

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.8 - phase: 0 /pseudo
(82 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
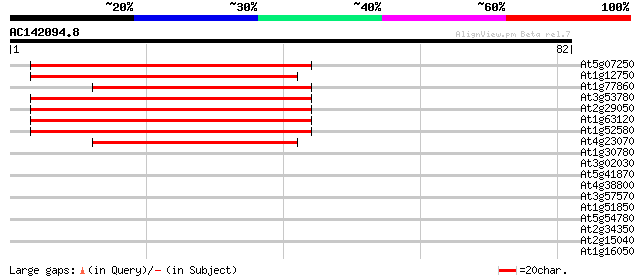
Score E
Sequences producing significant alignments: (bits) Value
At5g07250 membrane protein 47 2e-06
At1g12750 unknown protein 45 8e-06
At1g77860 hypothetical protein 44 1e-05
At3g53780 unknown protein 43 3e-05
At2g29050 hypothetical protein 42 4e-05
At1g63120 membrane protein, putative 42 7e-05
At1g52580 unknown protein 40 2e-04
At4g23070 putative membrane protein 39 5e-04
At1g30780 hypothetical protein 28 1.1
At3g02030 hypothetical protein 27 1.4
At5g41870 polygalacturonase-like protein 27 2.4
At4g38800 unknown protein 27 2.4
At3g57570 unknown protein 25 5.3
At1g51850 Putative protein kinase 25 5.3
At5g54780 putative protein 25 9.0
At2g34350 nodulin-like protein 25 9.0
At2g15040 putative disease resistance protein 25 9.0
At1g16050 hypothetical protein 25 9.0
>At5g07250 membrane protein
Length = 346
Score = 47.0 bits (110), Expect = 2e-06
Identities = 21/41 (51%), Positives = 30/41 (72%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
L++ V + NL IGI+P V+NF +GG + GFLLGF+LL +
Sbjct: 230 LTLLFVILINLAIGILPHVDNFAHVGGFVTGFLLGFILLAR 270
>At1g12750 unknown protein
Length = 307
Score = 44.7 bits (104), Expect = 8e-06
Identities = 21/39 (53%), Positives = 27/39 (68%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
LS + NL IG++P V+NF IGGL+ GF LGF+LL
Sbjct: 193 LSFLFIIAINLAIGLLPWVDNFAHIGGLLTGFCLGFILL 231
>At1g77860 hypothetical protein
Length = 319
Score = 43.9 bits (102), Expect = 1e-05
Identities = 20/32 (62%), Positives = 23/32 (71%)
Query: 13 NLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
N IG +P ++NF IGG I GFLLGFVLL K
Sbjct: 227 NFLIGFLPFIDNFANIGGFISGFLLGFVLLFK 258
>At3g53780 unknown protein
Length = 394
Score = 42.7 bits (99), Expect = 3e-05
Identities = 19/41 (46%), Positives = 29/41 (70%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ ++ NL +G++P V+NF IGG GFLLGFVLL +
Sbjct: 235 VTLVLIVAVNLGLGVLPGVDNFAHIGGFATGFLLGFVLLIR 275
>At2g29050 hypothetical protein
Length = 372
Score = 42.4 bits (98), Expect = 4e-05
Identities = 19/41 (46%), Positives = 27/41 (65%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
L++ + NL +GI+P V+NF +GG GFLLGFV L +
Sbjct: 279 LTLIFIIAINLAVGILPHVDNFAHLGGFTSGFLLGFVFLIR 319
>At1g63120 membrane protein, putative
Length = 317
Score = 41.6 bits (96), Expect = 7e-05
Identities = 18/41 (43%), Positives = 28/41 (67%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ + NL +G++P V+NF IGG + GF LGFVLL +
Sbjct: 206 ITLLFIIAINLALGMLPRVDNFAHIGGFLTGFCLGFVLLVR 246
>At1g52580 unknown protein
Length = 309
Score = 40.0 bits (92), Expect = 2e-04
Identities = 17/41 (41%), Positives = 28/41 (67%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCK 44
+++ ++ V NL++G +P V+N GG + GF LGFVLL +
Sbjct: 204 MTLILIIVLNLSVGFLPRVDNSAHFGGFLAGFFLGFVLLLR 244
>At4g23070 putative membrane protein
Length = 313
Score = 38.9 bits (89), Expect = 5e-04
Identities = 18/30 (60%), Positives = 22/30 (73%)
Query: 13 NLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
NL +G +P V+NF IGG GFLLGF+LL
Sbjct: 209 NLGLGTLPPVDNFAHIGGFFGGFLLGFLLL 238
>At1g30780 hypothetical protein
Length = 481
Score = 27.7 bits (60), Expect = 1.1
Identities = 12/48 (25%), Positives = 25/48 (52%)
Query: 21 IVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFILL 68
+ +N+ + ++ G +LG VL+C + F + L R + + F +L
Sbjct: 65 VSSNYSFVFNMVTGLILGQVLICYVNKFTFGVKSLIWRYVVFLSFFIL 112
>At3g02030 hypothetical protein
Length = 656
Score = 27.3 bits (59), Expect = 1.4
Identities = 19/55 (34%), Positives = 25/55 (44%)
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKR 58
L + M PVF T +V G I P L+G +L D LP Q +H+R
Sbjct: 371 LEVIMGPVFLSTTEDGKVVRGLGGIPSEGPVLLVGNHMLLASDKISLPGQFVHER 425
>At5g41870 polygalacturonase-like protein
Length = 449
Score = 26.6 bits (57), Expect = 2.4
Identities = 24/78 (30%), Positives = 34/78 (42%), Gaps = 10/78 (12%)
Query: 3 NLSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKK--------DPFVLPDQK 54
N+++ + ++N IGI I N G GG I G + V L K D PD K
Sbjct: 303 NVTVENITLYNSGIGI-HIKTNIGR-GGSIQGITISGVYLEKVRTGIKISGDTGDHPDDK 360
Query: 55 LHKRCLPIICFILLSTGW 72
+ LPI+ I + W
Sbjct: 361 FNTSALPIVRGITIKNVW 378
>At4g38800 unknown protein
Length = 267
Score = 26.6 bits (57), Expect = 2.4
Identities = 22/73 (30%), Positives = 31/73 (42%), Gaps = 13/73 (17%)
Query: 5 SIGMVPVFNLTIGIVP-----IVNNFGLIGGL-IPGFLLGFVLLCKKDPFVLPDQKLHKR 58
S+G VP +T + I+ N G GG + G +G D F++ D H R
Sbjct: 89 SVGTVPASLITFASIQALKPDIIINAGTCGGFKVKGANIG-------DVFLVSDVVFHDR 141
Query: 59 CLPIICFILLSTG 71
+PI F L G
Sbjct: 142 RIPIPMFDLYGVG 154
>At3g57570 unknown protein
Length = 1057
Score = 25.4 bits (54), Expect = 5.3
Identities = 17/46 (36%), Positives = 20/46 (42%)
Query: 27 LIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFILLSTGW 72
L G L P L VLL + + D K CL IC L+ GW
Sbjct: 760 LCGRLHPQVLFPTVLLQLEKATEIQDSLKIKACLFSICTSLMVRGW 805
>At1g51850 Putative protein kinase
Length = 875
Score = 25.4 bits (54), Expect = 5.3
Identities = 14/32 (43%), Positives = 18/32 (55%)
Query: 18 IVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFV 49
IVP+V + I LI +L F+L KK P V
Sbjct: 507 IVPVVASIASIAVLIGALVLFFILRKKKSPKV 538
>At5g54780 putative protein
Length = 435
Score = 24.6 bits (52), Expect = 9.0
Identities = 10/15 (66%), Positives = 12/15 (79%)
Query: 54 KLHKRCLPIICFILL 68
KLHK+ L +CFILL
Sbjct: 421 KLHKKYLKKVCFILL 435
>At2g34350 nodulin-like protein
Length = 2301
Score = 24.6 bits (52), Expect = 9.0
Identities = 12/32 (37%), Positives = 16/32 (49%), Gaps = 1/32 (3%)
Query: 9 VPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFV 40
+P FN T +V NF GG G + GF+
Sbjct: 740 LPFFN-TANVVTAARNFSQYGGTAVGIMQGFL 770
>At2g15040 putative disease resistance protein
Length = 1011
Score = 24.6 bits (52), Expect = 9.0
Identities = 9/25 (36%), Positives = 15/25 (60%)
Query: 21 IVNNFGLIGGLIPGFLLGFVLLCKK 45
I G I G+ G ++G++L+C K
Sbjct: 966 IAATIGFIPGIAFGLMMGYILVCYK 990
>At1g16050 hypothetical protein
Length = 160
Score = 24.6 bits (52), Expect = 9.0
Identities = 11/25 (44%), Positives = 15/25 (60%)
Query: 20 PIVNNFGLIGGLIPGFLLGFVLLCK 44
PI +G IGG I G +LG + C+
Sbjct: 84 PICVCYGAIGGYIGGQMLGLMRTCE 108
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.331 0.154 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,960,291
Number of Sequences: 26719
Number of extensions: 75934
Number of successful extensions: 185
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 169
Number of HSP's gapped (non-prelim): 21
length of query: 82
length of database: 11,318,596
effective HSP length: 58
effective length of query: 24
effective length of database: 9,768,894
effective search space: 234453456
effective search space used: 234453456
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC142094.8


