

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142507.1 - phase: 0
(542 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
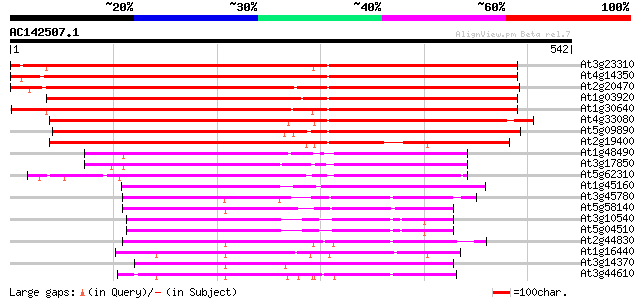
Score E
Sequences producing significant alignments: (bits) Value
At3g23310 protein kinase like protein 736 0.0
At4g14350 protein kinase 731 0.0
At2g20470 putative protein kinase 677 0.0
At1g03920 putative protein kinase 667 0.0
At1g30640 putative protein kinase 625 e-179
At4g33080 protein kinase like protein 580 e-166
At5g09890 protein kinase 556 e-159
At2g19400 putative protein kinase 527 e-150
At1g48490 IRE - like protein kinase 262 3e-70
At3g17850 protein kinase like protein 255 4e-68
At5g62310 IRE (root hair elongation) 254 1e-67
At1g45160 similar to IRE homolog 1 dbj|BAA89784.1 248 6e-66
At3g45780 nonphototropic hypocotyl 1 (NPH1) 184 1e-46
At5g58140 non phototropic hypocotyl 1-like (NPL1) 177 1e-44
At3g10540 3-phosphoinositide-dependent protein kinase-1 like 175 5e-44
At5g04510 3-phosphoinositide-dependent protein kinase-1 (PDK1) 172 3e-43
At2g44830 protein kinase like protein 171 9e-43
At1g16440 putative protein kinase 168 6e-42
At3g14370 putative protein kinase 157 1e-38
At3g44610 protein kinase-like protein 157 1e-38
>At3g23310 protein kinase like protein
Length = 568
Score = 736 bits (1901), Expect = 0.0
Identities = 362/500 (72%), Positives = 424/500 (84%), Gaps = 13/500 (2%)
Query: 1 METARRWFRKRWLKKKEATSTSNDHQCLALDLVN-------EEPPSNETKLKVEAAKHFV 53
M+TAR W +K LK K +SN + ++ EE SN TK K AAK ++
Sbjct: 1 MDTARAWLKK--LKSKGKEKSSNKKETSRGNVKEGSKTAGGEEAVSNVTKQKAAAAKQYI 58
Query: 54 ESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQRLKMGAD 113
E++YK Q Q+QQ+RKERR++LENKLA AEVS+E++KNLLK+ E+ ETE MRRQR KMG D
Sbjct: 59 ENHYKKQVQSQQQRKERRDMLENKLAAAEVSEEEQKNLLKDLEKKETEYMRRQRHKMGTD 118
Query: 114 DFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDSN 173
DFEPLTMIGKGAFGEVRICREKTTG VYAMKKL+KSEMLRRGQVE+VK+ERNLLAEVDSN
Sbjct: 119 DFEPLTMIGKGAFGEVRICREKTTGNVYAMKKLKKSEMLRRGQVEHVKAERNLLAEVDSN 178
Query: 174 CIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHKHN 233
CIVKLYCSFQD+EYLYLIMEYLPGGDMMTLLMR++ LTE+EA+FY+GETVLAIESIHKHN
Sbjct: 179 CIVKLYCSFQDEEYLYLIMEYLPGGDMMTLLMRKDTLTEDEARFYVGETVLAIESIHKHN 238
Query: 234 YIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDELTLSP- 292
YIHRDIKPDNLLLDR+GH+KLSDFGLCKPLDCS LQEKD N SG LQSD ++P
Sbjct: 239 YIHRDIKPDNLLLDRSGHMKLSDFGLCKPLDCSILQEKDFVVAHNLSGALQSDGRPVAPR 298
Query: 293 -NQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVG 351
+SQ +QL++W++N RMLAYSTVGTPDYIAPEV+LK+GYG+ECDWWSLGAIMYEMLVG
Sbjct: 299 RTRSQMEQLQNWQRNR-RMLAYSTVGTPDYIAPEVLLKKGYGMECDWWSLGAIMYEMLVG 357
Query: 352 YPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLLCNVKQRLGAKGADEIKA 410
+PPF+SD+P TTCRKI +WK YLKFP + +LSPEAKDLICRLLCNV+QR+G KGA+EIK
Sbjct: 358 FPPFYSDEPMTTCRKIVNWKNYLKFPDEVRLSPEAKDLICRLLCNVEQRIGTKGANEIKE 417
Query: 411 HPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEEDQKTEPSPKAGPWRKMLQSKDI 470
HPWF VEWEKLYQM+A FIP+VNDELDT+NF+KFEE D++ +PK+GPWRKML SKDI
Sbjct: 418 HPWFSGVEWEKLYQMKAAFIPQVNDELDTQNFEKFEETDKQVPKTPKSGPWRKMLSSKDI 477
Query: 471 NFVGYTYKNYEMVDANAIPG 490
NFVGYTYKN E+V+ + +PG
Sbjct: 478 NFVGYTYKNVEIVNHDQLPG 497
>At4g14350 protein kinase
Length = 551
Score = 731 bits (1887), Expect = 0.0
Identities = 364/498 (73%), Positives = 418/498 (83%), Gaps = 12/498 (2%)
Query: 1 METARRWFRK-------RWLKKKEATSTSNDHQCLALDLVNEEPPSNETKLKVEAAKHFV 53
META+ W K + KKKEATS + A EE SN TK K AAK ++
Sbjct: 1 METAKAWLSKLKSKDKVKSSKKKEATSNVKEGPKTA---GGEEALSNITKEKAAAAKLYI 57
Query: 54 ESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQRLKMGAD 113
E++YK Q Q+ QERKERR +LE KLA AEVS+E++ NLLK+ E ETE MRRQR KMGAD
Sbjct: 58 ENHYKMQMQSLQERKERRKMLEKKLAAAEVSEEEQNNLLKDLEMKETEYMRRQRHKMGAD 117
Query: 114 DFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDSN 173
DFEPLTMIGKGAFGEVRICREK TG VYAMKKL+KSEMLRRGQVE+VK+ERNLLAEVDSN
Sbjct: 118 DFEPLTMIGKGAFGEVRICREKGTGNVYAMKKLKKSEMLRRGQVEHVKAERNLLAEVDSN 177
Query: 174 CIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHKHN 233
CIVKLYCSFQD+EYLYLIMEYLPGGDMMTLLMR++ LTE+EA+FYIGETVLAIESIHKHN
Sbjct: 178 CIVKLYCSFQDEEYLYLIMEYLPGGDMMTLLMRKDTLTEDEARFYIGETVLAIESIHKHN 237
Query: 234 YIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDELTLSPN 293
YIHRDIKPDNLLLD++GH+KLSDFGLCKPLDCSNLQEKD + N SG LQSD ++
Sbjct: 238 YIHRDIKPDNLLLDKDGHMKLSDFGLCKPLDCSNLQEKDFTVARNVSGALQSDGRPVATR 297
Query: 294 QSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYP 353
++QQ+QL +W++N RMLAYSTVGTPDYIAPEV+LK+GYG+ECDWWSLGAIMYEMLVG+P
Sbjct: 298 RTQQEQLLNWQRNR-RMLAYSTVGTPDYIAPEVLLKKGYGMECDWWSLGAIMYEMLVGFP 356
Query: 354 PFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLLCNVKQRLGAKGADEIKAHP 412
PF+SDDP TTCRKI +W+ YLKFP + +LSPEAKDLICRLLCNV+QRLG KGADEIK HP
Sbjct: 357 PFYSDDPMTTCRKIVNWRNYLKFPDEVRLSPEAKDLICRLLCNVEQRLGTKGADEIKGHP 416
Query: 413 WFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEEDQKTEPSPKAGPWRKMLQSKDINF 472
WF+ EW KLYQM+A FIP+VNDELDT+NF+KFEE D++ S K+GPWRKML SKDINF
Sbjct: 417 WFRGTEWGKLYQMKAAFIPQVNDELDTQNFEKFEETDKQVPKSAKSGPWRKMLSSKDINF 476
Query: 473 VGYTYKNYEMVDANAIPG 490
VGYTYKN E+V+ + IPG
Sbjct: 477 VGYTYKNVEIVNDDQIPG 494
>At2g20470 putative protein kinase
Length = 596
Score = 677 bits (1747), Expect = 0.0
Identities = 329/505 (65%), Positives = 412/505 (81%), Gaps = 19/505 (3%)
Query: 1 METARRWFRKRWLKKKE------------ATSTSNDHQCLALDLVNEEPPSNETKLKVEA 48
M++A+ WF+KR ++ ++ S D + D EE SN TK KV A
Sbjct: 1 MDSAKGWFQKRQMRGGSRYKGASGGAGGGGSNGSADEHNVETD---EEAVSNTTKQKVAA 57
Query: 49 AKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQRL 108
AK ++E++YK Q + QERKERR++LE KLADA+VS+ED+ NLLK E+ ETE MR QR
Sbjct: 58 AKQYIENHYKEQMKILQERKERRSMLEQKLADADVSEEDQNNLLKFLEKKETEYMRLQRH 117
Query: 109 KMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLA 168
K+G DF+ LTMIGKGAFGEVR+CREKTTGQVYAMKKL+K+EMLRRGQVE+V++ERNLLA
Sbjct: 118 KLGVADFDLLTMIGKGAFGEVRVCREKTTGQVYAMKKLKKAEMLRRGQVEHVRAERNLLA 177
Query: 169 EVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIES 228
EVDSN IVKLYCSFQDD++LYL+MEYLPGGDMMTLLMR++ LTE EAKFY+ ETVLAIES
Sbjct: 178 EVDSNYIVKLYCSFQDDDHLYLVMEYLPGGDMMTLLMRKDTLTEEEAKFYVAETVLAIES 237
Query: 229 IHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDEL 288
IH+HNYIHRDIKPDNLLLDR GH++LSDFGLCKPLDCS + E D S+ N +G + +
Sbjct: 238 IHRHNYIHRDIKPDNLLLDRYGHLRLSDFGLCKPLDCSAIGENDFSN--NSNGSTEQEAG 295
Query: 289 TLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEM 348
+ +P ++QQ+QLEHW++N R LAYSTVGTPDYIAPEV+LK+GYG+ECDWWSLGAIMYEM
Sbjct: 296 STAPKRTQQEQLEHWQRNR-RTLAYSTVGTPDYIAPEVLLKKGYGMECDWWSLGAIMYEM 354
Query: 349 LVGYPPFHSDDPRTTCRKIAHWKTYLKFPKDK-LSPEAKDLICRLLCNVKQRLGAKGADE 407
LVGYPPF+SDDP +TCRKI +WK++LKFP++ LS EAKDLI LLC+V++RLG+KGADE
Sbjct: 355 LVGYPPFYSDDPMSTCRKIVNWKSHLKFPEEAILSREAKDLINSLLCSVRRRLGSKGADE 414
Query: 408 IKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEEDQKTEPSPKAGPWRKMLQS 467
+KAH WF+ V+W+ ++ M A F+P+VND+LDT+NF+KF+E + +T+ S K+GPWRKML S
Sbjct: 415 LKAHTWFETVDWDTIFDMDAAFVPEVNDDLDTQNFEKFDESESETQTSSKSGPWRKMLSS 474
Query: 468 KDINFVGYTYKNYEMVDANAIPGFG 492
KDINFVGYTYKN+E+V+ +PG G
Sbjct: 475 KDINFVGYTYKNFEIVNDYQVPGMG 499
>At1g03920 putative protein kinase
Length = 569
Score = 667 bits (1722), Expect = 0.0
Identities = 323/456 (70%), Positives = 388/456 (84%), Gaps = 3/456 (0%)
Query: 36 EPPSNETKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNF 95
E SN TK KV AAK ++E++YK Q +N ERKERR LE KLADA+V +ED+ NL+K
Sbjct: 58 EALSNSTKQKVAAAKQYIENHYKEQMKNLNERKERRTTLEKKLADADVCEEDQTNLMKFL 117
Query: 96 EEMETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRG 155
E+ ETE MR QR KMGADDFE LTMIGKGAFGEVR+ RE TG V+AMKKL+KSEMLRRG
Sbjct: 118 EKKETEYMRLQRHKMGADDFELLTMIGKGAFGEVRVVREINTGHVFAMKKLKKSEMLRRG 177
Query: 156 QVEYVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEA 215
QVE+V++ERNLLAEVDSNCIVKLYCSFQD+EYLYLIMEYLPGGDMMTLLMR++ L+E+EA
Sbjct: 178 QVEHVRAERNLLAEVDSNCIVKLYCSFQDNEYLYLIMEYLPGGDMMTLLMRKDTLSEDEA 237
Query: 216 KFYIGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSD 275
KFYI E+VLAIESIH NYIHRDIKPDNLLLDR GH++LSDFGLCKPLDCS + +D +
Sbjct: 238 KFYIAESVLAIESIHNRNYIHRDIKPDNLLLDRYGHLRLSDFGLCKPLDCSVIDGEDFTV 297
Query: 276 GVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVE 335
G SG S+ ++ +P +SQQ+QLEHW+KN RMLAYSTVGTPDYIAPEV+LK+GYG+E
Sbjct: 298 GNAGSGG-GSESVSTTPKRSQQEQLEHWQKNR-RMLAYSTVGTPDYIAPEVLLKKGYGME 355
Query: 336 CDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLLC 394
CDWWSLGAIMYEMLVGYPPF++DDP +TCRKI +WKT+LKFP++ +LS A+DLI +LLC
Sbjct: 356 CDWWSLGAIMYEMLVGYPPFYADDPMSTCRKIVNWKTHLKFPEESRLSRGARDLIGKLLC 415
Query: 395 NVKQRLGAKGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEEDQKTEP 454
+V QRLG+ GA +IKAHPWF+ V+WEK+YQM A FIP+VND+LDT+NF+KF+EED +T+
Sbjct: 416 SVNQRLGSTGASQIKAHPWFEGVQWEKIYQMEAAFIPEVNDDLDTQNFEKFDEEDNQTQA 475
Query: 455 SPKAGPWRKMLQSKDINFVGYTYKNYEMVDANAIPG 490
+ GPWRKML SKDINFVGYTYKN+E+V+ +PG
Sbjct: 476 PSRTGPWRKMLSSKDINFVGYTYKNFEIVNDYQVPG 511
>At1g30640 putative protein kinase
Length = 562
Score = 625 bits (1611), Expect = e-179
Identities = 312/498 (62%), Positives = 390/498 (77%), Gaps = 11/498 (2%)
Query: 2 ETARRWFRKRWLKKKEATSTSNDHQCLALDLVN---EEPPSNETKLKVEAAKHFVESYYK 58
E R WF+ R K +++ T + +L + PSN TK KV AAK ++E++YK
Sbjct: 4 EPTRSWFQIRQQKPDKSSPTKKGQEGNVKNLGRPPMNDAPSNATKQKVAAAKQYIENHYK 63
Query: 59 NQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQRLKMGADDFEPL 118
QK++ QERKERR+ILE LADA+V+ EDK ++LKNFE+ E E MR QR KMG DDFE L
Sbjct: 64 IQKKSLQERKERRSILEQNLADADVTVEDKMDILKNFEKKEMEYMRLQRQKMGVDDFELL 123
Query: 119 TMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDSNCIVKL 178
++IG+GAFGEVRIC+EK+TG VYAMKKL+KSEMLRRGQVE+VK+ERN+LAEVDS IVKL
Sbjct: 124 SIIGRGAFGEVRICKEKSTGSVYAMKKLKKSEMLRRGQVEHVKAERNVLAEVDSPFIVKL 183
Query: 179 YCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHKHNYIHRD 238
SFQDDE+LYLIMEYLPGGDMMTLLMR++ L E+E +FY+ +T+LAIESIHKHNY+HRD
Sbjct: 184 CYSFQDDEHLYLIMEYLPGGDMMTLLMRKDTLREDETRFYVAQTILAIESIHKHNYVHRD 243
Query: 239 IKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDELTLS-----PN 293
IKPDNLL+ RNGH+KLSDFGL K L+ N + ++ V+RS ++ LS P
Sbjct: 244 IKPDNLLITRNGHIKLSDFGLSKSLESKNFPDFK-AELVDRSTKPAAEHDRLSKPPSAPR 302
Query: 294 QSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYP 353
++QQ+QL HW++N R LA+STVGTPDYIAPEV+LK+GYG+ECDWWSLGAIM+EMLVG+P
Sbjct: 303 RTQQEQLLHWQQNR-RTLAFSTVGTPDYIAPEVLLKKGYGMECDWWSLGAIMFEMLVGFP 361
Query: 354 PFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLLCNVKQRLGAKGADEIKAHP 412
PF+S++P TCRKI +WKT LKFP + KLS E KDLI RLLCNV+QRLG KG EIKAHP
Sbjct: 362 PFYSEEPLATCRKIVNWKTCLKFPDEAKLSIEVKDLIRRLLCNVEQRLGTKGVHEIKAHP 421
Query: 413 WFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEEDQKTEPSPKAGPWRKMLQSKDINF 472
WF+ VEWE+LY+ AP+IP+V ELDT+NF+KF+E + S K+ PWRKM+ SKD NF
Sbjct: 422 WFRGVEWERLYESNAPYIPQVKHELDTQNFEKFDEVPSTCQTSSKSSPWRKMISSKDANF 481
Query: 473 VGYTYKNYEMVDANAIPG 490
+GYT+KN E+VD + IPG
Sbjct: 482 LGYTFKNLEIVDEHHIPG 499
>At4g33080 protein kinase like protein
Length = 519
Score = 580 bits (1494), Expect = e-166
Identities = 295/479 (61%), Positives = 369/479 (76%), Gaps = 17/479 (3%)
Query: 39 SNETKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEM 98
S+ T KV AAK ++E++YK Q +N QERKERR ILE KLA + V KE++ N++K+ E
Sbjct: 18 SSLTMEKVAAAKQYIENHYKAQNKNIQERKERRWILERKLASSGVPKEEQINMIKDLERK 77
Query: 99 ETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVE 158
ETE MR +R K+ DDFE LT+IG+GAFGEVR+CRE+ +G +YAMKKL+KSEM+ RGQVE
Sbjct: 78 ETEFMRLKRNKISVDDFELLTIIGRGAFGEVRLCRERKSGNIYAMKKLKKSEMVMRGQVE 137
Query: 159 YVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFY 218
+V++ERNLLAEV+S+ IVKLY SFQD EYLYLIMEYLPGGDMMTLLMR + L E+ A+FY
Sbjct: 138 HVRAERNLLAEVESHYIVKLYYSFQDPEYLYLIMEYLPGGDMMTLLMREDTLREDVARFY 197
Query: 219 IGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNL---QEKDLSD 275
I ++VLAIESIH++NYIHRDIKPDNLLLD++GH+KLSDFGLCKPLDC NL QE +D
Sbjct: 198 IAQSVLAIESIHRYNYIHRDIKPDNLLLDKDGHMKLSDFGLCKPLDCRNLPSIQENRATD 257
Query: 276 GVNRSGVLQSDELTLSPN-----QSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKR 330
S + D + +S Q+QL+HW+ N R LA+STVGTPDYIAPEV+LK+
Sbjct: 258 DETMSEPMDVDRCFPDTDNKRSWRSPQEQLQHWQMNR-RKLAFSTVGTPDYIAPEVLLKK 316
Query: 331 GYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLI 389
GYG+ECDWWSLGAIMYEMLVGYPPF++DDP +TCRKI HW+ +LKFP+D K S EAKDLI
Sbjct: 317 GYGMECDWWSLGAIMYEMLVGYPPFYADDPISTCRKIVHWRNHLKFPEDAKFSSEAKDLI 376
Query: 390 CRLLCNVKQRLG-AKGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEE 448
CRLLCNV RLG GA +IK HPWFK V WEKLY+M A + P+VNDELDT+NF KF+E
Sbjct: 377 CRLLCNVDHRLGTGGGAQQIKDHPWFKDVVWEKLYEMEAAYKPEVNDELDTQNFMKFDEV 436
Query: 449 DQKTEPSPKAGPWRKMLQS-KDINFVGYTYKNYEMVDANAIPGFGMFISLSLSLSLSHS 506
+ ++G RKML + KD++FVGYTYKN++ A+ G + ++ ++SL S
Sbjct: 437 NSPAPERTRSGLSRKMLLAPKDLSFVGYTYKNFD-----AVKGLRHSLEMARTMSLDRS 490
>At5g09890 protein kinase
Length = 515
Score = 556 bits (1434), Expect = e-159
Identities = 274/459 (59%), Positives = 350/459 (75%), Gaps = 11/459 (2%)
Query: 42 TKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEMETE 101
T+ K AAK F+E++YKN Q ER ERR + K+ +A++ E++ +++N ETE
Sbjct: 29 TRQKAAAAKQFIENHYKNYLQGLHERMERRREFQRKVQEAQLPVEEQDEMMRNLARRETE 88
Query: 102 IMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVK 161
MR QR K+G DDFE LT+IGKGAFGEVR+CR ++T +VYAMKKL+K+EML RGQVE+V+
Sbjct: 89 YMRLQRRKIGIDDFELLTVIGKGAFGEVRLCRLRSTSEVYAMKKLKKTEMLSRGQVEHVR 148
Query: 162 SERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGE 221
SERNLLAEVDS IVKL+ SFQD E LYLIMEYLPGGD+MTLLMR ++L+E+ A+FYI E
Sbjct: 149 SERNLLAEVDSRYIVKLFYSFQDSECLYLIMEYLPGGDIMTLLMREDILSEDVARFYIAE 208
Query: 222 TVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLD---CSNLQEKD---LSD 275
++LAI SIH+HNY+HRDIKPDNL+LD++GH+KLSDFGLCKPLD S L E D D
Sbjct: 209 SILAIHSIHQHNYVHRDIKPDNLILDKSGHLKLSDFGLCKPLDDKYSSLLLEDDEMLSQD 268
Query: 276 GVNRSGVLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVE 335
N+SG +D+ +P Q ++QL W++N R LAYSTVGT DY+APEV+LK+GYG+E
Sbjct: 269 SENQSGKSDADK---APWQMPKEQLLQWKRNR-RALAYSTVGTLDYMAPEVLLKKGYGME 324
Query: 336 CDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLLC 394
CDWWSLGAI+YEMLVGYPPF SDDPR TCRKI +W+ LKFP++ K+S EA+DLICRLLC
Sbjct: 325 CDWWSLGAILYEMLVGYPPFCSDDPRITCRKIINWRVCLKFPEEPKISDEARDLICRLLC 384
Query: 395 NVKQRLGAKGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEEDQKTEP 454
+V RLG +G +EIK+HPWFK W+KLY M A + P V+ ELDT+NF+KF E +
Sbjct: 385 DVDSRLGTRGVEEIKSHPWFKGTPWDKLYDMEAAYRPIVDGELDTQNFEKFPEVEGSPSE 444
Query: 455 SPKAGPWRKMLQSKDINFVGYTYKNYEMVDANAIPGFGM 493
+P+ GPWRKML SKD NF+G+T+K ++ + G M
Sbjct: 445 APQVGPWRKMLTSKDTNFIGFTFKKSDITRSMESSGADM 483
>At2g19400 putative protein kinase
Length = 509
Score = 527 bits (1358), Expect = e-150
Identities = 266/456 (58%), Positives = 342/456 (74%), Gaps = 29/456 (6%)
Query: 39 SNETKLKVEAAKHFVESYYKNQKQNQQERKERRNILENKLADAEVSKEDKKNLLKNFEEM 98
SN T KV AAK ++E++Y + ++ Q+RKERR +LE K+A +VS++++ LL++ +
Sbjct: 29 SNSTLEKVAAAKKYIENHYNRRMRHIQQRKERRWVLEQKIASLDVSEKEQLELLEDLQRK 88
Query: 99 ETEIMRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVE 158
ETE R R ++ DDF+ L++IG+GAFGEVR+CREK TG +YAMKKL+KSEML RGQVE
Sbjct: 89 ETEYTRLMRNRLCVDDFDLLSIIGRGAFGEVRLCREKKTGNIYAMKKLKKSEMLSRGQVE 148
Query: 159 YVKSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFY 218
+V++ERNLLAEV S+CIVKLY SFQD EYLYLIMEYL GGD+MTLLMR E LTE A+FY
Sbjct: 149 HVRAERNLLAEVASDCIVKLYYSFQDPEYLYLIMEYLSGGDVMTLLMREETLTETVARFY 208
Query: 219 IGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVN 278
I ++VLAIESIHKHNY+HRDIKPDNLLLD+ GH+KLSDFGLCKPLDC N+ ++++ +N
Sbjct: 209 IAQSVLAIESIHKHNYVHRDIKPDNLLLDKYGHMKLSDFGLCKPLDCRNISAMNVNEPLN 268
Query: 279 RSGVLQS----DELTLSPN----QSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKR 330
+ +S + ++ +S +QL+HW+ N R LAYSTVGTPDYIAPEV+LK+
Sbjct: 269 DENINESIDGDENCSIGRRGRRWKSPLEQLQHWQINR-RKLAYSTVGTPDYIAPEVLLKK 327
Query: 331 GYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKDKLSPEAKDLIC 390
GYGVECDWWSLGAIMYEMLVGYPPF+SDDP +L+PEA+DLIC
Sbjct: 328 GYGVECDWWSLGAIMYEMLVGYPPFYSDDPGA-----------------RLTPEARDLIC 370
Query: 391 RLLCNVKQRLGA--KGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEE 448
RLLC+ + RLG+ GA++IKAH WFK VEWEKLY+M A F P VN ELDT+NF KF+E
Sbjct: 371 RLLCDSEHRLGSHGAGAEQIKAHTWFKDVEWEKLYEMDAAFKPVVNGELDTQNFMKFDEV 430
Query: 449 DQKTEPSPKAGP-WRKMLQSKDINFVGYTYKNYEMV 483
+ +GP W+ + ++INFVGYTY+N++ V
Sbjct: 431 ECPKPARTGSGPSWKVSITPQNINFVGYTYRNFDAV 466
>At1g48490 IRE - like protein kinase
Length = 1235
Score = 262 bits (670), Expect = 3e-70
Identities = 149/378 (39%), Positives = 219/378 (57%), Gaps = 25/378 (6%)
Query: 73 ILENKLADAEVSKEDKKNLLKNFEEMETEIMRRQRL-------KMGADDFEPLTMIGKGA 125
I E L E+ ++K ++ +E +++R R ++ DDFE + I +GA
Sbjct: 779 IQEKYLQLCELMDDEKGTIIDEDAPLEDDVVRSLRTSPVHLRDRISIDDFEVMKSISRGA 838
Query: 126 FGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDSNCIVKLYCSFQDD 185
FG V + R+ TTG ++A+K L+K++M+R+ VE + +ER++L + +V+ + SF
Sbjct: 839 FGHVILARKNTTGDLFAIKVLRKADMIRKNAVESILAERDILINARNPFVVRFFYSFTCS 898
Query: 186 EYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHKHNYIHRDIKPDNLL 245
E LYL+MEYL GGD ++L + L E A+ YI E VLA+E +H +HRD+KPDNLL
Sbjct: 899 ENLYLVMEYLNGGDFYSMLRKIGCLDEANARVYIAEVVLALEYLHSEGVVHRDLKPDNLL 958
Query: 246 LDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRK 305
+ +GHVKL+DFGL K +N DLS V+ + L +E P L+H R
Sbjct: 959 IAHDGHVKLTDFGLSKVGLINNTD--DLSGPVSSATSLLVEEKPKLPT------LDHKR- 1009
Query: 306 NHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCR 365
S VGTPDY+APE++L G+G DWWS+G I+YE LVG PPF++D P+
Sbjct: 1010 --------SAVGTPDYLAPEILLGTGHGATADWWSVGIILYEFLVGIPPFNADHPQQIFD 1061
Query: 366 KIAHWKTYLKFPKDKLSPEAKDLICRLLC-NVKQRLGAKGADEIKAHPWFKVVEWEKLYQ 424
I + + +S EA+DLI RLL + QRLGA+GA E+K H +FK ++W L Q
Sbjct: 1062 NILNRNIQWPPVPEDMSHEARDLIDRLLTEDPHQRLGARGAAEVKQHSFFKDIDWNTLAQ 1121
Query: 425 MRAPFIPKVNDELDTRNF 442
+A F+P + DT F
Sbjct: 1122 QKAAFVPDSENAFDTSYF 1139
>At3g17850 protein kinase like protein
Length = 1296
Score = 255 bits (652), Expect = 4e-68
Identities = 141/382 (36%), Positives = 227/382 (58%), Gaps = 23/382 (6%)
Query: 73 ILENKLADAEVSKEDKKNLLKNFEE----METEIMRRQRL-------KMGADDFEPLTMI 121
I E + E+ ++K +LL + +E +++R R + DDFE + I
Sbjct: 829 IREKYVHMCELMDDEKVDLLSTVIDEDAPLEDDVVRSLRTSPVHPRDRTSIDDFEIIKPI 888
Query: 122 GKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDSNCIVKLYCS 181
+GAFG V + +++TTG ++A+K L+K++M+R+ VE + +ER++L V + +V+ + S
Sbjct: 889 SRGAFGRVFLAKKRTTGDLFAIKVLKKADMIRKNAVESILAERDILINVRNPFVVRFFYS 948
Query: 182 FQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHKHNYIHRDIKP 241
F + LYL+MEYL GGD+ +LL L E+ + YI E VLA+E +H +HRD+KP
Sbjct: 949 FTCRDNLYLVMEYLNGGDLYSLLRNLGCLEEDIVRVYIAEVVLALEYLHSEGVVHRDLKP 1008
Query: 242 DNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDELTLSPNQSQQQQLE 301
DNLL+ +GH+KL+DFGL K + N + V+ + +L +E L+ + ++QLE
Sbjct: 1009 DNLLIAHDGHIKLTDFGLSK-VGLINSTDDLAGPAVSGTSLLDEEESRLA---ASEEQLE 1064
Query: 302 HWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPR 361
+K S VGTPDY+APE++L G+G DWWS+G I++E++VG PPF+++ P+
Sbjct: 1065 RRKKR-------SAVGTPDYLAPEILLGTGHGATADWWSVGIILFELIVGIPPFNAEHPQ 1117
Query: 362 TTCRKIAHWKTYLKFPKDKLSPEAKDLICRLLC-NVKQRLGAKGADEIKAHPWFKVVEWE 420
I + K +++S EA D+I R L + QRLGA+GA E+K H +FK + W+
Sbjct: 1118 QIFDNILNRKIPWPHVPEEMSAEAHDIIDRFLTEDPHQRLGARGAAEVKQHIFFKDINWD 1177
Query: 421 KLYQMRAPFIPKVNDELDTRNF 442
L + +A F+P +DT F
Sbjct: 1178 TLARQKAAFVPASESAIDTSYF 1199
>At5g62310 IRE (root hair elongation)
Length = 1168
Score = 254 bits (648), Expect = 1e-67
Identities = 155/438 (35%), Positives = 254/438 (57%), Gaps = 30/438 (6%)
Query: 18 ATSTSNDHQC--LALDLVNEEPPSNETKLKVEAAKH---FVESYYKN-QKQNQQERKERR 71
A S +N + C +LD + E+ +E K ++ K VE++ + +K Q++ E
Sbjct: 650 ARSVANVNVCGYSSLDFMIEQ--LDELKYVIQDRKADALVVETFGRRIEKLLQEKYIELC 707
Query: 72 NILENKLADAEVSKEDKKNLLKNFEEMETEIMR------RQRLKMGADDFEPLTMIGKGA 125
+++++ D+ + D+++ + +E +R R + + +DFE + I +GA
Sbjct: 708 GLIDDEKVDSSNAMPDEES---SADEDTVRSLRASPLNPRAKDRTSIEDFEIIKPISRGA 764
Query: 126 FGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDSNCIVKLYCSFQDD 185
FG V + +++ TG ++A+K L+K++M+R+ VE + +ERN+L V + +V+ + SF
Sbjct: 765 FGRVFLAKKRATGDLFAIKVLKKADMIRKNAVESILAERNILISVRNPFVVRFFYSFTCR 824
Query: 186 EYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHKHNYIHRDIKPDNLL 245
E LYL+MEYL GGD+ +LL L E+ A+ YI E VLA+E +H N IHRD+KPDNLL
Sbjct: 825 ENLYLVMEYLNGGDLFSLLRNLGCLDEDMARIYIAEVVLALEYLHSVNIIHRDLKPDNLL 884
Query: 246 LDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDELTLSPNQSQQQQLEHWRK 305
++++GH+KL+DFGL K ++ + + SG D +++Q Q + RK
Sbjct: 885 INQDGHIKLTDFGLSKVGLINSTDDLSGESSLGNSGFFAED-----GSKAQHSQGKDSRK 939
Query: 306 NHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCR 365
H + VGTPDY+APE++L G+G DWWS+G I++E+LVG PPF+++ P+
Sbjct: 940 KH------AVVGTPDYLAPEILLGMGHGKTADWWSVGVILFEVLVGIPPFNAETPQQIFE 993
Query: 366 KIAHWKTYLKFPKDKLSPEAKDLICRLLC-NVKQRLGAKGADEIKAHPWFKVVEWEKLYQ 424
I + +++S EA DLI +LL N QRLGA GA E+K H +FK + W+ L +
Sbjct: 994 NIINRDIPWPNVPEEISYEAHDLINKLLTENPVQRLGATGAGEVKQHHFFKDINWDTLAR 1053
Query: 425 MRAPFIPKVNDELDTRNF 442
+A F+P + DT F
Sbjct: 1054 QKAMFVPSAEPQ-DTSYF 1070
>At1g45160 similar to IRE homolog 1 dbj|BAA89784.1
Length = 1067
Score = 248 bits (633), Expect = 6e-66
Identities = 140/355 (39%), Positives = 207/355 (57%), Gaps = 20/355 (5%)
Query: 109 KMGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLA 168
++ DDFE + I +GAFG+V + R++TTG +A+K L+K +M+R+ +E + ERN+L
Sbjct: 664 RISIDDFEIIKPISRGAFGKVFLARKRTTGDFFAIKVLKKLDMIRKNDIERILQERNILI 723
Query: 169 EVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIES 228
V +V+ + SF + LYL+MEYL GGD+ +LL + L E A+ YI E VLA+E
Sbjct: 724 TVRYPFLVRFFYSFTCRDNLYLVMEYLNGGDLYSLLQKVGCLDEEIARIYIAELVLALEY 783
Query: 229 IHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDEL 288
+H +HRD+KPDNLL+ NGH+KL+DFGL K +N + L E
Sbjct: 784 LHSLKIVHRDLKPDNLLIAYNGHIKLTDFGLSK------------IGLINNTIDLSGHES 831
Query: 289 TLSPNQSQQQQLEHWRKN-HSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYE 347
+SP + H++KN + +S VGTPDY+APE++L +G DWWS G +++E
Sbjct: 832 DVSPRTNS----HHFQKNQEEERIRHSAVGTPDYLAPEILLGTEHGYAADWWSAGIVLFE 887
Query: 348 MLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKDKLSPEAKDLICRLLCN-VKQRLGAKGAD 406
+L G PPF + P I + K ++S EA+DLI RLL + ++RLGA GA
Sbjct: 888 LLTGIPPFTASRPEKIFDNILNGKMPWPDVPGEMSYEAQDLINRLLVHEPEKRLGANGAA 947
Query: 407 EIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNF-DKFEEED-QKTEPSPKAG 459
E+K+HP+F+ V+WE L +A F+P+ DT F +F E TE +G
Sbjct: 948 EVKSHPFFQGVDWENLALQKAAFVPQPESINDTSYFVSRFSESSCSDTETGNNSG 1002
>At3g45780 nonphototropic hypocotyl 1 (NPH1)
Length = 996
Score = 184 bits (466), Expect = 1e-46
Identities = 125/353 (35%), Positives = 182/353 (51%), Gaps = 39/353 (11%)
Query: 110 MGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAE 169
+G F+P+ +G G G V + T Q++AMK + K+ ML R +V ++ER +L
Sbjct: 658 IGLKHFKPVKPLGSGDTGSVHLVELVGTDQLFAMKAMDKAVMLNRNKVHRARAEREILDL 717
Query: 170 VDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMR--REVLTENEAKFYIGETVLAIE 227
+D + LY SFQ ++ LI +Y PGG++ LL R R+VL E+ +FY + V+A+E
Sbjct: 718 LDHPFLPALYASFQTKTHICLITDYYPGGELFMLLDRQPRKVLKEDAVRFYAAQVVVALE 777
Query: 228 SIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGL-----CKP-LDCSNLQEKDLSDGVNRSG 281
+H I+RD+KP+N+L+ NG + LSDF L CKP L ++ EK
Sbjct: 778 YLHCQGIIYRDLKPENVLIQGNGDISLSDFDLSCLTSCKPQLLIPSIDEKK--------- 828
Query: 282 VLQSDELTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSL 341
Q + QQ + R + S VGT +YIAPE+I G+ DWW+L
Sbjct: 829 ---------KKKQQKSQQTPIFMAEPMR-ASNSFVGTEEYIAPEIISGAGHTSAVDWWAL 878
Query: 342 GAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLL-CNVKQR 399
G +MYEML GY PF + T + + LKFP S + K LI RLL + K+R
Sbjct: 879 GILMYEMLYGYTPFRGKTRQKTFTNVL--QKDLKFPASIPASLQVKQLIFRLLQRDPKKR 936
Query: 400 LGA-KGADEIKAHPWFKVVEWEKLYQMRAPFIPKVNDELDTRNFDKFEEEDQK 451
LG +GA+E+K H +FK + W + P EL+T F E +K
Sbjct: 937 LGCFEGANEVKQHSFFKGINWALIRCTNPP-------ELETPIFSGEAENGEK 982
>At5g58140 non phototropic hypocotyl 1-like (NPL1)
Length = 915
Score = 177 bits (449), Expect = 1e-44
Identities = 111/324 (34%), Positives = 173/324 (53%), Gaps = 22/324 (6%)
Query: 110 MGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAE 169
+G F+P+ +G G G V + K TG++YAMK ++K+ ML R + ER +++
Sbjct: 572 VGLHHFKPIKPLGSGDTGSVHLVELKGTGELYAMKAMEKTMMLNRNKAHRACIEREIISL 631
Query: 170 VDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRR--EVLTENEAKFYIGETVLAIE 227
+D + LY SFQ ++ LI ++ PGG++ LL R+ ++LTE+ A+FY E V+ +E
Sbjct: 632 LDHPFLPTLYASFQTSTHVCLITDFCPGGELFALLDRQPMKILTEDSARFYAAEVVIGLE 691
Query: 228 SIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDE 287
+H ++RD+KP+N+LL ++GH+ L+DF L C+ + R
Sbjct: 692 YLHCLGIVYRDLKPENILLKKDGHIVLADFDLSFMTTCTPQLIIPAAPSKRR-------- 743
Query: 288 LTLSPNQSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYE 347
+S+ Q L + S + S VGT +YIAPE+I G+ DWW+LG ++YE
Sbjct: 744 ------RSKSQPLPTFVAEPSTQ-SNSFVGTEEYIAPEIITGAGHTSAIDWWALGILLYE 796
Query: 348 MLVGYPPFHSDDPRTTCRKIAHWKTYLKFPKD-KLSPEAKDLICRLL-CNVKQRLGAK-G 404
ML G PF + + T I H L FP +S + LI LL + RLG+K G
Sbjct: 797 MLYGRTPFRGKNRQKTFANILH--KDLTFPSSIPVSLVGRQLINTLLNRDPSSRLGSKGG 854
Query: 405 ADEIKAHPWFKVVEWEKLYQMRAP 428
A+EIK H +F+ + W + M P
Sbjct: 855 ANEIKQHAFFRGINWPLIRGMSPP 878
>At3g10540 3-phosphoinositide-dependent protein kinase-1 like
Length = 486
Score = 175 bits (444), Expect = 5e-44
Identities = 110/318 (34%), Positives = 166/318 (51%), Gaps = 38/318 (11%)
Query: 114 DFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDSN 173
DFE + G G++ +V ++K G VYA+K + K + + + YVK ER +L +++
Sbjct: 44 DFELGKIYGVGSYSKVVRAKKKDNGTVYALKIMDKKFITKENKTAYVKLERIVLDQLEHP 103
Query: 174 CIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHKHN 233
IVKL+ +FQD + LY+ +E GG++ + R+ L+E+EA+FY E V A+E IH
Sbjct: 104 GIVKLFFTFQDTQSLYMALESCEGGELFDQITRKGRLSEDEARFYSAEVVDALEYIHNMG 163
Query: 234 YIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDELTLSPN 293
IHRDIKP+NLLL +GH+K++DFG KP +Q ++T+ PN
Sbjct: 164 LIHRDIKPENLLLTLDGHIKIADFGSVKP--------------------MQDSQITVLPN 203
Query: 294 QSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYP 353
+ + A + VGT Y+ PEV+ D W+LG +Y+ML G
Sbjct: 204 AASDDK------------ACTFVGTAAYVPPEVLNSSPATFGNDLWALGCTLYQMLSGTS 251
Query: 354 PFHSDDPRTTCRKIAHWKTYLKFPKDKLSPEAKDLICRLLCNVKQR---LGAKGADEIKA 410
PF ++I +KFP + S A+DLI RLL R G++G D +K
Sbjct: 252 PFKDASEWLIFQRII--ARDIKFP-NHFSEAARDLIDRLLDTDPSRRPGAGSEGYDSLKR 308
Query: 411 HPWFKVVEWEKLYQMRAP 428
HP+FK V+W+ L P
Sbjct: 309 HPFFKGVDWKNLRSQTPP 326
>At5g04510 3-phosphoinositide-dependent protein kinase-1 (PDK1)
Length = 491
Score = 172 bits (437), Expect = 3e-43
Identities = 109/318 (34%), Positives = 164/318 (51%), Gaps = 38/318 (11%)
Query: 114 DFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDSN 173
DFE + G G++ +V ++K TG VYA+K + K + + + YVK ER +L +++
Sbjct: 43 DFEFGKIYGVGSYSKVVRAKKKETGTVYALKIMDKKFITKENKTAYVKLERIVLDQLEHP 102
Query: 174 CIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRREVLTENEAKFYIGETVLAIESIHKHN 233
I+KLY +FQD LY+ +E GG++ + R+ L+E+EA+FY E V A+E IH
Sbjct: 103 GIIKLYFTFQDTSSLYMALESCEGGELFDQITRKGRLSEDEARFYTAEVVDALEYIHSMG 162
Query: 234 YIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDELTLSPN 293
IHRDIKP+NLLL +GH+K++DFG KP +Q ++T+ PN
Sbjct: 163 LIHRDIKPENLLLTSDGHIKIADFGSVKP--------------------MQDSQITVLPN 202
Query: 294 QSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYP 353
+ + A + VGT Y+ PEV+ D W+LG +Y+ML G
Sbjct: 203 AASDDK------------ACTFVGTAAYVPPEVLNSSPATFGNDLWALGCTLYQMLSGTS 250
Query: 354 PFHSDDPRTTCRKIAHWKTYLKFPKDKLSPEAKDLICRLLCNVKQR---LGAKGADEIKA 410
PF ++I +KFP + S A+DLI RLL R G++G +K
Sbjct: 251 PFKDASEWLIFQRII--ARDIKFP-NHFSEAARDLIDRLLDTEPSRRPGAGSEGYVALKR 307
Query: 411 HPWFKVVEWEKLYQMRAP 428
HP+F V+W+ L P
Sbjct: 308 HPFFNGVDWKNLRSQTPP 325
>At2g44830 protein kinase like protein
Length = 765
Score = 171 bits (433), Expect = 9e-43
Identities = 121/389 (31%), Positives = 185/389 (47%), Gaps = 57/389 (14%)
Query: 110 MGADDFEPLTMIGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAE 169
+G F+ L +G G G V + T +A+K + K+ + R ++ ++ER++L
Sbjct: 358 LGMSHFKLLKRLGCGDIGSVYLAELSGTRCHFAVKVMDKASLEDRKKLNRAQTERDILQL 417
Query: 170 VDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRR--EVLTENEAKFYIGETVLAIE 227
+D + LY F+ D + L+MEY PGGD+ TL R+ + +E A+FY E +LA+E
Sbjct: 418 LDHPFLPTLYTHFETDRFSCLVMEYCPGGDLHTLRQRQPGKHFSEYAARFYAAEVLLALE 477
Query: 228 SIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVNRSGVLQSDE 287
+H ++RD+KP+N+L+ +GH+ LSDF L S K +R G
Sbjct: 478 YLHMLGVVYRDLKPENVLVRDDGHIMLSDFDLSLRCAVSPTLIKTFDSDPSRRGAFCVQP 537
Query: 288 LTLSP-----------------------NQSQQQQLEHWRKNHSRML----------AYS 314
+ P N+S++ Q + + K+HS L + S
Sbjct: 538 ACMEPTSACIIQPSCFLPRSIFPNKNKKNKSRKTQADFF-KSHSGSLPELVAEPNTRSMS 596
Query: 315 TVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYL 374
VGT +Y+APE+I G+G DWW+ G ++E+L G PF R T + L
Sbjct: 597 FVGTHEYLAPEIIKGEGHGSAVDWWTFGIFVHELLYGKTPFKGSGNRATLFNVV--GEQL 654
Query: 375 KFPKDKLSPEA-KDLICRLLC-NVKQRLGAK-GADEIKAHPWFKVVEWEKLYQMRAPFIP 431
KFP+ + A +DLI LL + K RLG K GA EIK HP+F+ V W + P +P
Sbjct: 655 KFPESPATSYAGRDLIQALLVKDPKNRLGTKRGATEIKQHPFFEGVNWALIRCSTPPEVP 714
Query: 432 KVNDELDTRNFDKFEEEDQKTEPSPKAGP 460
+ +TEP PK GP
Sbjct: 715 R----------------QMETEPPPKYGP 727
>At1g16440 putative protein kinase
Length = 431
Score = 168 bits (426), Expect = 6e-42
Identities = 106/360 (29%), Positives = 184/360 (50%), Gaps = 30/360 (8%)
Query: 103 MRRQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQV--YAMKKLQKSEMLRRGQVEYV 160
++ + +K+G DF L +G G G V + K +AMK + K+ ++ R ++
Sbjct: 33 LKSRGIKLGISDFRVLKRLGYGDIGSVYLVELKGANPTTYFAMKVMDKASLVSRNKLLRA 92
Query: 161 KSERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRR--EVLTENEAKFY 218
++ER +L+++D + LY F+ D++ L+ME+ GG++ +L ++ + TE+ A+F+
Sbjct: 93 QTEREILSQLDHPFLPTLYSHFETDKFYCLVMEFCSGGNLYSLRQKQPNKCFTEDAARFF 152
Query: 219 IGETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSNLQEKDLSDGVN 278
E +LA+E +H ++RD+KP+N+L+ +GH+ LSDF L + K + G
Sbjct: 153 ASEVLLALEYLHMLGIVYRDLKPENVLVRDDGHIMLSDFDLSLRCSVNPTLVKSFNGG-G 211
Query: 279 RSGVLQSDELT---LSPNQSQQQQLEHWRKNH-----------------SRMLAYSTVGT 318
+G++ + P+ + L+ +KN + + + S VGT
Sbjct: 212 TTGIIDDNAAVQGCYQPSAFFPRMLQSSKKNRKSKSDFDGSLPELMAEPTNVKSMSFVGT 271
Query: 319 PDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKIAHWKTYLKFPK 378
+Y+APE+I G+G DWW+ G +YE+L G PF + T + L+FP+
Sbjct: 272 HEYLAPEIIKNEGHGSAVDWWTFGIFIYELLHGATPFKGQGNKATLYNVIGQP--LRFPE 329
Query: 379 -DKLSPEAKDLICRLLCNVKQRLGA--KGADEIKAHPWFKVVEWEKLYQMRAPFIPKVND 435
++S AKDLI LL Q A +GA EIK HP+F+ V W + P +P+ D
Sbjct: 330 YSQVSSTAKDLIKGLLVKEPQNRIAYKRGATEIKQHPFFEGVNWALIRGETPPHLPEPVD 389
>At3g14370 putative protein kinase
Length = 480
Score = 157 bits (398), Expect = 1e-38
Identities = 92/317 (29%), Positives = 159/317 (50%), Gaps = 9/317 (2%)
Query: 121 IGKGAFGEVRICREKTTGQVYAMKKLQKSEMLRRGQVEYVKSERNLLAEVDSNCIVKLYC 180
+G G G V +C + + +A+K + ++ + ++ V++E +L+ +D + LY
Sbjct: 94 LGTGNLGRVFLCNLRDSSARFALKVIDRNCLTTEKKLSQVETEAEILSLLDHPFLPTLYA 153
Query: 181 SFQDDEYLYLIMEYLPGGDMMTLLMRR--EVLTENEAKFYIGETVLAIESIHKHNYIHRD 238
+ Y L+++Y P GD+ +LL ++ L +F+ E ++A+E +H ++RD
Sbjct: 154 RIDESHYTCLLIDYAPNGDLHSLLRKQPGNRLPIQPVRFFAAEVLVALEYLHAMGIVYRD 213
Query: 239 IKPDNLLLDRNGHVKLSDFGLCKPLDC-----SNLQEKDLSDGVNRSGVLQSDELTLSPN 293
+KP+N+LL +GHV LSDF LC D S + S R +
Sbjct: 214 LKPENVLLREDGHVMLSDFDLCFKSDVVPTFKSRRYRRSSSSPSLRRRRSGCFSVAAEKK 273
Query: 294 QSQQQQLEHWRKNHSRMLAYSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYP 353
+++ + + + S VGT +Y+APE++ G+G DWW+ G +YE+L G
Sbjct: 274 YEREEIVSEFAAEPVTAFSRSCVGTHEYLAPELVSGNGHGSGVDWWAFGIFLYELLYGTT 333
Query: 354 PFHSDDPRTTCRKIAHWKTYLKFPKDKLSPEAKDLICRLLC-NVKQRLG-AKGADEIKAH 411
PF + T R I F D EA+DLI +LL + ++RLG A+GA +IK H
Sbjct: 334 PFKGESKEQTLRNIVSTTKTASFHMDGDLDEARDLIEKLLVKDPRKRLGCARGAQDIKRH 393
Query: 412 PWFKVVEWEKLYQMRAP 428
P+F ++W + + P
Sbjct: 394 PFFDGIKWPLIRHYKPP 410
>At3g44610 protein kinase-like protein
Length = 451
Score = 157 bits (397), Expect = 1e-38
Identities = 113/367 (30%), Positives = 183/367 (49%), Gaps = 46/367 (12%)
Query: 105 RQRLKMGADDFEPLTMIGKGAFGEVRICREKTTGQV--YAMKKLQKSEMLRRGQVEYVKS 162
R RL++G+ D + + F + E T ++ A K + K E+ R + K+
Sbjct: 70 RFRLRLGSGDIGSVFL---AEFKSLTAVTETTAVKLPLLAAKVMDKKELASRSKEGRAKT 126
Query: 163 ERNLLAEVDSNCIVKLYCSFQDDEYLYLIMEYLPGGDMMTLLMRR--EVLTENEAKFYIG 220
ER +L +D + LY + ++L L+ E+ PGGD+ L ++ + E+ +FY+
Sbjct: 127 EREILESLDHPFLPTLYAAIDSPKWLCLLTEFCPGGDLHVLRQKQTHKRFHESAVRFYVS 186
Query: 221 ETVLAIESIHKHNYIHRDIKPDNLLLDRNGHVKLSDFGLCKPLDCSN------LQEKDLS 274
E ++AIE +H ++RD+KP+N+L+ +GH+ L+DF L D S L +L
Sbjct: 187 EVIVAIEYLHMLGIVYRDLKPENVLVRSDGHIMLTDFDLSLKCDESTSTPQIVLNRNNLP 246
Query: 275 DGV---NRSGVLQSDELTLS----PN-------------QSQQQQLEHWRKNHSRMLA-- 312
+G N + + + T S PN + ++++ +H R N ++A
Sbjct: 247 NGSSDQNENQGMDHRQTTSSSCMIPNCIVPAVSCFHPRIRRRKKKTDH-RNNGPELVAEP 305
Query: 313 -----YSTVGTPDYIAPEVILKRGYGVECDWWSLGAIMYEMLVGYPPFHSDDPRTTCRKI 367
S VGT +Y+APE++ G+G DWW+LG M+E+ G PF D T I
Sbjct: 306 VDVRSMSFVGTHEYLAPEIVSGEGHGSAVDWWTLGIFMFELFYGTTPFKGMDHELTLANI 365
Query: 368 AHWKTYLKFPKDKLSPE-AKDLICRLLC-NVKQRLGAK-GADEIKAHPWFKVVEWEKLYQ 424
L+FPK+ P AKDLI +LL + +RLG+ GA +K HP+F+ V W L
Sbjct: 366 V--ARALEFPKEPTIPSAAKDLISQLLAKDPSRRLGSSLGATAVKRHPFFQGVNWALLMC 423
Query: 425 MRAPFIP 431
R PF+P
Sbjct: 424 TRPPFLP 430
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.135 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,774,594
Number of Sequences: 26719
Number of extensions: 575064
Number of successful extensions: 4566
Number of sequences better than 10.0: 1013
Number of HSP's better than 10.0 without gapping: 885
Number of HSP's successfully gapped in prelim test: 128
Number of HSP's that attempted gapping in prelim test: 1989
Number of HSP's gapped (non-prelim): 1851
length of query: 542
length of database: 11,318,596
effective HSP length: 104
effective length of query: 438
effective length of database: 8,539,820
effective search space: 3740441160
effective search space used: 3740441160
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC142507.1


