSPANSPPAPTPTP-TPSPHSDSP--PAPSPD-- 75
+A SP P+T+S L+ S SP SP++SP +P P + SP DSP P+ SP+
Sbjct: 31 AAVSPGREPSTDSPLSPSSSPEEDSPLSPSSSPEEDSPLPPSSSPEEDSPLAPSSSPEVD 90
Query: 76 --NSPSSSPSPSPSSSPAPSPD 95
+PSSSP P+ SP+
Sbjct: 91 SPLAPSSSPEVDSPQPPSSSPE 112
Score = 38.1 bits (87), Expect = 0.002
Identities = 24/60 (40%), Positives = 30/60 (50%), Gaps = 10/60 (16%)
Query: 38 SPSPTPIPTPSPANSPPAPTP-TPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDE 96
SPSP + SP P +P +P+ SP DSP SPSSSP P+ SP+E
Sbjct: 27 SPSPAAV---SPGREPSTDSPLSPSSSPEEDSPL------SPSSSPEEDSPLPPSSSPEE 77
>At3g24550 protein kinase, putative
Length = 652
Score = 82.8 bits (203), Expect = 8e-17
Identities = 50/145 (34%), Positives = 78/145 (53%), Gaps = 12/145 (8%)
Query: 21 SSADSPAPTPATNSSL---NSPSPTPIPTPSPANSPPAPTPTPTPS-PHSDSPPAPSPDN 76
++ SP P+P+TNS+ +SP P +P PSP S P P P+PS P + SPP+P+ +
Sbjct: 38 TTPSSPPPSPSTNSTSPPPSSPLPPSLPPPSPPGSLTPPLPQPSPSAPITPSPPSPTTPS 97
Query: 77 SPSSSPSPS--PSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAA 134
+P S PSP+ P ++P+ S N S S G+S+G GIA+G +A
Sbjct: 98 NPRSPPSPNQGPPNTPSGSTPRTPSNTKPS------PPSDSSDGLSTGVVVGIAIGGVAI 151
Query: 135 VGVVALGAMVVKKRRQNIQRSEYGY 159
+ ++ L ++ KK+R+ E Y
Sbjct: 152 LVILTLICLLCKKKRRRRHDDEAAY 176
Score = 34.7 bits (78), Expect = 0.024
Identities = 33/110 (30%), Positives = 45/110 (40%), Gaps = 15/110 (13%)
Query: 27 APTPATNSSLNS-PSPTPIPTPSPANSPPAPTPTPT------PSPHSDSP--PAPSPDNS 77
A P+ N + S P P P PSP PP P P P S +SD P P PSP
Sbjct: 204 ASRPSDNHVVTSLPPPKP---PSPPRKPPPPPPPPAFMSSSGGSDYSDLPVLPPPSPGLV 260
Query: 78 PSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGI 127
S S + + + ++ N + G G K G + SGK+ +
Sbjct: 261 LGFSKSTFTYEELSRATNGFSEANLLGQGGFGYVHK---GILPSGKEVAV 307
>At4g27850 putative proline-rich protein
Length = 577
Score = 73.9 bits (180), Expect = 4e-14
Identities = 48/99 (48%), Positives = 57/99 (57%), Gaps = 14/99 (14%)
Query: 9 VFLLLSFLVNIASSADSPAPTPATNSSLN---SPSPTP-----IPTPSPANSP---PAPT 57
V LLLS +V +S P P P S L SPSPTP +P+P P +SP P P
Sbjct: 147 VLLLLSVIVLWSSDPPLPPPPPPYPSPLPPPPSPSPTPGPDSPLPSPGP-DSPLPLPGPP 205
Query: 58 PTPTPSPHSDSP-PAPSPDNSPSSSPSPSPSSSPAPSPD 95
P+P+P+P DSP P+P PD SP P P PSSSP P PD
Sbjct: 206 PSPSPTPGPDSPLPSPGPD-SPLPLPGPPPSSSPTPGPD 243
Score = 72.0 bits (175), Expect = 1e-13
Identities = 41/88 (46%), Positives = 51/88 (57%), Gaps = 15/88 (17%)
Query: 25 SPAPTPATNSSLNSPSP-TPIPTPSPANSPPAPTPTP-----TPSPHSDSP---PAPSPD 75
SP+PTP +S L SP P +P+P P P PP+P+PTP PSP DSP P P P
Sbjct: 179 SPSPTPGPDSPLPSPGPDSPLPLPGP---PPSPSPTPGPDSPLPSPGPDSPLPLPGPPPS 235
Query: 76 NSPS---SSPSPSPSSSPAPSPDEAADN 100
+SP+ SP PSP P+PSP D+
Sbjct: 236 SSPTPGPDSPLPSPGPPPSPSPTPGPDS 263
Score = 68.9 bits (167), Expect = 1e-12
Identities = 40/84 (47%), Positives = 49/84 (57%), Gaps = 18/84 (21%)
Query: 25 SPAPTPATNSSLNSP---SPTPIPTPSPANSP----------PAPTPTPTPSPHSDSP-P 70
SP+PTP +S L SP SP P+P P P++SP P P P+P+P+P DSP P
Sbjct: 207 SPSPTPGPDSPLPSPGPDSPLPLPGPPPSSSPTPGPDSPLPSPGPPPSPSPTPGPDSPLP 266
Query: 71 APSPDNSPSSSPSPSPSSSPAPSP 94
+P PD SP SP P P P PSP
Sbjct: 267 SPGPD-SPLPSPGPDP---PLPSP 286
Score = 65.5 bits (158), Expect = 1e-11
Identities = 36/88 (40%), Positives = 44/88 (49%), Gaps = 19/88 (21%)
Query: 24 DSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTP---------------TPTPSPHSDS 68
DSP P+P +S L P P P P+P+P P P+P +PTP P S
Sbjct: 187 DSPLPSPGPDSPLPLPGPPPSPSPTPGPDSPLPSPGPDSPLPLPGPPPSSSPTPGPDS-- 244
Query: 69 PPAPSPDNSPSSSPSPSPSSS-PAPSPD 95
P PSP PS SP+P P S P+P PD
Sbjct: 245 -PLPSPGPPPSPSPTPGPDSPLPSPGPD 271
Score = 41.2 bits (95), Expect = 3e-04
Identities = 20/55 (36%), Positives = 26/55 (46%), Gaps = 14/55 (25%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTP--------------TPTPSPH 65
S +PTP +S L SP P P P+P+P P P+P P+P PH
Sbjct: 235 SSSPTPGPDSPLPSPGPPPSPSPTPGPDSPLPSPGPDSPLPSPGPDPPLPSPGPH 289
>At5g56330 putative protein
Length = 350
Score = 70.5 bits (171), Expect = 4e-13
Identities = 48/158 (30%), Positives = 66/158 (41%), Gaps = 37/158 (23%)
Query: 1 MAIPRFSLVFLLLSFLVNIASSADSPAPT---PATNSSLNSPSPTPIPT----------- 46
M I V +L+ + I SSA +P P PA + P PTP PT
Sbjct: 1 MKISSLGWVLVLIFISITIVSSAPAPKPPKPKPAPAPTPPKPKPTPAPTPPKPKPKPAPT 60
Query: 47 -----PSPANSPPAPTPTPTPSPHSDSP-PAPSPDN--------------SPSSSPSPSP 86
P+PA +PP P P P P+P P PAP+P N +P+ +P+P+P
Sbjct: 61 PPKPKPAPAPTPPKPKPAPAPTPPKPKPKPAPTPPNPKPTPAPTPPKPKPAPAPAPTPAP 120
Query: 87 SSSPAPSP---DEAADNNAISHTGIGEDGKSSGGGMSS 121
PAP P E D S+ G G + G + +
Sbjct: 121 KPKPAPKPAPGGEVEDETEFSYETKGNKGPAKWGTLDA 158
>At3g13965 unknown protein
Length = 209
Score = 70.1 bits (170), Expect = 5e-13
Identities = 43/130 (33%), Positives = 60/130 (46%), Gaps = 14/130 (10%)
Query: 28 PTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPS---- 83
P P + S++N SP P P PSP PP PTP+P P P S PP P SP S +
Sbjct: 79 PPPPSKSTINVMSPPPPPLPSPPPPPPPPTPSPPPPPISKPPPPPRAQASPPHSNNRESH 138
Query: 84 ---PSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAAVGVVAL 140
P P PSP + + +S G+++GK G+ AA+ + +
Sbjct: 139 RNQPPPRRLRPPSPPPPPPPSRVFK-------QSEKSGLNTGKTVGLVFAGFAAMLQICV 191
Query: 141 GAMVVKKRRQ 150
A +V KR+Q
Sbjct: 192 VAFLVFKRKQ 201
>At1g23540 putative serine/threonine protein kinase
Length = 731
Score = 69.7 bits (169), Expect = 7e-13
Identities = 47/140 (33%), Positives = 64/140 (45%), Gaps = 9/140 (6%)
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPT-PSPHSDSPPAPSPDNSPSS 80
S+ SP P S SP P P NSPPA PT P P S PA SP +P++
Sbjct: 136 SSPSPNVGPTNPESPPLQSPPAPPASDPTNSPPASPLDPTNPPPIQPSGPATSPPANPNA 195
Query: 81 SPSP-------SPSSSPAPSPDEAADNNAISHTGIGE-DGKSSGGGMSSGKKAGIAVGVI 132
PSP +PSS P SP + + G DG GGG G+AV
Sbjct: 196 PPSPFPTVPPKTPSSGPVVSPSLTSPSKGTPTPNQGNGDGGGGGGGYQGKTMVGMAVAGF 255
Query: 133 AAVGVVALGAMVVKKRRQNI 152
A + ++ + +V +K+++NI
Sbjct: 256 AIMALIGVVFLVRRKKKRNI 275
Score = 53.9 bits (128), Expect = 4e-08
Identities = 32/88 (36%), Positives = 40/88 (45%), Gaps = 10/88 (11%)
Query: 24 DSPAPTPATNSSLNSPSPTPIPTPSPANSPPAP------TPTPTP----SPHSDSPPAPS 73
DS P+P +SS P P P + PP P T +P P SP DS P+P
Sbjct: 36 DSSPPSPPADSSSTPPLSEPSTPPPDSQLPPLPSILPPLTDSPPPPSDSSPPVDSTPSPP 95
Query: 74 PDNSPSSSPSPSPSSSPAPSPDEAADNN 101
P S S P S +P P+E+ DNN
Sbjct: 96 PPTSNESPSPPEDSETPPAPPNESNDNN 123
Score = 48.5 bits (114), Expect = 2e-06
Identities = 39/103 (37%), Positives = 52/103 (49%), Gaps = 26/103 (25%)
Query: 22 SADSPAP---------TPATNSSL-----NSPSP------TPIPTPSPANSPPAPTPTPT 61
S+ PAP TP+ NS+L + PSP TP P P+ PP P
Sbjct: 9 SSSPPAPPADTAPPPETPSENSALPPVDSSPPSPPADSSSTP-PLSEPSTPPPDSQLPPL 67
Query: 62 PS---PHSDSPPAPSPDNSP-SSSPS-PSPSSSPAPSPDEAAD 99
PS P +DSPP PS + P S+PS P P+S+ +PSP E ++
Sbjct: 68 PSILPPLTDSPPPPSDSSPPVDSTPSPPPPTSNESPSPPEDSE 110
Score = 47.8 bits (112), Expect = 3e-06
Identities = 32/100 (32%), Positives = 47/100 (47%), Gaps = 17/100 (17%)
Query: 24 DSPAP----TPATNSSLNSPSPTPIPTPSP---ANSPPAP-------TPTPTPSPHSDSP 69
DSP P +P +S+ + P PT +PSP + +PPAP P P+ S P
Sbjct: 76 DSPPPPSDSSPPVDSTPSPPPPTSNESPSPPEDSETPPAPPNESNDNNPPPSQDLQSPPP 135
Query: 70 PAPSPD---NSPSSSPSPSPSSSPAPSPDEAADNNAISHT 106
+PSP+ +P S P SP + PA P + + + T
Sbjct: 136 SSPSPNVGPTNPESPPLQSPPAPPASDPTNSPPASPLDPT 175
Score = 47.0 bits (110), Expect = 5e-06
Identities = 27/77 (35%), Positives = 40/77 (51%), Gaps = 12/77 (15%)
Query: 28 PTPATNSSLNSPSPTPIPTPSPANSPPAPTP-----TPTPSPHSDSPPAPSPDNSPSSSP 82
P P+ L P P + P +S P+P P +P+P S++PPAP P+ S ++P
Sbjct: 66 PLPSILPPLTDSPPPPSDSSPPVDSTPSPPPPTSNESPSPPEDSETPPAP-PNESNDNNP 124
Query: 83 SPS------PSSSPAPS 93
PS P SSP+P+
Sbjct: 125 PPSQDLQSPPPSSPSPN 141
Score = 46.6 bits (109), Expect = 6e-06
Identities = 26/84 (30%), Positives = 38/84 (44%), Gaps = 9/84 (10%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAP---------TPTPTPSPHSDSPPA 71
SS+ P P+T + P P P +SPP P TP+P P ++SP
Sbjct: 46 SSSTPPLSEPSTPPPDSQLPPLPSILPPLTDSPPPPSDSSPPVDSTPSPPPPTSNESPSP 105
Query: 72 PSPDNSPSSSPSPSPSSSPAPSPD 95
P +P + P+ S ++P PS D
Sbjct: 106 PEDSETPPAPPNESNDNNPPPSQD 129
Score = 43.1 bits (100), Expect = 7e-05
Identities = 31/84 (36%), Positives = 40/84 (46%), Gaps = 6/84 (7%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P TP+ NS+L +P P+ ++S P + TP P S PP PS + SP P
Sbjct: 22 PPETPSENSALPPVDSSPPSPPADSSSTPPLSEPSTPPPDSQLPPLPSILPPLTDSPPP- 80
Query: 86 PS-SSP----APSPDEAADNNAIS 104
PS SSP PSP N + S
Sbjct: 81 PSDSSPPVDSTPSPPPPTSNESPS 104
Score = 40.8 bits (94), Expect = 3e-04
Identities = 25/55 (45%), Positives = 31/55 (55%), Gaps = 7/55 (12%)
Query: 48 SPANSPPAPT-----PTPTPSPHSDSPPA-PSPDNSPSSSPSPSPSSSPA-PSPD 95
SP++SPPAP P TPS +S PP SP + P+ S S P S P+ P PD
Sbjct: 7 SPSSSPPAPPADTAPPPETPSENSALPPVDSSPPSPPADSSSTPPLSEPSTPPPD 61
Score = 39.3 bits (90), Expect = 0.001
Identities = 24/60 (40%), Positives = 30/60 (50%), Gaps = 6/60 (10%)
Query: 38 SPSPTPIPTPSPANSPPAPTPTPTPS--PHSDSPPAPSPDNS---PSSSPSPSPSSSPAP 92
SPS +P P P +PP TP+ + P SPP+P D+S P S PS P S P
Sbjct: 7 SPSSSP-PAPPADTAPPPETPSENSALPPVDSSPPSPPADSSSTPPLSEPSTPPPDSQLP 65
Score = 29.6 bits (65), Expect = 0.78
Identities = 13/27 (48%), Positives = 20/27 (73%), Gaps = 1/27 (3%)
Query: 77 SPSSSPSPSPSSSPAPSPDEAADNNAI 103
SPSSSP P+P + AP P+ ++N+A+
Sbjct: 7 SPSSSP-PAPPADTAPPPETPSENSAL 32
>At2g22470 arabinogalactan-protein AGP2
Length = 131
Score = 68.6 bits (166), Expect = 2e-12
Identities = 44/100 (44%), Positives = 56/100 (56%), Gaps = 17/100 (17%)
Query: 13 LSFLVNIASS--ADSPAPTPATNSSLNSPSPTPIP-----TPSPANSPPAPTPTPTPSPH 65
L FL +A+S A +PAP P T + P PT +P TPSP SPP P PTP+P
Sbjct: 9 LIFLGFLATSCLAQAPAPAPTTVT----PPPTALPPVTAETPSPIASPPVPVNEPTPAPT 64
Query: 66 SD--SPPAPSPDNSPSSSPSPS----PSSSPAPSPDEAAD 99
+ + P SP + + +P PS P+SSPAP PD AAD
Sbjct: 65 TSPTTSPVASPPQTDAPAPGPSAGLTPTSSPAPGPDGAAD 104
>At2g18470 putative protein kinase
Length = 633
Score = 68.6 bits (166), Expect = 2e-12
Identities = 58/183 (31%), Positives = 81/183 (43%), Gaps = 45/183 (24%)
Query: 19 IASSADSPAPTPATNSSLNSPSPTPIPTPSP-ANSPPAPTP-----TPTPSPHSDSPP-- 70
+ASS +S PT +T SS + PS T T SP A SPP+PTP + +P P S SPP
Sbjct: 1 MASSPESAPPTNST-SSPSPPSNTNSTTSSPPAPSPPSPTPPQGDSSSSPPPDSTSPPAP 59
Query: 71 -APSPDNSPSSSPS----------------------PSPSSSPAPSPDEAADNNAISHTG 107
AP+P NS ++SPS PS S P+P DN +
Sbjct: 60 QAPNPPNSSNNSPSPPSQGGGGERGNGGNNGGNDTPPSRGSPPSPPSRSNGDNGGSRSSP 119
Query: 108 IGEDGKS-------------SGGGMSSGKKAGIAVGVIAAVGVVALGAMVVKKRRQNIQR 154
G+ G S GGG S I VGV+ G++ + ++V RR+ ++
Sbjct: 120 PGDTGGSRSDNPPSSGGSSGGGGGGRSNTNTAIIVGVLVGAGLLMIVLIIVCLRRKKKRK 179
Query: 155 SEY 157
+
Sbjct: 180 DSF 182
>At4g09030 arabinogalactan-protein AGP10
Length = 127
Score = 67.8 bits (164), Expect = 3e-12
Identities = 41/117 (35%), Positives = 59/117 (50%), Gaps = 6/117 (5%)
Query: 11 LLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPP 70
++L FL IASSA + AP PA S P P P P + P P+ TPTP+P + P
Sbjct: 7 VVLLFLALIASSAIAQAPGPAPTRS-----PLPSPAQPPRTAAPTPSITPTPTPTPSATP 61
Query: 71 APSPDNSPSSSPSPSPSSSPAPSPDEAADNNAIS-HTGIGEDGKSSGGGMSSGKKAG 126
+P + P+ SP PS +S PAP D ++ TG ++ +++G AG
Sbjct: 62 TAAPVSPPAGSPLPSSASPPAPPTSLTPDGAPVAGPTGSTPVDNNNAATLAAGSLAG 118
>At5g38560 putative protein
Length = 681
Score = 66.6 bits (161), Expect = 6e-12
Identities = 47/162 (29%), Positives = 71/162 (43%), Gaps = 28/162 (17%)
Query: 27 APTPATNSSLNSPSPTPI-PTPSPANSPPAPTPTP---TPSPHSDSPPAPSPDNSPSSSP 82
+P P ++S + P+PT P P P+ SPP TP+P TPSP SP P+P + S P
Sbjct: 114 SPPPPPDASPSPPAPTTTNPPPKPSPSPPGETPSPPGETPSPPKPSPSTPTPTTTTSPPP 173
Query: 83 SPSPSSS---------------PAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGI 127
P+ S+S P P P + TG + ++ SS K+ +
Sbjct: 174 PPATSASPPSSNPTDPSTLAPPPTPLPVVPREKPIAKPTGPASNNGNNTLPSSSPGKSEV 233
Query: 128 AVGVIAAVGVV---------ALGAMVVKKRRQNIQRSEYGYT 160
G I A+GV+ +G +KR++ + GYT
Sbjct: 234 GTGGIVAIGVIVGLVFLSLFVMGVWFTRKRKRKDPGTFVGYT 275
Score = 53.9 bits (128), Expect = 4e-08
Identities = 28/65 (43%), Positives = 33/65 (50%), Gaps = 3/65 (4%)
Query: 30 PATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSS 89
P++NSS +P P +P+ PP TP PSP PP S P SP PSSS
Sbjct: 13 PSSNSSTTAPPPLQTQPTTPSAPPPV---TPPPSPPQSPPPVVSSSPPPPVVSSPPPSSS 69
Query: 90 PAPSP 94
P PSP
Sbjct: 70 PPPSP 74
Score = 50.8 bits (120), Expect = 3e-07
Identities = 37/93 (39%), Positives = 46/93 (48%), Gaps = 18/93 (19%)
Query: 16 LVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPS--PHSDSPP 70
+V+ + SP P+P + SP PT +P P SPP TP TP P + SPP
Sbjct: 60 VVSSPPPSSSPPPSPPV---ITSPPPTVASSPPPPVVIASPPPSTPATTPPAPPQTVSPP 116
Query: 71 APSPDNSPS------SSPSPSPSSSP---APSP 94
P PD SPS ++P P PS SP PSP
Sbjct: 117 PP-PDASPSPPAPTTTNPPPKPSPSPPGETPSP 148
Score = 50.4 bits (119), Expect = 4e-07
Identities = 29/77 (37%), Positives = 41/77 (52%), Gaps = 4/77 (5%)
Query: 20 ASSADSPAPTPATNSSLNSPSPTPI-PTPSPANSPP-APTPTPTPSPHSDSPPAPSPDNS 77
+S++ + AP P +P P+ P PSP SPP + +P P S PP+ SP S
Sbjct: 14 SSNSSTTAPPPLQTQPTTPSAPPPVTPPPSPPQSPPPVVSSSPPPPVVSSPPPSSSPPPS 73
Query: 78 PS--SSPSPSPSSSPAP 92
P +SP P+ +SSP P
Sbjct: 74 PPVITSPPPTVASSPPP 90
Score = 48.1 bits (113), Expect = 2e-06
Identities = 28/76 (36%), Positives = 40/76 (51%), Gaps = 8/76 (10%)
Query: 22 SADSPAPTPATNSSLNSPSPTP----IPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNS 77
S+ P P ++ +SP P+P P P+ A+SPP P +P P S PA +P
Sbjct: 53 SSSPPPPVVSSPPPSSSPPPSPPVITSPPPTVASSPPPPVVIASPPP---STPATTPPAP 109
Query: 78 PSS-SPSPSPSSSPAP 92
P + SP P P +SP+P
Sbjct: 110 PQTVSPPPPPDASPSP 125
Score = 45.8 bits (107), Expect = 1e-05
Identities = 24/79 (30%), Positives = 39/79 (48%), Gaps = 9/79 (11%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTP----TPTPSPHSDSPP-----APSPDN 76
P+P + ++S P P+ + P +S P P+P +P P+ S PP +P P
Sbjct: 42 PSPPQSPPPVVSSSPPPPVVSSPPPSSSPPPSPPVITSPPPTVASSPPPPVVIASPPPST 101
Query: 77 SPSSSPSPSPSSSPAPSPD 95
++ P+P + SP P PD
Sbjct: 102 PATTPPAPPQTVSPPPPPD 120
Score = 45.8 bits (107), Expect = 1e-05
Identities = 35/99 (35%), Positives = 45/99 (45%), Gaps = 29/99 (29%)
Query: 25 SPAPTPATNSSLNSPSP---TPIPTPSPANSPP---APTPT-------------PTPSPH 65
SP +P S + P P +P P+ SP SPP +P PT P PS
Sbjct: 43 SPPQSPPPVVSSSPPPPVVSSPPPSSSPPPSPPVITSPPPTVASSPPPPVVIASPPPSTP 102
Query: 66 SDSPPAP----SPDNSPSSSPSP------SPSSSPAPSP 94
+ +PPAP SP P +SPSP +P P+PSP
Sbjct: 103 ATTPPAPPQTVSPPPPPDASPSPPAPTTTNPPPKPSPSP 141
Score = 42.0 bits (97), Expect = 2e-04
Identities = 27/81 (33%), Positives = 37/81 (45%), Gaps = 9/81 (11%)
Query: 27 APTPATNSS-----LNSPSPTPIPTPSP---ANSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
+P P +SS ++SP P+ P PSP + PP +P P SPP +P +P
Sbjct: 47 SPPPVVSSSPPPPVVSSPPPSSSPPPSPPVITSPPPTVASSPPPPVVIASPPPSTPATTP 106
Query: 79 SSSPSP-SPSSSPAPSPDEAA 98
+ P SP P SP A
Sbjct: 107 PAPPQTVSPPPPPDASPSPPA 127
Score = 32.0 bits (71), Expect = 0.16
Identities = 14/42 (33%), Positives = 22/42 (52%)
Query: 56 PTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEA 97
P P +P + S AP P + ++PS P +P PSP ++
Sbjct: 6 PLPILSPPSSNSSTTAPPPLQTQPTTPSAPPPVTPPPSPPQS 47
Score = 30.4 bits (67), Expect = 0.46
Identities = 18/51 (35%), Positives = 22/51 (42%), Gaps = 3/51 (5%)
Query: 43 PIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPS 93
P+P SP +S + T P +P AP P P PSP S P S
Sbjct: 6 PLPILSPPSSNSSTTAPPPLQTQPTTPSAPPPVTPP---PSPPQSPPPVVS 53
>At5g14380 agp6
Length = 150
Score = 66.6 bits (161), Expect = 6e-12
Identities = 51/151 (33%), Positives = 69/151 (44%), Gaps = 34/151 (22%)
Query: 8 LVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSP 64
+V +LL+ + A +AD+P+ +P SPSPT PT +P +P APT P +P
Sbjct: 6 VVLVLLTLTIATAFAADAPSASPK-----KSPSPTAAPTKAPTATTKAPSAPTKAPAAAP 60
Query: 65 HS---DSPPAPSP--------DNSPSSSPS-----PSPSSSPAPSPDEAADNNAISHTGI 108
S SP A SP D+ +SSPS P+ SS PAP+PD +
Sbjct: 61 KSSSASSPKASSPAAEGPVPEDDYSASSPSDSAEAPTVSSPPAPTPDSTS---------- 110
Query: 109 GEDGKSSGGGMSSGKKAGIAVGVIAAVGVVA 139
DG S G S K + + VG VA
Sbjct: 111 AADGPSDGPTAESPKSGAVTTAKFSVVGTVA 141
>At2g14890 arabinogalactan-protein AGP9
Length = 191
Score = 65.9 bits (159), Expect = 1e-11
Identities = 36/82 (43%), Positives = 45/82 (53%), Gaps = 6/82 (7%)
Query: 28 PTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPS--SSPSPS 85
P P + +P P P P+P SPPA P P P+ DSP +PSP +SP SS +P
Sbjct: 94 PPPVASPPPATPPPVATPPPAPLASPPAQVPAPAPTTKPDSP-SPSPSSSPPLPSSDAPG 152
Query: 86 PSS---SPAPSPDEAADNNAIS 104
PS+ SPAPSP + D N S
Sbjct: 153 PSTDSISPAPSPTDVNDQNGAS 174
Score = 64.7 bits (156), Expect = 2e-11
Identities = 43/103 (41%), Positives = 54/103 (51%), Gaps = 11/103 (10%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTP--SPHSDSPPAPSPDNSP 78
SS +P PAT + SP P P+ +P PA PP TP P P SP + P AP+P P
Sbjct: 75 SSPPPASPPPATPPPVASPPP-PVASPPPATPPPVATPPPAPLASPPAQVP-APAPTTKP 132
Query: 79 SSSPSPSPSSSP------APSPDEAADNNAISHTGIGEDGKSS 115
S PSPSPSSSP AP P + + A S T + + +S
Sbjct: 133 DS-PSPSPSSSPPLPSSDAPGPSTDSISPAPSPTDVNDQNGAS 174
Score = 52.4 bits (124), Expect = 1e-07
Identities = 33/83 (39%), Positives = 45/83 (53%), Gaps = 8/83 (9%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTP---TPSPHSDSPPAPSPDNS 77
++A PA P SS SP P TP P SPP P +P TP P + PPAP +
Sbjct: 62 TTAPPPANPPPPVSSPPPASPPPA-TPPPVASPPPPVASPPPATPPPVATPPPAPLA-SP 119
Query: 78 PSSSPSPSPSS---SPAPSPDEA 97
P+ P+P+P++ SP+PSP +
Sbjct: 120 PAQVPAPAPTTKPDSPSPSPSSS 142
Score = 51.2 bits (121), Expect = 3e-07
Identities = 33/95 (34%), Positives = 43/95 (44%), Gaps = 14/95 (14%)
Query: 10 FLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPT--PSPHSD 67
F + + + + APT + P+ TP P P+P PPA TP P P P +
Sbjct: 5 FAIAVICIVLIAGVTGQAPT-------SPPTATPAP-PTPTTPPPAATPPPVSAPPPVTT 56
Query: 68 SPP----APSPDNSPSSSPSPSPSSSPAPSPDEAA 98
SPP AP P N P SP P+S P +P A
Sbjct: 57 SPPPVTTAPPPANPPPPVSSPPPASPPPATPPPVA 91
Score = 50.1 bits (118), Expect = 6e-07
Identities = 30/100 (30%), Positives = 44/100 (44%), Gaps = 13/100 (13%)
Query: 6 FSLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPP----------- 54
F++ + + + + A + PT AT + +P P TP P ++PP
Sbjct: 5 FAIAVICIVLIAGVTGQAPTSPPT-ATPAPPTPTTPPPAATPPPVSAPPPVTTSPPPVTT 63
Query: 55 APTPTPTPSPHSDSPPA-PSPDNSPSSSPSPSPSSSPAPS 93
AP P P P S PPA P P P + P P +SP P+
Sbjct: 64 APPPANPPPPVSSPPPASPPPATPPPVASPPPPVASPPPA 103
Score = 46.2 bits (108), Expect = 8e-06
Identities = 26/72 (36%), Positives = 36/72 (49%), Gaps = 10/72 (13%)
Query: 27 APTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPS--PHSDSPPAPSPDNSPSSSP-- 82
AP P T S P P+ T P +PP P +P P+ P + PP SP +S P
Sbjct: 50 APPPVTTS------PPPVTTAPPPANPPPPVSSPPPASPPPATPPPVASPPPPVASPPPA 103
Query: 83 SPSPSSSPAPSP 94
+P P ++P P+P
Sbjct: 104 TPPPVATPPPAP 115
Score = 45.1 bits (105), Expect = 2e-05
Identities = 29/84 (34%), Positives = 37/84 (43%), Gaps = 9/84 (10%)
Query: 22 SADSPAPTPATNSSLNSPSPTPIP-----TPSPANS--PPAPTPTPTPSPHSDSPPAPSP 74
+A PTP T +P P P +P P + PPA P P SP SPP +P
Sbjct: 28 TATPAPPTPTTPPPAATPPPVSAPPPVTTSPPPVTTAPPPANPPPPVSSPPPASPPPATP 87
Query: 75 DNSPSSSPSPSPSSSPAPSPDEAA 98
P +SP P +S P +P A
Sbjct: 88 --PPVASPPPPVASPPPATPPPVA 109
>At1g20135 anter-specific proline-rich protein APG precursor
Length = 534
Score = 65.5 bits (158), Expect = 1e-11
Identities = 31/70 (44%), Positives = 37/70 (52%), Gaps = 3/70 (4%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPP-APTPTPTPSPHSDSPPAPSPDNSPSSSPSP 84
P P PA + P P P P P P +PP AP P P PSP PPAP+P P P P
Sbjct: 119 PVPPPACPPTPPKPQPKPAPPPEPKPAPPPAPKPVPCPSP--PKPPAPTPKPVPPHGPPP 176
Query: 85 SPSSSPAPSP 94
P+ +P P+P
Sbjct: 177 KPAPAPTPAP 186
Score = 64.7 bits (156), Expect = 2e-11
Identities = 38/110 (34%), Positives = 49/110 (44%), Gaps = 23/110 (20%)
Query: 6 FSLVFLLLSFLVNIASSADS---------------PAPTP-ATNSSLNSPSPTPIPTPSP 49
F +F LLSF + ++ ++ P P P N PSP P+ P P
Sbjct: 16 FRSIFCLLSFCIFFLTTTNAQVMHRRLWPWPLWPRPYPQPWPMNPPTPDPSPKPVAPPGP 75
Query: 50 ANSP-----PAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
++ P P+P P+P P P PPAPSP SP SP P P P P P
Sbjct: 76 SSKPVAPPGPSPCPSPPPKPQPKPPPAPSP--SPCPSPPPKPQPKPVPPP 123
Score = 62.0 bits (149), Expect = 1e-10
Identities = 34/69 (49%), Positives = 41/69 (59%), Gaps = 10/69 (14%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
PAP P + P+P P+P PSP PPAPTP P P PH PP P+P +P+P+
Sbjct: 136 PAPPPEPKPA-PPPAPKPVPCPSPPK-PPAPTPKPVP-PHG-PPPKPAP------APTPA 185
Query: 86 PSSSPAPSP 94
PS PAPSP
Sbjct: 186 PSPKPAPSP 194
Score = 61.2 bits (147), Expect = 2e-10
Identities = 31/69 (44%), Positives = 37/69 (52%), Gaps = 3/69 (4%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
PAP+P+ S P P P P P PA P P P P P+P + PAP P +P P PS
Sbjct: 101 PAPSPSPCPS-PPPKPQPKPVPPPACPPTPPKPQPKPAPPPEPKPAPPP--APKPVPCPS 157
Query: 86 PSSSPAPSP 94
P PAP+P
Sbjct: 158 PPKPPAPTP 166
Score = 60.5 bits (145), Expect = 4e-10
Identities = 32/75 (42%), Positives = 36/75 (47%), Gaps = 11/75 (14%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTPSPHSDSPPA-----PSPDNSPS 79
P P+P SP P P P P PA SP P P+P P P P PPA P P P+
Sbjct: 83 PGPSPCP-----SPPPKPQPKPPPAPSPSPCPSPPPKPQPKPVPPPACPPTPPKPQPKPA 137
Query: 80 SSPSPSPSSSPAPSP 94
P P P+ PAP P
Sbjct: 138 PPPEPKPAPPPAPKP 152
Score = 57.4 bits (137), Expect = 3e-09
Identities = 28/75 (37%), Positives = 38/75 (50%), Gaps = 4/75 (5%)
Query: 27 APTPATNSSLNSPSPTPIPTPSP---ANSPPAPTPTPTPSPHSDSPPAPSPDNS-PSSSP 82
AP ++ + P P+P P+P P PPAP+P+P PSP P P P + P + P
Sbjct: 71 APPGPSSKPVAPPGPSPCPSPPPKPQPKPPPAPSPSPCPSPPPKPQPKPVPPPACPPTPP 130
Query: 83 SPSPSSSPAPSPDEA 97
P P +P P P A
Sbjct: 131 KPQPKPAPPPEPKPA 145
Score = 47.8 bits (112), Expect = 3e-06
Identities = 30/71 (42%), Positives = 33/71 (46%), Gaps = 15/71 (21%)
Query: 26 PAPTPATN-----SSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
PAP PA S P+PTP P P P PP P P PTP+P PAPSP
Sbjct: 144 PAPPPAPKPVPCPSPPKPPAPTPKPVP-PHGPPPKPAPAPTPAP--SPKPAPSP------ 194
Query: 81 SPSPSPSSSPA 91
P P + PA
Sbjct: 195 -PKPENKTIPA 204
Score = 30.8 bits (68), Expect = 0.35
Identities = 13/29 (44%), Positives = 15/29 (50%)
Query: 70 PAPSPDNSPSSSPSPSPSSSPAPSPDEAA 98
P P P N P+ PSP P + P PS A
Sbjct: 53 PQPWPMNPPTPDPSPKPVAPPGPSSKPVA 81
>At5g60630 unknown protein
Length = 139
Score = 63.5 bits (153), Expect = 5e-11
Identities = 42/100 (42%), Positives = 52/100 (52%), Gaps = 3/100 (3%)
Query: 65 HSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGG--GMSSG 122
HS S P + SPS + S + E+ D N T G + S GG G G
Sbjct: 36 HSSSLPQSETEMSPSPTMSNDYDYPSSSQLTESNDLNYTDSTRPGGEEASVGGENGGGGG 95
Query: 123 KKAGIA-VGVIAAVGVVALGAMVVKKRRQNIQRSEYGYTA 161
KK GIA VG IAA +V G V+KKRR+NI+RS YGY +
Sbjct: 96 KKTGIAVVGSIAAASMVGFGGYVLKKRRENIRRSRYGYAS 135
>At5g10430 AtAGP4
Length = 135
Score = 63.2 bits (152), Expect = 6e-11
Identities = 44/111 (39%), Positives = 61/111 (54%), Gaps = 16/111 (14%)
Query: 9 VFLLLSFLVNIASSADSPAPTPATNSSLNSP----SPTPIPTPSPANSPPAPTPTP--TP 62
VFL+L+ A A +PAPTP +P +P P+ TP PA +P TP P TP
Sbjct: 8 VFLMLALFATSAL-AQAPAPTPTATPPPATPPPVATPPPVATPPPAATPAPATPPPAATP 66
Query: 63 SPHSDSPP--APSPDNSPSSS------PSPSPSSSPAPSPDEAADNNAISH 105
+P + +PP APSP + P++S P+ SPSS+P PS A + A S+
Sbjct: 67 AP-ATTPPSVAPSPADVPTASPPAPEGPTVSPSSAPGPSDASPAPSAAFSN 116
>At4g18670 extensin-like protein
Length = 839
Score = 62.8 bits (151), Expect = 8e-11
Identities = 39/87 (44%), Positives = 47/87 (53%), Gaps = 17/87 (19%)
Query: 25 SPAPTPATNSSLNSPS---------PTPIPTPSPANSPPAPT-----PTPTPSPHS--DS 68
SP TP+ S SPS P+P TPSP SPP+P+ P+ TPSP S S
Sbjct: 412 SPPTTPSPGGSPPSPSISPSPPITVPSPPTTPSPGGSPPSPSIVPSPPSTTPSPGSPPTS 471
Query: 69 PPAPSPDNSPSSSP-SPSPSSSPAPSP 94
P P+P SP SSP +P+P SP SP
Sbjct: 472 PTTPTPGGSPPSSPTTPTPGGSPPSSP 498
Score = 62.0 bits (149), Expect = 1e-10
Identities = 38/75 (50%), Positives = 44/75 (58%), Gaps = 6/75 (8%)
Query: 25 SPAPTPATNSSLNSPSPTPIP---TPSPANSPPAPT-PTPTPSPHSDSPPAPSPDNSPSS 80
SP TP+ S SPS P P TPSP + P +PT PTP SP S SP P+P SP S
Sbjct: 438 SPPTTPSPGGSPPSPSIVPSPPSTTPSPGSPPTSPTTPTPGGSPPS-SPTTPTPGGSPPS 496
Query: 81 SPS-PSPSSSPAPSP 94
SP+ P+P SP SP
Sbjct: 497 SPTTPTPGGSPPSSP 511
Score = 60.1 bits (144), Expect = 5e-10
Identities = 39/78 (50%), Positives = 46/78 (58%), Gaps = 7/78 (8%)
Query: 22 SADSPAPTPATNSSLNSPS--PTPIPTPSPANSPPAPTPTPTP--SPHSDSPPAPSPDNS 77
S SP+ P+ S+ SP PT TP+P SPP+ TPTP SP S SP P+P S
Sbjct: 448 SPPSPSIVPSPPSTTPSPGSPPTSPTTPTPGGSPPSSPTTPTPGGSPPS-SPTTPTPGGS 506
Query: 78 PSSSP-SPSPSSSPAPSP 94
P SSP +PSP SP PSP
Sbjct: 507 PPSSPTTPSPGGSP-PSP 523
Score = 58.2 bits (139), Expect = 2e-09
Identities = 35/80 (43%), Positives = 46/80 (56%), Gaps = 7/80 (8%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIP-----TPSPANSPPAPTPTPTPSPHSDSPPA-PSP 74
SS +P P + SS +P+P P TPSP SPP+P+ +P+P SPP+ P+
Sbjct: 483 SSPTTPTPGGSPPSSPTTPTPGGSPPSSPTTPSPGGSPPSPSISPSPPITVPSPPSTPTS 542
Query: 75 DNSPSSSPSPSPSSSPAPSP 94
SP S SP+P SSP PSP
Sbjct: 543 PGSPPSPSSPTP-SSPIPSP 561
Score = 55.1 bits (131), Expect = 2e-08
Identities = 38/78 (48%), Positives = 43/78 (54%), Gaps = 10/78 (12%)
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTP--SPHSDSPPAPSPDNSPS 79
S SP +P T + SP +P TP+P SPP+ TPTP SP S SP PSP SP
Sbjct: 464 SPGSPPTSPTTPTPGGSPPSSPT-TPTPGGSPPSSPTTPTPGGSPPS-SPTTPSPGGSP- 520
Query: 80 SSPSPSPSSSP---APSP 94
PSPS S SP PSP
Sbjct: 521 --PSPSISPSPPITVPSP 536
Score = 54.7 bits (130), Expect = 2e-08
Identities = 30/72 (41%), Positives = 38/72 (52%), Gaps = 3/72 (4%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPA--PTPTPTPSPHSDSPPAPSPDN-SPSSSP 82
P PTP + P PTPI +P P + PP +P P P+ H + PP PSP + SP SP
Sbjct: 724 PPPTPTYHYISPPPPPTPIHSPPPQSHPPCIEYSPPPPPTVHYNPPPPPSPAHYSPPPSP 783
Query: 83 SPSPSSSPAPSP 94
+SP P P
Sbjct: 784 PVYYYNSPPPPP 795
Score = 54.3 bits (129), Expect = 3e-08
Identities = 36/90 (40%), Positives = 45/90 (50%), Gaps = 19/90 (21%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSP------ 74
SS +P+P + S SPSP PI PSP ++P +P P+PS + S P PSP
Sbjct: 509 SSPTTPSPGGSPPSPSISPSP-PITVPSPPSTPTSPGSPPSPSSPTPSSPIPSPPTPSTP 567
Query: 75 -------DNSPSSSPS-----PSPSSSPAP 92
NSP PS PSP SSP+P
Sbjct: 568 PTPISPGQNSPPIIPSPPFTGPSPPSSPSP 597
Score = 53.5 bits (127), Expect = 5e-08
Identities = 31/75 (41%), Positives = 42/75 (55%), Gaps = 8/75 (10%)
Query: 28 PTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTP--SPHS----DSPPA--PSPDNSPS 79
P+P T S P+P +PSP + P+P TP+P SP S SPP+ PSP + P+
Sbjct: 411 PSPPTTPSPGGSPPSPSISPSPPITVPSPPTTPSPGGSPPSPSIVPSPPSTTPSPGSPPT 470
Query: 80 SSPSPSPSSSPAPSP 94
S +P+P SP SP
Sbjct: 471 SPTTPTPGGSPPSSP 485
Score = 53.1 bits (126), Expect = 7e-08
Identities = 37/74 (50%), Positives = 42/74 (56%), Gaps = 6/74 (8%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPS-PANSPPAPTPTPTP-SPHSDSPP-APSPD-NSPSSS 81
P+P P+T +S SP PTPS P SPP P+ PTP SP +SPP PSP PS
Sbjct: 534 PSP-PSTPTSPGSPPSPSSPTPSSPIPSPPTPSTPPTPISPGQNSPPIIPSPPFTGPSPP 592
Query: 82 PSPSPSSSPA-PSP 94
SPSP P PSP
Sbjct: 593 SSPSPPLPPVIPSP 606
Score = 52.4 bits (124), Expect = 1e-07
Identities = 34/76 (44%), Positives = 40/76 (51%), Gaps = 4/76 (5%)
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPT-PSPH-SDSPP--APSPDNS 77
S S TP S S TP P SP +SP P+P + PSP S SPP PSP ++
Sbjct: 480 SPPSSPTTPTPGGSPPSSPTTPTPGGSPPSSPTTPSPGGSPPSPSISPSPPITVPSPPST 539
Query: 78 PSSSPSPSPSSSPAPS 93
P+S SP SSP PS
Sbjct: 540 PTSPGSPPSPSSPTPS 555
Score = 50.8 bits (120), Expect = 3e-07
Identities = 30/68 (44%), Positives = 40/68 (58%), Gaps = 3/68 (4%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSP 84
SP +P T S SP P+P +PSP + P+P TPT SP +P+P +SP SP P
Sbjct: 506 SPPSSPTTPSPGGSP-PSPSISPSPPITVPSPPSTPTSPGSPPSPSSPTP-SSPIPSP-P 562
Query: 85 SPSSSPAP 92
+PS+ P P
Sbjct: 563 TPSTPPTP 570
Score = 48.1 bits (113), Expect = 2e-06
Identities = 32/87 (36%), Positives = 46/87 (52%), Gaps = 13/87 (14%)
Query: 41 PTPIPTPSPANSPPAPTPTPTPSPHSDSPPA--------PSPD---NSPSSSPSP-SPSS 88
P+P TPSP SPP+P+ +P+P SPP PSP + PS++PSP SP +
Sbjct: 411 PSPPTTPSPGGSPPSPSISPSPPITVPSPPTTPSPGGSPPSPSIVPSPPSTTPSPGSPPT 470
Query: 89 SP-APSPDEAADNNAISHTGIGEDGKS 114
SP P+P + ++ + T G S
Sbjct: 471 SPTTPTPGGSPPSSPTTPTPGGSPPSS 497
Score = 44.3 bits (103), Expect = 3e-05
Identities = 33/85 (38%), Positives = 39/85 (45%), Gaps = 16/85 (18%)
Query: 25 SPAPTPATNSSLNSPS-PTP---IPTPSPANSPPAP------TPTPTPSPHSDSPPAPSP 74
SP TP + S SPS PTP IP+P ++PP P +P PSP P PS
Sbjct: 535 SPPSTPTSPGSPPSPSSPTPSSPIPSPPTPSTPPTPISPGQNSPPIIPSPPFTGPSPPSS 594
Query: 75 DNSPSSSPSPSP------SSSPAPS 93
+ P PSP SSP PS
Sbjct: 595 PSPPLPPVIPSPPIVGPTPSSPPPS 619
Score = 43.5 bits (101), Expect = 5e-05
Identities = 26/74 (35%), Positives = 36/74 (48%), Gaps = 8/74 (10%)
Query: 25 SPAPTPATNSSLNSP------SPTPIPTPSPANSPPAPTPTPTPSPHSDSPP--APSPDN 76
SP P+P T S+ +P SP IP+P P +P+P P SPP P+P +
Sbjct: 556 SPIPSPPTPSTPPTPISPGQNSPPIIPSPPFTGPSPPSSPSPPLPPVIPSPPIVGPTPSS 615
Query: 77 SPSSSPSPSPSSSP 90
P S+P+P P
Sbjct: 616 PPPSTPTPGTLLHP 629
Score = 43.5 bits (101), Expect = 5e-05
Identities = 32/83 (38%), Positives = 38/83 (45%), Gaps = 12/83 (14%)
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPA----NSPPA----PTPTPTPSPHSDSPPAP 72
S P+P+ T SS PTP P+P NSPP P P+P P S SPP P
Sbjct: 542 SPGSPPSPSSPTPSSPIPSPPTPSTPPTPISPGQNSPPIIPSPPFTGPSP-PSSPSPPLP 600
Query: 73 SPDNSP---SSSPSPSPSSSPAP 92
SP +PS P S+P P
Sbjct: 601 PVIPSPPIVGPTPSSPPPSTPTP 623
Score = 42.7 bits (99), Expect = 9e-05
Identities = 36/100 (36%), Positives = 44/100 (44%), Gaps = 11/100 (11%)
Query: 2 AIPRFSLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPT----PIPTPSPANSPPAPT 57
A+P L +L +N+AS P P P SS P P P PTP+ P P
Sbjct: 683 ALPLLPL-YLTCRHRLNLASP---PPPAPYYYSSPQPPPPPHYSLPPPTPTYHYISPPPP 738
Query: 58 PTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEA 97
PTP SP S P P + SP P P+ +P P P A
Sbjct: 739 PTPIHSPPPQSHP-PCIEYSPP--PPPTVHYNPPPPPSPA 775
Score = 41.6 bits (96), Expect = 2e-04
Identities = 26/60 (43%), Positives = 34/60 (56%), Gaps = 11/60 (18%)
Query: 43 PIPTPS------PANSPPAPTPTPTPSPHSDSPP-APSPDNSPSSS---PSPSPSSSPAP 92
P+ PS P SPP+P+ +P+P SPP PSP SP S PSP PS++P+P
Sbjct: 407 PVVVPSPPTTPSPGGSPPSPSISPSPPITVPSPPTTPSPGGSPPSPSIVPSP-PSTTPSP 465
Score = 38.5 bits (88), Expect = 0.002
Identities = 31/83 (37%), Positives = 36/83 (43%), Gaps = 15/83 (18%)
Query: 25 SPAPTPATNSSLNSPSPTPI---PTPSPA----NSPPAP-----TPTPTPSPHSDSPPAP 72
SP P P + + P P+P P PSP NSPP P +P P P H PP P
Sbjct: 757 SPPPPPTVHYN-PPPPPSPAHYSPPPSPPVYYYNSPPPPPAVHYSPPPPPVIHHSQPPPP 815
Query: 73 SPDNSPSSSPSPSPS-SSPAPSP 94
P P P S +SP P P
Sbjct: 816 PIYEGP-LPPIPGISYASPPPPP 837
Score = 36.2 bits (82), Expect = 0.008
Identities = 26/80 (32%), Positives = 29/80 (35%), Gaps = 12/80 (15%)
Query: 25 SPAPTPATNSSLNSPSPTPI------PTPSPANSPPAPTP------TPTPSPHSDSPPAP 72
SP P SP P P P PSPA+ P P+P +P P P P P
Sbjct: 744 SPPPQSHPPCIEYSPPPPPTVHYNPPPPPSPAHYSPPPSPPVYYYNSPPPPPAVHYSPPP 803
Query: 73 SPDNSPSSSPSPSPSSSPAP 92
P S P P P P
Sbjct: 804 PPVIHHSQPPPPPIYEGPLP 823
Score = 34.7 bits (78), Expect = 0.024
Identities = 20/42 (47%), Positives = 26/42 (61%), Gaps = 4/42 (9%)
Query: 53 PPAPTPTP--TPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAP 92
PP P+P TPSP SPP+PS SP + PSP ++P+P
Sbjct: 406 PPVVVPSPPTTPSP-GGSPPSPSISPSPPITV-PSPPTTPSP 445
Score = 29.6 bits (65), Expect = 0.78
Identities = 20/54 (37%), Positives = 23/54 (42%), Gaps = 11/54 (20%)
Query: 24 DSPAPTPATNSSLNSPSPTPI-----PTPSPANSPPAPTPTPTPSPHSDSPPAP 72
+SP P PA + SP P P+ P P P P P P P SPP P
Sbjct: 789 NSPPPPPAVH---YSPPPPPVIHHSQPPPPPIYEGPLP---PIPGISYASPPPP 836
>At1g26150 Pto kinase interactor, putative
Length = 760
Score = 62.8 bits (151), Expect = 8e-11
Identities = 36/83 (43%), Positives = 45/83 (53%), Gaps = 12/83 (14%)
Query: 25 SPAPTPATNS-SLNSPSPTPIP-TPSPANSPPAPTPTPTPSPHS--------DSPPAPSP 74
SP P P+ S SL P PT IP +P P SPP P PT P P + SPP P P
Sbjct: 70 SPPPEPSPPSPSLTGPPPTTIPVSPPPEPSPPPPLPTEAPPPANPVSSPPPESSPPPPPP 129
Query: 75 DNSPSSSP--SPSPSSSPAPSPD 95
+P ++P SPSP ++P P P+
Sbjct: 130 TEAPPTTPITSPSPPTNPPPPPE 152
Score = 59.3 bits (142), Expect = 9e-10
Identities = 55/166 (33%), Positives = 71/166 (42%), Gaps = 43/166 (25%)
Query: 29 TPATNSSLNSPSPTPIPTPSPANSPPAP-TPTPTPSPHSDS------------------- 68
TP ++S PSP P P PP P + PTPSP S S
Sbjct: 209 TPPSDSE--HPSPPPPGHPKRREQPPPPGSKRPTPSPPSPSDSKRPVHPSPPSPPEETLP 266
Query: 69 PPAPSPDNSPSSSPSP----------SPSSSPAPS---PDEAADNNAISHTGIGEDGKSS 115
PP PSPD PS+S SP SP S P S PD + NN T SS
Sbjct: 267 PPKPSPDPLPSNSSSPPTLLPPSSVVSPPSPPRKSVSGPDNPSPNNPTPVT-----DNSS 321
Query: 116 GGGMSSGKKAGIAVGVIAAVGVVALGAMV--VKKRRQNIQRSEYGY 159
G+S G+++GV A V + +G +V +KKR++ + GY
Sbjct: 322 SSGISIAAVVGVSIGV-ALVLLTLIGVVVCCLKKRKKRLSTIGGGY 366
Score = 57.4 bits (137), Expect = 3e-09
Identities = 27/69 (39%), Positives = 35/69 (50%), Gaps = 3/69 (4%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P + ++SP P P P P P TP +PSP ++ PP P SP S P+P
Sbjct: 105 PTEAPPPANPVSSPPPESSPPPPPPTEAPPTTPITSPSPPTNPPP---PPESPPSLPAPD 161
Query: 86 PSSSPAPSP 94
P S+P P P
Sbjct: 162 PPSNPLPPP 170
Score = 56.6 bits (135), Expect = 6e-09
Identities = 33/85 (38%), Positives = 43/85 (49%), Gaps = 5/85 (5%)
Query: 18 NIASSADSPAPT-PATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDN 76
N + ++PA + P + L+SP P P P PSP+ + P PT P P SPP P P
Sbjct: 49 NPPETTNTPAQSSPPPETPLSSPPPEPSP-PSPSLTGPPPTTIPVSPPPEPSPPPPLPTE 107
Query: 77 SPSSS---PSPSPSSSPAPSPDEAA 98
+P + SP P SSP P P A
Sbjct: 108 APPPANPVSSPPPESSPPPPPPTEA 132
Score = 52.8 bits (125), Expect = 9e-08
Identities = 32/92 (34%), Positives = 41/92 (43%), Gaps = 13/92 (14%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPPAP------------TPTPTPSPHSDSPPAP 72
+P T T + + P TP+ +P P SPP+P +P P PSP P
Sbjct: 49 NPPETTNTPAQSSPPPETPLSSPPPEPSPPSPSLTGPPPTTIPVSPPPEPSPPPPLPTEA 108
Query: 73 SPDNSPSSSPSPSPSSSPAPSPDEAADNNAIS 104
P +P SSP P SS P P P EA I+
Sbjct: 109 PPPANPVSSPPPE-SSPPPPPPTEAPPTTPIT 139
Score = 51.2 bits (121), Expect = 3e-07
Identities = 31/76 (40%), Positives = 36/76 (46%), Gaps = 10/76 (13%)
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNS---PSSS 81
SP P P T + P TPI +PSP +PP P +P P D P P P PS S
Sbjct: 123 SPPPPPPTEA----PPTTPITSPSPPTNPPPPPESPPSLPAPDPPSNPLPPPKLVPPSHS 178
Query: 82 PS---PSPSSSPAPSP 94
P PSP +S P P
Sbjct: 179 PPRHLPSPPASEIPPP 194
Score = 48.1 bits (113), Expect = 2e-06
Identities = 34/100 (34%), Positives = 40/100 (40%), Gaps = 26/100 (26%)
Query: 21 SSADSPAPTPATNSS-LNSPSPTPIPTPSPANSPPAPTPTPTPSP--------------- 64
SS P PT A ++ + SPSP P P P + P P P P +P
Sbjct: 122 SSPPPPPPTEAPPTTPITSPSPPTNPPPPPESPPSLPAPDPPSNPLPPPKLVPPSHSPPR 181
Query: 65 HSDSPPA----------PSPDNSPSSSPSPSPSSSPAPSP 94
H SPPA PSP S S PS S P+P P
Sbjct: 182 HLPSPPASEIPPPPRHLPSPPASERPSTPPSDSEHPSPPP 221
Score = 47.0 bits (110), Expect = 5e-06
Identities = 33/97 (34%), Positives = 43/97 (44%), Gaps = 17/97 (17%)
Query: 26 PAPTPATNS----SLNSPSPTP---IPTPSPANSPPAPTPTPTP-------SPHSDSP-P 70
PAP P +N L PS +P +P+P + PP P P+P +P SDS P
Sbjct: 158 PAPDPPSNPLPPPKLVPPSHSPPRHLPSPPASEIPPPPRHLPSPPASERPSTPPSDSEHP 217
Query: 71 APSPDNSPSSSPSPSPSSS--PAPSPDEAADNNAISH 105
+P P P P P S P PSP +D+ H
Sbjct: 218 SPPPPGHPKRREQPPPPGSKRPTPSPPSPSDSKRPVH 254
Score = 46.6 bits (109), Expect = 6e-06
Identities = 28/70 (40%), Positives = 36/70 (51%), Gaps = 9/70 (12%)
Query: 27 APTPATNSSLNSPSPTPIPTPSPANSPP-----APTPTPTPSPHSDSPPA-PSPDNSPSS 80
AP PA S P +P P P P +PP +P+P P P +SPP+ P+PD P S
Sbjct: 108 APPPANPVSSPPPESSP-PPPPPTEAPPTTPITSPSPPTNPPPPPESPPSLPAPD--PPS 164
Query: 81 SPSPSPSSSP 90
+P P P P
Sbjct: 165 NPLPPPKLVP 174
Score = 46.6 bits (109), Expect = 6e-06
Identities = 32/84 (38%), Positives = 39/84 (46%), Gaps = 10/84 (11%)
Query: 24 DSPAPTPATNSS--LNSPSPTPIPTPSPANSPPA-PTPTPTPSPHSDSPP--APSPDNS- 77
++P TP T+ S N P P P PA PP+ P P P P S SPP PSP S
Sbjct: 131 EAPPTTPITSPSPPTNPPPPPESPPSLPAPDPPSNPLPPPKLVPPSHSPPRHLPSPPASE 190
Query: 78 ----PSSSPSPSPSSSPAPSPDEA 97
P PSP S P+ P ++
Sbjct: 191 IPPPPRHLPSPPASERPSTPPSDS 214
Score = 42.7 bits (99), Expect = 9e-05
Identities = 28/78 (35%), Positives = 36/78 (45%), Gaps = 11/78 (14%)
Query: 26 PAPTPATNSSLNSPSPTPIPT----PSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSS 81
P P P+ N+ SPT PT P N+P +P P +P S PP PSP + +
Sbjct: 27 PPPQPSFPGD-NATSPTREPTNGNPPETTNTPAQSSP-PPETPLSSPPPEPSPPSPSLTG 84
Query: 82 PSP-----SPSSSPAPSP 94
P P SP P+P P
Sbjct: 85 PPPTTIPVSPPPEPSPPP 102
Score = 42.7 bits (99), Expect = 9e-05
Identities = 24/76 (31%), Positives = 37/76 (48%), Gaps = 1/76 (1%)
Query: 19 IASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
+ + A +P + + SL SP +P P P+ P +PT P + +PP + +
Sbjct: 1 MTTPAQAPREEVSLSPSLASPPLMALPPPQPS-FPGDNATSPTREPTNGNPPETTNTPAQ 59
Query: 79 SSSPSPSPSSSPAPSP 94
SS P +P SSP P P
Sbjct: 60 SSPPPETPLSSPPPEP 75
Score = 41.6 bits (96), Expect = 2e-04
Identities = 28/74 (37%), Positives = 34/74 (45%), Gaps = 9/74 (12%)
Query: 29 TPATNSSLNSPSPTPIPTPSPANSPP-APTPTPTPSPHSDSPPAPSPDNSPSS----SPS 83
T T N P TP+ ++ PP P +P P P SPP+PS P + SP
Sbjct: 39 TSPTREPTNGNPPETTNTPAQSSPPPETPLSSPPPEP---SPPSPSLTGPPPTTIPVSPP 95
Query: 84 PSPSSSPAPSPDEA 97
P P S P P P EA
Sbjct: 96 PEP-SPPPPLPTEA 108
>At5g64310 arabinogalactan-protein AGP1 (gb|AAC77823.1)
Length = 131
Score = 62.4 bits (150), Expect = 1e-10
Identities = 51/135 (37%), Positives = 72/135 (52%), Gaps = 12/135 (8%)
Query: 7 SLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHS 66
SLVF+LL+ L+ ++ A SPAP P S++ +P P+P +P AP P +PSP
Sbjct: 6 SLVFVLLAALLISSAVAQSPAPAP---SNVGGRRISPAPSPKKMTAP-APAPEVSPSP-- 59
Query: 67 DSPPAPSPDNSPSSSPSPSPSSSP-APSPDEAADNNAISHTGIGEDGKSSGGGMSS--GK 123
SP A S +S PSP + SP A SP A +AIS + G + GG +S+
Sbjct: 60 -SPAAALTPESSASPPSPPLADSPTADSP--ALSPSAISDSPTEAPGPAQGGAVSNKFAS 116
Query: 124 KAGIAVGVIAAVGVV 138
+AV + AAV V+
Sbjct: 117 FGSVAVMLTAAVLVI 131
>At2g24450 predicted GPI-anchored protein
Length = 280
Score = 62.4 bits (150), Expect = 1e-10
Identities = 36/105 (34%), Positives = 57/105 (54%), Gaps = 14/105 (13%)
Query: 40 SPTPIPTPSPANSPPAP-------TPTPTPS---PHSDSPPAPSPDNSPSSSPSPSPSSS 89
SP +P P P +SPPAP TP P P+ ++D+PP +P+ +P+S +PS S S
Sbjct: 176 SPVKVPPPPPMSSPPAPSPKKGAATPAPAPADEGDYADAPPGLAPETAPAS--APSESDS 233
Query: 90 PAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAA 134
PAP+PD++ + +S G+S G A + +G +A+
Sbjct: 234 PAPAPDKSGKKKMAAADEAEPPSSASNTGLSFG--AVLVLGFVAS 276
Score = 42.7 bits (99), Expect = 9e-05
Identities = 25/81 (30%), Positives = 38/81 (46%), Gaps = 8/81 (9%)
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPP--APTPTPTPSPHSDSPPAPSPDNSPSSSPS 83
PAP+P ++ +P+P P A++PP AP P +P PAP+PD S +
Sbjct: 190 PAPSPKKGAA--TPAPAPADEGDYADAPPGLAPETAPASAPSESDSPAPAPDKSGKKKMA 247
Query: 84 PSPSSSPAPSPDEAADNNAIS 104
+ + P S A N +S
Sbjct: 248 AADEAEPPSS----ASNTGLS 264
Score = 31.2 bits (69), Expect = 0.27
Identities = 18/55 (32%), Positives = 26/55 (46%), Gaps = 3/55 (5%)
Query: 48 SPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSP---SSSPAPSPDEAAD 99
+P N P +P SP P SS P+PSP +++PAP+P + D
Sbjct: 156 NPYNLSVVQISMPIVAPGLGSPVKVPPPPPMSSPPAPSPKKGAATPAPAPADEGD 210
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.305 0.124 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,674,046
Number of Sequences: 26719
Number of extensions: 328249
Number of successful extensions: 21239
Number of sequences better than 10.0: 1029
Number of HSP's better than 10.0 without gapping: 698
Number of HSP's successfully gapped in prelim test: 351
Number of HSP's that attempted gapping in prelim test: 2751
Number of HSP's gapped (non-prelim): 6465
length of query: 166
length of database: 11,318,596
effective HSP length: 92
effective length of query: 74
effective length of database: 8,860,448
effective search space: 655673152
effective search space used: 655673152
T: 11
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144405.9


