

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
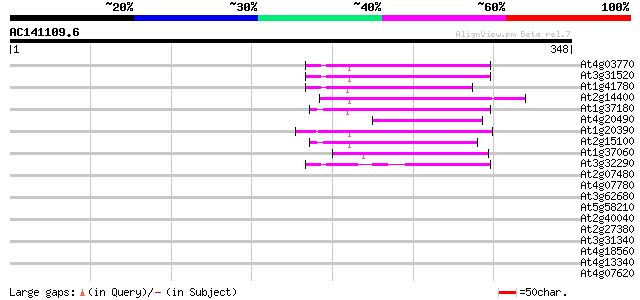
Query= AC141109.6 - phase: 0
(348 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g03770 hypothetical protein 65 7e-11
At3g31520 hypothetical protein 62 5e-10
At1g41780 hypothetical protein 54 2e-07
At2g14400 putative retroelement pol polyprotein 52 4e-07
At1g37180 hypothetical protein 52 6e-07
At4g20490 putative protein 48 9e-06
At1g20390 hypothetical protein 47 2e-05
At2g15100 putative retroelement pol polyprotein 42 6e-04
At1g37060 Athila retroelment ORF 1, putative 42 6e-04
At3g32290 hypothetical protein 41 8e-04
At2g07480 F9A16.15 39 0.005
At4g07780 putative athila transposon protein 38 0.009
At3g62680 proline-rich protein 35 0.047
At5g58210 unknown protein 35 0.061
At2g40040 unknown protein 34 0.10
At2g27380 hypothetical protein 34 0.10
At3g31340 Athila ORF 1, putative 34 0.14
At4g18560 pherophorin - like protein 33 0.18
At4g13340 extensin-like protein 33 0.18
At4g07620 putative athila transposon protein 33 0.23
>At4g03770 hypothetical protein
Length = 464
Score = 64.7 bits (156), Expect = 7e-11
Identities = 34/119 (28%), Positives = 63/119 (52%), Gaps = 7/119 (5%)
Query: 184 PHKFKVPYFEKYKGNSCPEEHLKMYV----RRMSTYARNDQVFIYYFQESLASPASKWYT 239
P K ++ YF G+S P+ HLK+++ R D + F E L PA W+T
Sbjct: 231 PGKLRIEYFN---GSSDPKGHLKLFIISVARAKFRPEERDAGLCHLFVEHLKGPALNWFT 287
Query: 240 NLDKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRDTATQV 298
L + +F +LS +F++QY+ +D + +DL +++Q+ E +++ ++R T +V
Sbjct: 288 RLKGNSVDSFQELSTLFLKQYSVLIDPSTSDADLWSLSQQPNEPLRDFLAKFRSTLAKV 346
>At3g31520 hypothetical protein
Length = 528
Score = 62.0 bits (149), Expect = 5e-10
Identities = 33/119 (27%), Positives = 62/119 (51%), Gaps = 7/119 (5%)
Query: 184 PHKFKVPYFEKYKGNSCPEEHLKMYV----RRMSTYARNDQVFIYYFQESLASPASKWYT 239
P K ++ YF G+S P+ HLK ++ R D + F E L PA W++
Sbjct: 239 PGKLRIEYFN---GSSDPKRHLKSFIISVARTKFRPEERDAGLCHLFVEHLKGPALCWFS 295
Query: 240 NLDKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRDTATQV 298
L+ + +F +LS +F++QY+ +D+ +DL +++Q+ E +++ + R T +V
Sbjct: 296 RLEGNSMDSFQELSTLFLKQYSVLIDLGTSDADLWSLSQQPNEPLRDFLAKLRSTLAKV 354
>At1g41780 hypothetical protein
Length = 246
Score = 53.5 bits (127), Expect = 2e-07
Identities = 29/108 (26%), Positives = 53/108 (48%), Gaps = 7/108 (6%)
Query: 184 PHKFKVPYFEKYKGNSCPEEHLKMY----VRRMSTYARNDQVFIYYFQESLASPASKWYT 239
P K ++ YF G+S P+ HLK + R +D + F E L P W++
Sbjct: 138 PEKLRIEYFS---GSSDPKGHLKSFKISVARAKFKPEESDAGLCHLFVEHLKGPVLDWFS 194
Query: 240 NLDKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEY 287
L+ + +F +LS +F++QY+ +D DL +++Q+ E ++
Sbjct: 195 RLEGNSVDSFQELSTLFLKQYSVLIDPGTSDVDLWSLSQQPNEPLSDF 242
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 52.4 bits (124), Expect = 4e-07
Identities = 36/132 (27%), Positives = 64/132 (48%), Gaps = 5/132 (3%)
Query: 193 EKYKGNSCPEEHLKMYV----RRMSTYARNDQVFIYYFQESLASPASKWYTNLDKTRIQT 248
E Y G P+ +L ++ R A D + F E+L A W+T L+ I
Sbjct: 163 ESYNGLEDPKGYLAAFLIAAGRVDLNEADEDVRYCKLFSENLCGQALMWFTQLEPGSISN 222
Query: 249 FCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRDTATQVSPRIEEKEMT 308
F +LS VF++QY+ +D + +DL ++Q ET + + +++ +++S RI ++
Sbjct: 223 FNELSVVFLKQYSILMDKSISDTDLWNLSQGPNETLRAFITKFKYVLSKLS-RISQQSAL 281
Query: 309 KIFLKTLSYTTR 320
K L Y +R
Sbjct: 282 SALRKGLWYDSR 293
>At1g37180 hypothetical protein
Length = 661
Score = 51.6 bits (122), Expect = 6e-07
Identities = 33/116 (28%), Positives = 53/116 (45%), Gaps = 7/116 (6%)
Query: 187 FKVPYFEKYKGNSCPEEHLKMY---VRRMSTYA-RNDQVFIYYFQESLASPASKWYTNLD 242
FK+P Y G P EHL + VRR+ D F E+L+ PA W+T L+
Sbjct: 314 FKLPV---YNGKGDPNEHLTSFQVIVRRVPLEPYEEDAGLCKLFSENLSGPALTWFTQLE 370
Query: 243 KTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRDTATQV 298
+ I F LS F++QY Y + + L ++Q + + Y +++ Q+
Sbjct: 371 EGSIDNFKQLSTAFIKQYGYFIKSDITEAHLWNLSQSADDPLRTYIIVFKEIMVQI 426
>At4g20490 putative protein
Length = 853
Score = 47.8 bits (112), Expect = 9e-06
Identities = 24/68 (35%), Positives = 35/68 (51%)
Query: 226 FQESLASPASKWYTNLDKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFK 285
F E L A ++ L+ I F LS F++QY + SDL +MTQEN ET
Sbjct: 225 FVEGLTGNALTCFSRLEANSIDNFTQLSTAFLKQYRVFIQPGASSSDLWSMTQENGETLN 284
Query: 286 EYAHRWRD 293
+Y R+++
Sbjct: 285 DYLGRFKE 292
>At1g20390 hypothetical protein
Length = 1791
Score = 47.0 bits (110), Expect = 2e-05
Identities = 32/126 (25%), Positives = 56/126 (44%), Gaps = 5/126 (3%)
Query: 178 VLNVQIPHKFKVPYFEKYKGNSCPEEHLKMYV----RRMSTYARNDQVFIYYFQESLASP 233
+ NV I K+ E Y G + P+E L + R T D F E L P
Sbjct: 161 ITNVSIRGAQKIK-LESYNGRNDPKEFLTSFNVAINRAELTIDNFDAGRCQIFIEHLTGP 219
Query: 234 ASKWYTNLDKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRD 293
A W++ L I +F L+ F++ Y ++ +DL +++Q KE+ + + R++
Sbjct: 220 AHNWFSRLKPNSIDSFHQLTSSFLKHYAPLIENQTSNADLWSISQGAKESLRSFVDRFKL 279
Query: 294 TATQVS 299
T ++
Sbjct: 280 VVTNIT 285
>At2g15100 putative retroelement pol polyprotein
Length = 1329
Score = 41.6 bits (96), Expect = 6e-04
Identities = 30/108 (27%), Positives = 45/108 (40%), Gaps = 7/108 (6%)
Query: 187 FKVPYFEKYKGNSCPEEHLKMYV----RRMSTYARNDQVFIYYFQESLASPASKWYTNLD 242
FK+P Y G +EHL + R D F E+L A W+T L+
Sbjct: 173 FKLPV---YNGKGDLKEHLTSFQVIAGRVPLEPHEEDAGLCKLFSENLFGLALTWFTQLE 229
Query: 243 KTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHR 290
+ I F LS F++QY Y ++ + L +Q E + Y +R
Sbjct: 230 EGSIDNFKQLSTAFIKQYEYFINSDITEAHLWNFSQSADEPLRTYIYR 277
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 41.6 bits (96), Expect = 6e-04
Identities = 30/101 (29%), Positives = 46/101 (44%), Gaps = 4/101 (3%)
Query: 201 PEEHLKMYVRRMSTYARN----DQVFIYYFQESLASPASKWYTNLDKTRIQTFCDLSEVF 256
P +HL + R S N D + F SL A +W +L + I ++ D + F
Sbjct: 62 PLDHLDEFDRLCSLTKINRVSEDGFKLRLFPFSLGDKAHQWEKSLPQGSITSWNDCKKAF 121
Query: 257 VEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRDTATQ 297
+ ++ N A R+D+ TQ N ETF E R++ TQ
Sbjct: 122 LAKFFSNSRTARLRNDISGFTQTNNETFYEAWERFKGYQTQ 162
>At3g32290 hypothetical protein
Length = 612
Score = 41.2 bits (95), Expect = 8e-04
Identities = 27/115 (23%), Positives = 57/115 (49%), Gaps = 21/115 (18%)
Query: 184 PHKFKVPYFEKYKGNSCPEEHLKMYVRRMSTYARNDQVFIYYFQESLASPASKWYTNLDK 243
P K ++ YF G+S P+ HLK ++ ++ + QE + S
Sbjct: 177 PGKLRIEYFN---GSSDPKGHLKSFIISVARAK--------FRQEERDAGNS-------- 217
Query: 244 TRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRDTATQV 298
+ +F +LS +F++QY+ +D+ +DL +++Q+ E +++ ++R T +V
Sbjct: 218 --VDSFQELSTLFLKQYSVLIDLGTSDADLWSLSQQPNEPLRDFLAKFRSTLAKV 270
>At2g07480 F9A16.15
Length = 1012
Score = 38.5 bits (88), Expect = 0.005
Identities = 28/101 (27%), Positives = 43/101 (41%), Gaps = 4/101 (3%)
Query: 201 PEEHLKMYVRRMSTYARN----DQVFIYYFQESLASPASKWYTNLDKTRIQTFCDLSEVF 256
P +HL + R S N + + F SL A W L I T+ D + F
Sbjct: 57 PLDHLDNFDRLCSLTKINGVSEESFKLRLFPFSLGDKAHLWEKTLPVKSIDTWDDCKKAF 116
Query: 257 VEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRDTATQ 297
+ ++ N A RS++ Q+N E+F E R++ TQ
Sbjct: 117 LAKFFSNSRKARLRSEISGFNQKNSESFSEAWERFKGYTTQ 157
>At4g07780 putative athila transposon protein
Length = 446
Score = 37.7 bits (86), Expect = 0.009
Identities = 22/78 (28%), Positives = 36/78 (45%)
Query: 219 DQVFIYYFQESLASPASKWYTNLDKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQ 278
D + F SL A W L I ++ D + F+ ++ N A R+++ TQ
Sbjct: 92 DMFKLRLFPFSLGDKAHHWKKTLPPDSITSWDDCKKDFLAKFFSNARTARLRNEISGFTQ 151
Query: 279 ENKETFKEYAHRWRDTAT 296
+N ETF E + R++ T
Sbjct: 152 KNNETFFEASERFKSYTT 169
>At3g62680 proline-rich protein
Length = 313
Score = 35.4 bits (80), Expect = 0.047
Identities = 27/97 (27%), Positives = 38/97 (38%), Gaps = 5/97 (5%)
Query: 39 PPPPPVRTQVEASSSTISE-WTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPT 97
P PPV E + S +T PT S P ++ P PP + P +
Sbjct: 27 PSSPPVYKSPEHKPTLPSPVYTPPVYKPTLSPPVYTKPTIPP----PVYTPPVYKHTPSP 82
Query: 98 HQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEPIF 134
Y PPPV P V +P ++T P I P++
Sbjct: 83 PVYTKPTIPPPVYTPPVYKPTLSPPVYTKPTIPPPVY 119
Score = 32.7 bits (73), Expect = 0.30
Identities = 18/81 (22%), Positives = 33/81 (40%), Gaps = 5/81 (6%)
Query: 54 TISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTHQYVAHVPPPPVRAPL 113
T +++ + P + +P+H P + P + + Y PPPV P
Sbjct: 20 TTADYYSPSSPPVYKSPEHK-----PTLPSPVYTPPVYKPTLSPPVYTKPTIPPPVYTPP 74
Query: 114 VVMTYSAPMIHTVPQIEEPIF 134
V +P ++T P I P++
Sbjct: 75 VYKHTPSPPVYTKPTIPPPVY 95
Score = 29.3 bits (64), Expect = 3.3
Identities = 28/94 (29%), Positives = 34/94 (35%), Gaps = 11/94 (11%)
Query: 40 PPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTHQ 99
P PPV T+ +T PT S P ++ P PP + P PT
Sbjct: 80 PSPPVYTKPTIPPPV---YTPPVYKPTLSPPVYTKPTIPP----PVYTP---PVYKPTPV 129
Query: 100 YVAHVPPPPVRAPLVVM-TYSAPMIHTVPQIEEP 132
Y PPPV P V T S P+ P P
Sbjct: 130 YTKPTIPPPVYTPPVYKPTPSPPVYKKSPSYSSP 163
>At5g58210 unknown protein
Length = 380
Score = 35.0 bits (79), Expect = 0.061
Identities = 34/129 (26%), Positives = 57/129 (43%), Gaps = 13/129 (10%)
Query: 17 EAQAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPW 76
EA+A ++V A A + P P P+ VEA + + ++ ADT +H +P
Sbjct: 136 EAKALSEVSPSVPADASSHLS-PSPVPI---VEAKALSEVSPSVPADTSSHFSPAP---- 187
Query: 77 FPPFTTGEILCPIACEAQMPTHQYVAHVPPPPVRA-PLVVMTYSAP---MIHTVPQIEEP 132
P E L ++ T + + P P V A L ++ S P H P + EP
Sbjct: 188 -VPIVEAEALSVVSPSVPADTSSHFSPAPVPIVEAKTLSEVSPSVPDDASSHFSPVVVEP 246
Query: 133 IFHSGNMEV 141
F +G++++
Sbjct: 247 EFQAGSVDI 255
>At2g40040 unknown protein
Length = 839
Score = 34.3 bits (77), Expect = 0.10
Identities = 16/61 (26%), Positives = 38/61 (62%), Gaps = 4/61 (6%)
Query: 13 KTMAEAQAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQH 72
++ +++Q+Q+Q +Q+Q+Q+ +Q+Q P +TQ ++ S T ++ A +P+ +P
Sbjct: 781 QSQSQSQSQSQSQSQSQSQSQSQSQSQSQSPSQTQTQSPSQTQAQ----AQSPSSQSPSQ 836
Query: 73 S 73
+
Sbjct: 837 T 837
Score = 31.2 bits (69), Expect = 0.88
Identities = 22/84 (26%), Positives = 40/84 (47%), Gaps = 1/84 (1%)
Query: 17 EAQAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQ-HSMP 75
+ Q Q+Q +Q +AQ+ +QAQ P ++Q ++ S + S+ + + + S Q S
Sbjct: 749 QTQTQSQSPSQTRAQSPSQAQAQSPSQTQSQSQSQSQSQSQSQSQSQSQSQSQSQSQSQS 808
Query: 76 WFPPFTTGEILCPIACEAQMPTHQ 99
P T + +AQ P+ Q
Sbjct: 809 QSPSQTQTQSPSQTQAQAQSPSSQ 832
Score = 28.5 bits (62), Expect = 5.7
Identities = 16/56 (28%), Positives = 31/56 (54%)
Query: 16 AEAQAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQ 71
++AQAQ+ Q+Q+Q+ +Q+Q ++Q ++ S + S+ + T T S Q
Sbjct: 766 SQAQAQSPSQTQSQSQSQSQSQSQSQSQSQSQSQSQSQSQSQSQSPSQTQTQSPSQ 821
Score = 27.7 bits (60), Expect = 9.7
Identities = 14/59 (23%), Positives = 33/59 (55%)
Query: 13 KTMAEAQAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQ 71
+T A++ +QAQ + +Q Q+ +Q+Q ++Q ++ S + S+ + +P+ + Q
Sbjct: 759 QTRAQSPSQAQAQSPSQTQSQSQSQSQSQSQSQSQSQSQSQSQSQSQSQSQSPSQTQTQ 817
>At2g27380 hypothetical protein
Length = 761
Score = 34.3 bits (77), Expect = 0.10
Identities = 30/103 (29%), Positives = 39/103 (37%), Gaps = 19/103 (18%)
Query: 38 VPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMP--------WFPPFTTGEILCPI 89
V PPP + S I + TPT+S P P + PP I P
Sbjct: 453 VKPPPVHKPPTPTYSPPIKPPPVKPPTPTYSPPVQPPPVQKPPTPTYSPPVKPPPIQKP- 511
Query: 90 ACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEP 132
PT Y + PPPV+ P TYS P+ P + +P
Sbjct: 512 ------PTPTYSPPIKPPPVKPP--TPTYSPPI--KPPPVHKP 544
Score = 32.3 bits (72), Expect = 0.39
Identities = 30/103 (29%), Positives = 38/103 (36%), Gaps = 18/103 (17%)
Query: 38 VPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMP--------WFPPFTTGEILCPI 89
V PPP + S I + TPT+S P P + PP I P
Sbjct: 503 VKPPPIQKPPTPTYSPPIKPPPVKPPTPTYSPPIKPPPVHKPPTPTYSPPIKPPPIHKP- 561
Query: 90 ACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEP 132
PT Y + PPPV P TYS P+ P + +P
Sbjct: 562 ------PTPTYSPPIKPPPVHKP-PTPTYSPPI--KPPPVHKP 595
Score = 29.3 bits (64), Expect = 3.3
Identities = 25/77 (32%), Positives = 32/77 (41%), Gaps = 18/77 (23%)
Query: 64 TPTHSAPQHSMP--------WFPPFTTGEILCPIACEAQMPTHQYVAHVPPPPVRAPLVV 115
TPT+S P + P + PP I P PT Y V PPPV+ P
Sbjct: 109 TPTYSPPIYPPPIQKPPTPTYSPPIYPPPIQKP-------PTPSYSPPVKPPPVQMP-PT 160
Query: 116 MTYSAPMIHTVPQIEEP 132
TYS P+ P + +P
Sbjct: 161 PTYSPPI--KPPPVHKP 175
Score = 29.3 bits (64), Expect = 3.3
Identities = 23/77 (29%), Positives = 32/77 (40%), Gaps = 19/77 (24%)
Query: 64 TPTHSAPQHSMP--------WFPPFTTGEILCPIACEAQMPTHQYVAHVPPPPVRAPLVV 115
TPT+S P S P + PP + P PT Y + PPPV+ P +
Sbjct: 295 TPTYSPPVKSPPVQKPPTPTYSPPIKPPPVQKP-------PTPTYSPPIKPPPVKPPTPI 347
Query: 116 MTYSAPMIHTVPQIEEP 132
YS P+ P + +P
Sbjct: 348 --YSPPV--KPPPVHKP 360
Score = 28.9 bits (63), Expect = 4.4
Identities = 24/93 (25%), Positives = 33/93 (34%), Gaps = 16/93 (17%)
Query: 38 VPPPPPVRTQVEASSSTISEWTI-CADTPTHSAPQHSMP--------WFPPFTTGEILCP 88
+ PPP + S + + TPT+S P P + PP + P
Sbjct: 655 IKPPPVQKPPTPTYSPPVKPPPVQLPPTPTYSPPVKPPPVQVPPTPTYSPPVKPPPVQVP 714
Query: 89 IACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAP 121
PT Y + PPPV+ P T S P
Sbjct: 715 -------PTPTYSPPIKPPPVQVPPTPTTPSPP 740
Score = 28.5 bits (62), Expect = 5.7
Identities = 26/95 (27%), Positives = 34/95 (35%), Gaps = 19/95 (20%)
Query: 38 VPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPT 97
+ PPP + S I + TP +S P P P PT
Sbjct: 319 IKPPPVQKPPTPTYSPPIKPPPVKPPTPIYSPPVKPPPVHKP----------------PT 362
Query: 98 HQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEP 132
Y V PPPV P + YS P+ P I++P
Sbjct: 363 PIYSPPVKPPPVHKPPTPI-YSPPV--KPPPIQKP 394
Score = 28.5 bits (62), Expect = 5.7
Identities = 27/96 (28%), Positives = 37/96 (38%), Gaps = 21/96 (21%)
Query: 39 PP--PPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMP 96
PP PPP++ S TPT+S P + P P P
Sbjct: 65 PPIYPPPIQKPPTYSPPIYPPPIQKPPTPTYSPPIYPPPIQKP----------------P 108
Query: 97 THQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEP 132
T Y + PPP++ P TYS P+ P I++P
Sbjct: 109 TPTYSPPIYPPPIQKP-PTPTYSPPIY--PPPIQKP 141
>At3g31340 Athila ORF 1, putative
Length = 781
Score = 33.9 bits (76), Expect = 0.14
Identities = 19/65 (29%), Positives = 32/65 (49%)
Query: 229 SLASPASKWYTNLDKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYA 288
SLA A +W +LD ++++ D F+ QY A R+ + + Q E+F E
Sbjct: 142 SLADNAHRWLKSLDPINLRSWEDYKAAFLGQYFTQSRTAILRNKISSFQQGGTESFPEAW 201
Query: 289 HRWRD 293
R++D
Sbjct: 202 ERFKD 206
>At4g18560 pherophorin - like protein
Length = 637
Score = 33.5 bits (75), Expect = 0.18
Identities = 31/120 (25%), Positives = 55/120 (45%), Gaps = 18/120 (15%)
Query: 20 AQAQVLAQAQAQAHAQAQVP--PPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWF 77
+ ++ L ++ + + +++VP P PP + + ST + AD P PQ S+P
Sbjct: 244 SNSEELTESSSLSTVRSRVPRVPKPPPKRSISLGDSTENR----ADPP----PQKSIPPP 295
Query: 78 PPFTTGEILCPIACEAQMPTHQYVAHVPPPPVR-APLVVMTYSAPMIHTVPQIEEPIFHS 136
PP +L Q P V+ PPPP P ++ ++ + VP++ E +HS
Sbjct: 296 PPPPPPPLL------QQPPPPPSVSKAPPPPPPPPPPKSLSIASAKVRRVPEVVE-FYHS 348
>At4g13340 extensin-like protein
Length = 760
Score = 33.5 bits (75), Expect = 0.18
Identities = 28/95 (29%), Positives = 33/95 (34%), Gaps = 5/95 (5%)
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTH 98
PPPPPV + S C P P HS P PP + P + P H
Sbjct: 512 PPPPPVYSSPPPPPSPAPTPVYCTRPPP--PPPHSPP--PPQFSPPPPEPYYYSSPPPPH 567
Query: 99 QYVA-HVPPPPVRAPLVVMTYSAPMIHTVPQIEEP 132
H PPPP P + Y +P P P
Sbjct: 568 SSPPPHSPPPPHSPPPPIYPYLSPPPPPTPVSSPP 602
Score = 32.3 bits (72), Expect = 0.39
Identities = 30/94 (31%), Positives = 36/94 (37%), Gaps = 18/94 (19%)
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTH 98
PPPPPV SS E + +P S +S P PP CE P
Sbjct: 662 PPPPPVHY----SSPPPPE--VHYHSPPPSPVHYSSPPPPPSAP--------CEESPPPA 707
Query: 99 QYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEP 132
V H PPP P+V + P+IH P P
Sbjct: 708 PVVHHSPPP----PMVHHSPPPPVIHQSPPPPSP 737
Score = 30.4 bits (67), Expect = 1.5
Identities = 29/96 (30%), Positives = 38/96 (39%), Gaps = 17/96 (17%)
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQ-HSMPWFPPFTTGEILCPIACEAQMPT 97
PPPP + S + + P HS+P HS P PP + P
Sbjct: 539 PPPPHSPPPPQFSPPPPEPYYYSSPPPPHSSPPPHSPP--PPHSPPP-----------PI 585
Query: 98 HQYVAHVPPP-PVRAPLVVMTYSAPMIHTVPQIEEP 132
+ Y++ PPP PV +P YS P P IE P
Sbjct: 586 YPYLSPPPPPTPVSSPPPTPVYSPP--PPPPCIEPP 619
>At4g07620 putative athila transposon protein
Length = 683
Score = 33.1 bits (74), Expect = 0.23
Identities = 17/52 (32%), Positives = 29/52 (55%)
Query: 246 IQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETFKEYAHRWRDTATQ 297
I ++ D +VF+ ++ NV A R+++ Q+N ETF E R++ TQ
Sbjct: 90 INSWDDCKKVFLAKFFSNVRTARLRNEISGFNQKNNETFCEAWERFKSYTTQ 141
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.319 0.130 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,174,381
Number of Sequences: 26719
Number of extensions: 357678
Number of successful extensions: 1976
Number of sequences better than 10.0: 80
Number of HSP's better than 10.0 without gapping: 29
Number of HSP's successfully gapped in prelim test: 53
Number of HSP's that attempted gapping in prelim test: 1551
Number of HSP's gapped (non-prelim): 367
length of query: 348
length of database: 11,318,596
effective HSP length: 100
effective length of query: 248
effective length of database: 8,646,696
effective search space: 2144380608
effective search space used: 2144380608
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC141109.6


