

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.3 + phase: 0
(85 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
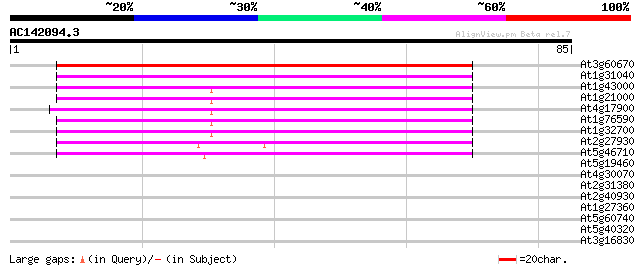
Sequences producing significant alignments: (bits) Value
At3g60670 putative protein 72 6e-14
At1g31040 F17F8.4 65 8e-12
At1g43000 hypothetical protein 58 7e-10
At1g21000 unknown protein 57 1e-09
At4g17900 unknown protein 54 1e-08
At1g76590 unknown protein 54 1e-08
At1g32700 unknown protein 52 5e-08
At2g27930 unknown protein 49 3e-07
At5g46710 unknown protein 47 1e-06
At5g19460 unknown protein 28 1.1
At4g30070 hypothetical protein 27 2.3
At2g31380 B-box zinc finger protein (STH) 27 2.3
At2g40930 ubiquitin-specific protease (UBP5) 26 3.1
At1g27360 squamosa-promoter binding protein-like 11 26 3.1
At5g60740 xABC transporter - like protein 26 4.0
At5g40320 putative protein 26 4.0
At3g16830 unknown protein 25 8.9
>At3g60670 putative protein
Length = 245
Score = 71.6 bits (174), Expect = 6e-14
Identities = 31/63 (49%), Positives = 38/63 (60%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C DH D + NEKN C+DC + C HC +H+ HR QI RY Y+DV R + K DCS
Sbjct: 22 CLDHEDDKKNEKNILCIDCCLTICPHCLSSHTSHRLLQIRRYVYRDVLRVEDGSKLMDCS 81
Query: 68 NIQ 70
IQ
Sbjct: 82 LIQ 84
>At1g31040 F17F8.4
Length = 241
Score = 64.7 bits (156), Expect = 8e-12
Identities = 28/63 (44%), Positives = 36/63 (56%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C H R +EKN FC+ C + C HC +H H Q+ RY Y DV R S+L+K DCS
Sbjct: 34 CGIHETRRKSEKNVFCLLCCLSVCPHCLPSHRSHPLLQVRRYVYHDVVRLSDLEKLIDCS 93
Query: 68 NIQ 70
+Q
Sbjct: 94 YVQ 96
>At1g43000 hypothetical protein
Length = 216
Score = 58.2 bits (139), Expect = 7e-10
Identities = 30/64 (46%), Positives = 35/64 (53%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H +E N FC+DC+ FC C H HR QI R SY +V R SE+QKH D
Sbjct: 24 CSIHSQSSKSECNLFCLDCSGNAFCSSCLAHHRTHRVIQIRRSSYHNVVRVSEIQKHIDI 83
Query: 67 SNIQ 70
S IQ
Sbjct: 84 SCIQ 87
>At1g21000 unknown protein
Length = 246
Score = 57.4 bits (137), Expect = 1e-09
Identities = 29/64 (45%), Positives = 36/64 (55%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D NE N FC+DCA FC +C H HR QI R SY +V R +E+QK D
Sbjct: 31 CSIHVDSNKNECNLFCLDCAGNAFCSYCLVKHKDHRVVQIRRSSYHNVVRVNEIQKFIDI 90
Query: 67 SNIQ 70
+ +Q
Sbjct: 91 ACVQ 94
>At4g17900 unknown protein
Length = 227
Score = 54.3 bits (129), Expect = 1e-08
Identities = 27/65 (41%), Positives = 34/65 (51%), Gaps = 1/65 (1%)
Query: 7 YCDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFD 65
+C H D +E N +C+DC C C H HR QI R SY DV R +E+QK+ D
Sbjct: 48 HCKFHGDSHKSECNMYCLDCTNGPLCSLCLAHHKDHRTIQIRRSSYHDVIRVNEIQKYLD 107
Query: 66 CSNIQ 70
IQ
Sbjct: 108 IGGIQ 112
>At1g76590 unknown protein
Length = 245
Score = 53.9 bits (128), Expect = 1e-08
Identities = 27/64 (42%), Positives = 36/64 (56%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H +E N FC+DC+ FC +C H HR QI R SY +V R +E+QK+ D
Sbjct: 33 CSIHAASNKSECNMFCLDCSSEAFCSYCLLNHRNHRVLQIRRSSYHNVVRVNEIQKYIDI 92
Query: 67 SNIQ 70
S +Q
Sbjct: 93 SCVQ 96
>At1g32700 unknown protein
Length = 213
Score = 52.0 bits (123), Expect = 5e-08
Identities = 26/64 (40%), Positives = 32/64 (49%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H H QI R SY DV R SE+QK D
Sbjct: 27 CKLHADSHKSECNMYCLDCTNGPLCSLCLSFHKDHHAIQIRRSSYHDVIRVSEIQKFLDI 86
Query: 67 SNIQ 70
+ +Q
Sbjct: 87 TGVQ 90
>At2g27930 unknown protein
Length = 135
Score = 49.3 bits (116), Expect = 3e-07
Identities = 26/65 (40%), Positives = 34/65 (52%), Gaps = 2/65 (3%)
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEA-HSIHRRFQIYRYSYQDVFRHSELQKHFD 65
C HR+ NE N FC+ C FC +C+ + H H QI R SY DV R SE++ D
Sbjct: 19 CPRHRETPRNECNMFCLSCQNAAFCFYCRSSFHIDHPVLQIRRSSYHDVVRVSEIENALD 78
Query: 66 CSNIQ 70
+Q
Sbjct: 79 IRGVQ 83
>At5g46710 unknown protein
Length = 226
Score = 47.4 bits (111), Expect = 1e-06
Identities = 25/64 (39%), Positives = 32/64 (49%), Gaps = 1/64 (1%)
Query: 8 CDDHRDLRSNEKNTFCVDCAV-RFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H L E +C+DC FC C H HR QI SY +V + E+QK+ D
Sbjct: 43 CKFHGHLPRTECKMYCLDCTNDSFCSLCLSEHENHRTIQIRISSYHNVTKVDEIQKYLDI 102
Query: 67 SNIQ 70
S+IQ
Sbjct: 103 SSIQ 106
>At5g19460 unknown protein
Length = 374
Score = 27.7 bits (60), Expect = 1.1
Identities = 15/39 (38%), Positives = 21/39 (53%)
Query: 40 IHRRFQIYRYSYQDVFRHSELQKHFDCSNIQVTLNALLQ 78
IH+RF Y + D+F S+ D + VTLN +LQ
Sbjct: 111 IHKRFTEYLREFHDIFTFSQNGSCPDRVDGYVTLNLMLQ 149
>At4g30070 hypothetical protein
Length = 161
Score = 26.6 bits (57), Expect = 2.3
Identities = 10/33 (30%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Query: 4 SSGYCDDHRDLRSNEKNTFCVDCAVRFCRHCKE 36
+SGYCD+ + + + C+D + +C +C E
Sbjct: 28 ASGYCDNDGAVVNQARGDTCID-GLGYCNNCDE 59
>At2g31380 B-box zinc finger protein (STH)
Length = 238
Score = 26.6 bits (57), Expect = 2.3
Identities = 20/59 (33%), Positives = 27/59 (44%), Gaps = 10/59 (16%)
Query: 22 FCVDCAVRFCRHCKEA-------HSIHRRFQI--YRYSYQDVFRHSELQK-HFDCSNIQ 70
FCV+ CR C EA + H+RF R + + E++K HFD SN Q
Sbjct: 68 FCVEDRALLCRDCDEATHAPNTRSANHQRFLATGIRVALSSTSCNQEVEKNHFDPSNQQ 126
>At2g40930 ubiquitin-specific protease (UBP5)
Length = 924
Score = 26.2 bits (56), Expect = 3.1
Identities = 21/72 (29%), Positives = 36/72 (49%), Gaps = 3/72 (4%)
Query: 10 DHRDLRSNEKNTFCVDCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCSNI 69
DH +R ++K T + R C + S H R Y Y +Q ++L K D +N+
Sbjct: 171 DHSAIRISKKETIR-ELHRRACE-IFDLDSEHVRIWDY-YGHQKYSLMNDLDKTLDDANL 227
Query: 70 QVTLNALLQVID 81
Q+ + L++V+D
Sbjct: 228 QMDQDILVEVLD 239
>At1g27360 squamosa-promoter binding protein-like 11
Length = 393
Score = 26.2 bits (56), Expect = 3.1
Identities = 14/45 (31%), Positives = 24/45 (53%), Gaps = 8/45 (17%)
Query: 1 MNSSSGYCDDHRDLRSNEKNTFCVDCAV-----RFCRHCKEAHSI 40
++S+ GY HR + EK++ C +V RFC+ C H++
Sbjct: 184 LSSAKGY---HRKHKVCEKHSKCPKVSVSGLERRFCQQCSRFHAV 225
>At5g60740 xABC transporter - like protein
Length = 1061
Score = 25.8 bits (55), Expect = 4.0
Identities = 9/54 (16%), Positives = 28/54 (51%), Gaps = 4/54 (7%)
Query: 32 RHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCSNIQVTLNALLQVIDSFSL 85
+H + ++H + Q++RY+Y + + +Q+ N +T + ++ + + +
Sbjct: 442 KHAPKGKALHTQSQMFRYAYGQIEKEKAMQE----QNKNLTFSGVISMANDIDI 491
>At5g40320 putative protein
Length = 594
Score = 25.8 bits (55), Expect = 4.0
Identities = 15/47 (31%), Positives = 23/47 (48%), Gaps = 6/47 (12%)
Query: 3 SSSGYCDDHRDLRSNEKNTF-CVDCAVRFCRHCKEA-----HSIHRR 43
SS+G CD +LR N + + C C + + C E+ H H+R
Sbjct: 61 SSTGQCDGCPELRENGLHGYNCNTCGLTVHKECAESSPEINHPSHQR 107
>At3g16830 unknown protein
Length = 1131
Score = 24.6 bits (52), Expect = 8.9
Identities = 11/31 (35%), Positives = 15/31 (47%), Gaps = 4/31 (12%)
Query: 38 HSIHRRFQIYRYSYQDVFRHSELQKHFDCSN 68
H IH +Y Y D+ +H E+ H C N
Sbjct: 431 HLIH----VYAYQGSDLRQHLEIDAHVGCVN 457
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.329 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,875,780
Number of Sequences: 26719
Number of extensions: 64590
Number of successful extensions: 250
Number of sequences better than 10.0: 17
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 233
Number of HSP's gapped (non-prelim): 17
length of query: 85
length of database: 11,318,596
effective HSP length: 61
effective length of query: 24
effective length of database: 9,688,737
effective search space: 232529688
effective search space used: 232529688
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC142094.3


