

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140023.16 - phase: 0
(1060 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
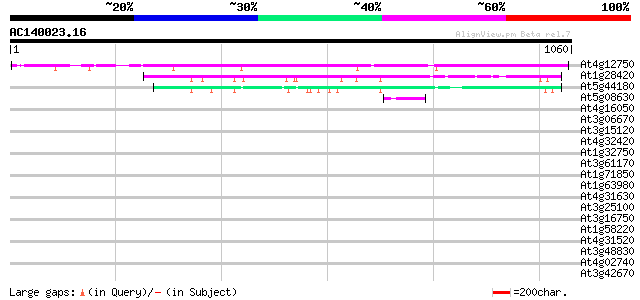
Score E
Sequences producing significant alignments: (bits) Value
At4g12750 putative protein 862 0.0
At1g28420 hypothetical protein 212 1e-54
At5g44180 unknown protein 162 1e-39
At5g08630 unknown protein 48 3e-05
At4g16050 hypothetical protein 37 0.078
At3g06670 unknown protein 35 0.17
At3g15120 chaperone-like ATPase 35 0.23
At4g32420 cyclophilin (AtCYP95) 35 0.29
At1g32750 unknown protein 34 0.50
At3g61170 putative protein 33 0.66
At1g71850 hypothetical protein 33 0.66
At1g63980 unknown protein 33 0.66
At4g31630 putative protein 33 0.86
At3g25100 Cdc45-like protein 33 0.86
At3g16750 hypothetical protein 33 0.86
At1g58220 MYB-family transcription factor-like protein 33 0.86
At4g31520 putative protein 33 1.1
At3g48830 putative protein 33 1.1
At4g02740 hypothetical protein 32 1.5
At3g42670 putative protein 32 1.5
>At4g12750 putative protein
Length = 1108
Score = 862 bits (2226), Expect = 0.0
Identities = 518/1128 (45%), Positives = 671/1128 (58%), Gaps = 145/1128 (12%)
Query: 3 GKGLIVPPCYSRGGKVLQREKVDATSSSSYEMVVNGNWKKNRKRKSVQELYTTGDIVNTV 62
G G P Y R L R +TSS + V ++ S Q L + I+ V
Sbjct: 36 GLGAKNPQLYDRS---LMRS---STSSRCVGVAVEERCIVGTRKASCQNLLPSSHILAKV 89
Query: 63 LLNDAPTLGSEFDSLPSGPKNYN-----SACQQDQEPVKRRKASKSAIQSHPNCNMKAPV 117
D P+LGSEFD LPSG + + S QQ Q+ ++RK S+ + +C +
Sbjct: 90 FRKDGPSLGSEFDHLPSGARKASWLGTSSVGQQKQKVARKRKISELMDHTSQDCIQE--- 146
Query: 118 ERHGMGKGLATNPNCKMKAPVKRHGMGKGLAT-----NPNCNMKAPVKRHGMGKGLAANP 172
A V +HG+GKGL T NPN +P + A P
Sbjct: 147 -----------------NATVMKHGIGKGLMTVWRVMNPNRRDVSPCV--DLLDERATLP 187
Query: 173 NSNMKAPVKRHGMGKGLMTIWRATNHDARDLPISFGSVDKDVHLTSNTKTPISVNRSQKA 232
S+ + P + + L +I + R S + +++
Sbjct: 188 QSSARNPPHQKKKQRQLASILKQKLLQKR-----------------------STEKKRRS 224
Query: 233 VTTNGESNQYATQNQLPIEKCELALDSSISDAGVDQISMLIDDEELELREIQEGSNLLIC 292
+ E N+ TQ + E CELA D + IS L+DDEELE+RE E N L C
Sbjct: 225 IHREAELNKDETQREFK-ENCELAADGEVFKETCQTISTLVDDEELEMRERHERGNPLTC 283
Query: 293 SDQLAANGMLGGSLC--------------PDVLVKFPPGDVKMKKPIHLQPWDSSPELVK 338
S ++G G LC PD+L KFPP V+M+ P L PW+SSPE VK
Sbjct: 284 SCHHPSSGSHGCFLCKGIAMRSSDSSLLFPDLLPKFPPNSVQMRMPFGLHPWNSSPESVK 343
Query: 339 KLFKDSMLLGQIHVALLTLLLSDIEVELSNGFCPHLNKSCNFLALLHSVENQEYSLDAWR 398
KLFKDS+LLG+IH++LL LLL D+E EL G +L+ SC FLALL SVE+Q LD WR
Sbjct: 344 KLFKDSLLLGKIHLSLLKLLLLDVETELERGSFSNLSISCKFLALLQSVESQILILDMWR 403
Query: 399 RSLNPLTWIEILRQVLVAAGFGSKQGAFQREGLGKEL----------------------- 435
SLN LTW E+LRQ+LVAAG+GS + A Q E L K+L
Sbjct: 404 NSLNSLTWTELLRQILVAAGYGSLKCAVQSEELSKQLASTCFVLGDRSVICGELKALARL 463
Query: 436 ---------DILVNYGLCPGTLKCELFKILSERGNNGCKVSELAKSMQIAELNLSSTTEE 486
++ YGL GTLK ELF++L+ +GNNG K+SELA + ++A LNL++ EE
Sbjct: 464 YFVIDDIHMKLMKKYGLRLGTLKGELFRMLNGQGNNGLKISELADAPEVAVLNLATVPEE 523
Query: 487 LESLIYSTLSSDITLFEKISSSAYRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSS 546
E+ I STL+SDITLFEKIS S YR+R++ ++D D SQSD++DSGSVDDE +D + SS
Sbjct: 524 RENSICSTLASDITLFEKISESTYRVRVNCFSEDPDKSQSDSDDSGSVDDE-SDDCSISS 582
Query: 547 GDDFGSGSIHSNIRKLRRHNSRKAKHNKLKVYTEIDESHAGEVWLLGLMDSEYSDLKIEE 606
GD+ S + +RK++ RK K +V +EIDESH GE WLLGLM+ EYSDL +EE
Sbjct: 583 GDEIEHVSENPALRKVKCRKRRKHKSKMREVCSEIDESHPGEPWLLGLMEGEYSDLSVEE 642
Query: 607 KLNALAALTGLLSSGSSIRMKDPVKVTADCSSSIQLRGSGAKIKRSVNPI---------- 656
KL+ AL LLSSGS+IRM+D + ADC+ SI GSG KIKRS +
Sbjct: 643 KLDVFVALIDLLSSGSTIRMEDLPRAVADCAPSIYSHGSGGKIKRSSSNQYSYPRGSWVH 702
Query: 657 -EQMQCTKEVHMNSHACPVDSSLLVSKFHIQEASLEKRKVSAYSHPIQSVFLGSDRRYNR 715
++ K + +S + PVDSS +V F A L + + HP+QSV+LGSDRR+NR
Sbjct: 703 GGELYGIKALSKSSDSHPVDSSSIVGAF----AKLAGNRANNV-HPMQSVYLGSDRRFNR 757
Query: 716 YWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLER 775
YWLFLG CN +DPGHR V+FESSEDGHWEVI+ +EAL ALLSVLDDRG+REA LIESLE+
Sbjct: 758 YWLFLGTCNANDPGHRCVFFESSEDGHWEVINNKEALRALLSVLDDRGRREARLIESLEK 817
Query: 776 RQTSLCRSMSRIKVSNIGMGCMSHSDQSE----LDRVAEDSCSPVSDVD-NLNLTEITD- 829
R++ LC++M +V+ QSE D V EDS SPVSD+D NL L EI +
Sbjct: 818 RESFLCQAMLSRQVT-----------QSETAHFTDIVREDSSSPVSDIDNNLCLNEIAND 866
Query: 830 -YLPSPGAVVIEAGKKEEEQLHKWIRVQEYDSWIWNSFYLDLNVVKYGRRSYLDSLARCR 888
+ A+V E G K E+ L W +QE+D WIW +F +LN VK+ RRSYLDSL RC+
Sbjct: 867 QFSSQHAAIVFEIGSKREKSL-LWSLIQEFDDWIWANFNFNLNSVKHRRRSYLDSLTRCK 925
Query: 889 SCHDLYWRDERHCKICHMTFELDFDLEEKYAIHIAMCREKEDSNTFPNHKVLPSQIQSLK 948
SCHDLYWRDE+HCKICH TFE+D DLEE+YAIH A C KE+ +TFP+HKVL SQ+QSLK
Sbjct: 926 SCHDLYWRDEKHCKICHATFEVDIDLEERYAIHAATCMRKEECDTFPDHKVLSSQLQSLK 985
Query: 949 AAIYAIESVMPEDALVGAWRKSAHNLWIKRLRRTSTLVELLQVLADFVGAFNDSWLFQCK 1008
AA+YAIES MPEDAL+GAWRKSAH LW KRLRR+S++ E+ QV+ DFVGA N+ WL+ C
Sbjct: 986 AAVYAIESAMPEDALIGAWRKSAHRLWAKRLRRSSSVSEITQVIGDFVGAINEEWLWHCS 1045
Query: 1009 FP-DGVVEETIASFASMPHTSSALALWLVKLDAIIAPYLDRVQTQKSQ 1055
++ E I F SMP T+SA+ALWLVKLD +IAPY+++ ++ Q
Sbjct: 1046 DQGQTLMGEIINCFPSMPQTTSAIALWLVKLDTLIAPYVEKAPPERDQ 1093
>At1g28420 hypothetical protein
Length = 1703
Score = 212 bits (539), Expect = 1e-54
Identities = 247/985 (25%), Positives = 399/985 (40%), Gaps = 220/985 (22%)
Query: 254 ELALDSSISDAGVDQISMLIDDEELELREIQEGSNLLICSDQLAANGMLGGSLCPDVLVK 313
+LA++ + + + LI+DE+LEL E+ S L QL + + + D L
Sbjct: 467 KLAIEKATARRIAKESMDLIEDEQLELMELAAISKGLPSVLQLDHDTLQNLEVYRDSLST 526
Query: 314 FPPGDVKMKKPIHLQPWDSSPELVKKLFK-----------------------------DS 344
FPP +++K P + PW S E V L DS
Sbjct: 527 FPPKSLQLKMPFAISPWKDSDETVGNLLMVWRFLISFSDVLDLWPFTLDEFIQAFHDYDS 586
Query: 345 MLLGQIHVALLTLLLSDIE-------VELSNGFCPHLNKSCNFLALLHSVENQEYSLDAW 397
LLG+IHV LL ++ D+E + N N ++ + + +W
Sbjct: 587 RLLGEIHVTLLRSIIRDVEDVARTPFSGIGNNQYTTANPEGGHPQIVEGAYAWGFDIRSW 646
Query: 398 RRSLNPLTWIEILRQVLVAAGFGSK----------------------------QGAFQRE 429
++ LNPLTW EILRQ+ ++AGFG K G
Sbjct: 647 KKHLNPLTWPEILRQLALSAGFGPKLKKKHSRLTNTGDKDEAKGCEDVISTIRNGTAAES 706
Query: 430 GLG--KELDILV----NYGLCPGTLKCELFKILSERGNNGCKVSELAKSMQIAELNLSST 483
+E +L + L PGT+K F +LS G+ G V ELA +Q + L +T
Sbjct: 707 AFASMREKGLLAPRKSRHRLTPGTVKFAAFHVLSLEGSKGLTVLELADKIQKSGLRDLTT 766
Query: 484 TEELESLIYSTLSSDITLFEKISSSAYRLRMSTVA--KD--------------------- 520
++ E+ I L+ D+ LFE+I+ S Y +R V KD
Sbjct: 767 SKTPEASISVALTRDVKLFERIAPSTYCVRAPYVKDPKDGEAILADARKKIRAFENGFTG 826
Query: 521 ---------DDDSQSDTEDSGSVDD--------------ELN--------------DSDT 543
D+D + D ++ VDD E N +D
Sbjct: 827 PEDVNDLERDEDFEIDIDEDPEVDDLATLASASKSAVLGEANVLSGKGVDTMFCDVKADV 886
Query: 544 CSSGDDFGSGSIHSNIRKLRRHNSRKAKHNKLK-VYTEIDESHAGEVWLLGLMDSEYSDL 602
S + S S ++ + +S + K+ + V IDES+ G+ W+ GL + +Y L
Sbjct: 887 KSELEKEFSSPPPSTMKSIVPQHSERHKNTVVGGVDAVIDESNQGQSWIQGLTEGDYCHL 946
Query: 603 KIEEKLNALAALTGLLSSGSSIR---------------------------MKDPVKVTAD 635
+EE+LNAL AL G+ + G+SIR M+D +K+
Sbjct: 947 SVEERLNALVALVGIANEGNSIRTGLEDRMEAANALKKQMWAEAQLDNSCMRDVLKLDLQ 1006
Query: 636 CSSSIQLRGS-GAKIKRSV---------NPIEQMQCTK---EVHMNSHACPVDSSLLVSK 682
+S + + G I +S +P + + TK ++ + H + +L+
Sbjct: 1007 NLASSKTESTIGLPIIQSSTRERDSFDRDPSQLLDETKPLEDLSNDLHKSSAERALINQD 1066
Query: 683 FHI-QEASLEKRKVSAYS----------HPIQSVFLGSDRRYNRYWLFLGPCNIDDPGHR 731
+I QE KR S +P +S+ LG DRR+NRYW F + DP R
Sbjct: 1067 ANISQENYASKRSRSQLKSYIGHKAEEVYPYRSLPLGQDRRHNRYWHFAVSVSKSDPCSR 1126
Query: 732 RVYFESSEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLERRQTSLCRSMSR-IKVS 790
++ E DG W +ID+EEA L++ LD RG RE+ L L++ + S + + IK++
Sbjct: 1127 LLFVEL-HDGKWLLIDSEEAFDILVASLDMRGIRESHLRIMLQKIEGSFKENACKDIKLA 1185
Query: 791 NIGMGCMSHSDQSELDRVAEDSCSPVSD-VDNLNLTEITDYLPSPGAVVIEAGKKEEEQL 849
+++S ++ DS SP S + N +D + + ++ ++ G+ + E
Sbjct: 1186 RNPF----LTEKSVVNHSPTDSVSPSSSAISGSN----SDSMETSTSIRVDLGRNDTENK 1237
Query: 850 HKWIRVQEYDSWIWNSFYLDLNVV--KYGRRSYLDSLARCRSCHDLYWRDERHCKICHMT 907
+ R ++ W+W Y L KYG++ + LA C +C Y + C CH
Sbjct: 1238 NLSKRFHDFQRWMWTETYSSLPSCARKYGKKRS-ELLATCDACVASYLSEYTFCSSCHQR 1296
Query: 908 FELDFDLEEKYAIHIAMCREKEDSNTFPNHKVLPSQIQSLKAAIYAIESVMPEDALVGAW 967
++ D E +A+ LP ++ LK + +E+ +P++AL W
Sbjct: 1297 LDV-VDSSEILDSGLAV-------------SPLPFGVRLLKPLLVFLEASVPDEALESFW 1342
Query: 968 RKSAHNLWIKRLRRTSTLVELLQVLADFVGAFN----DSWLFQCKFPDGVV------EET 1017
+ W RL +S+ ELLQVL A S K G + +
Sbjct: 1343 TEDQRKKWGFRLNTSSSPGELLQVLTSLESAIKKESLSSNFMSAKELLGAANAEADDQGS 1402
Query: 1018 IASFASMPHTSSALALWLVKLDAII 1042
+ +P T SA+AL L +LDA I
Sbjct: 1403 VDVLPWIPKTVSAVALRLSELDASI 1427
>At5g44180 unknown protein
Length = 1694
Score = 162 bits (409), Expect = 1e-39
Identities = 229/984 (23%), Positives = 366/984 (36%), Gaps = 244/984 (24%)
Query: 272 LIDDEELELREIQEGSNLLICSDQLAANGMLGGSLCPDVLVKFPPGDVKMKKPIHLQPWD 331
LI+DE LEL E+ + L L + D FPP VK+KKP ++PW+
Sbjct: 452 LIEDERLELMEVAALTKGLPSMLALDFETLQNLDEYRDKQAIFPPTSVKLKKPFAVKPWN 511
Query: 332 SSPELVKKLFK-----------------------------DSMLLGQIHVALLTLLLSDI 362
S E V L D L+G+IH+ LL ++ DI
Sbjct: 512 GSDENVANLLMVWRFLITFADVLGLWPFTLDEFAQAFHDYDPRLMGEIHIVLLKTIIKDI 571
Query: 363 EV---ELSNGFCPHLNKSCN----FLALLHSVENQEYSLDAWRRSLNPLTWIEILRQVLV 415
E LS G + N + N ++ + + +WR++LN TW EILRQ+ +
Sbjct: 572 EGVVRTLSTGVGANQNVAANPGGGHPHVVEGAYAWGFDIRSWRKNLNVFTWPEILRQLAL 631
Query: 416 AAGFGSK---------------------------------QGAF---QREGLGKELDILV 439
+AG G + + AF Q GL
Sbjct: 632 SAGLGPQLKKMNIRTVSVHDDNEANNSENVIFNLRKGVAAENAFAKMQERGLSNPRRS-- 689
Query: 440 NYGLCPGTLKCELFKILSERGNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDI 499
+ L PGT+K F +LS G G + E+A+ +Q + L +T+ E+ + + LS D
Sbjct: 690 RHRLTPGTVKFAAFHVLSLEGEKGLNILEVAEKIQKSGLRDLTTSRTPEASVAAALSRDT 749
Query: 500 TLFEKISSSAYRLRMSTVAKDDDDSQ--------------SDTEDSGSVDDELNDSDTCS 545
LFE+++ S Y +R S KD D++ S D VDD D D S
Sbjct: 750 KLFERVAPSTYCVRAS-YRKDAGDAETIFAEARERIRAFKSGITDVEDVDDAERDED--S 806
Query: 546 SGDDFGSGSIHSNIRK-----LRRHNS-------RKAKHNKLKVYTEI------------ 581
D + N++K L+ N K + + + TE+
Sbjct: 807 ESDVGEDPEVDVNLKKEDPNPLKVENLIGVEPLLENGKLDTVPMKTELGLPLTPSLPEEM 866
Query: 582 ---------------------------DESHAGEVWLLGLMDSEYSDLK----------- 603
DES GE W+ GL++ +YS+L
Sbjct: 867 KDEKRDDTLADQSLEDAVANGEDSACFDESKLGEQWVQGLVEGDYSNLSSEERLNALVAL 926
Query: 604 -------------IEEKLNALAALTGLL----------SSGSSIRMKDPVKVTADCSSSI 640
+EE+L +AL + S IR TA +I
Sbjct: 927 IGIATEGNTIRIALEERLEVASALKKQMWGEVQLDKRWKEESLIRANYLSYPTAKPGLNI 986
Query: 641 QLRGSGAKIKRS--VNPIEQMQCTKEVHMNSHACPVDSSLLVSKFHIQEASLEKRKVSAY 698
SG + S V PI ++ + SL + + +L+ ++ Y
Sbjct: 987 ATPASGNQESSSADVTPISSQDPVSLPQIDVNNVIAGPSLQLQENVPGVENLQYQQQQGY 1046
Query: 699 S---------------------HPIQSVFLGSDRRYNRYWLFLGPCNIDDPGHRRVYFES 737
+ + +S+ LG DRR NRYW F + +DPG R++ E
Sbjct: 1047 TADRERLRAQLKAYVGYKAEELYVYRSLPLGQDRRRNRYWRFSASASRNDPGCGRIFVEL 1106
Query: 738 SEDGHWEVIDTEEALCALLSVLDDRGKREALLIESLERRQTSLCRSMSRIKVSNIGMGCM 797
+DG W +ID+EEA L+ LD RG RE+ L L + + S ++ R +N G+ +
Sbjct: 1107 -QDGRWRLIDSEEAFDYLVKSLDVRGVRESHLHFMLLKIEASFKEALRRNVAANPGVCSI 1165
Query: 798 SHSDQSELDRVAEDSCSPVSDVDNLNLTEITDYLPSPGAVVIEAGKKEEEQLHKWIRVQE 857
S S S+ AE S + ++ + N E L R
Sbjct: 1166 SSSLDSD---TAEISTTFKIELGDSNAVERCSVLQ---------------------RFHS 1201
Query: 858 YDSWIWNSFY--LDLNVVKYGRRSYLDSLARCRSCHDLYWRDERHCKICHMTFELDFDLE 915
++ W+W++ L+ KYG + CR C +L++ + C C E
Sbjct: 1202 FEKWMWDNMLHPSALSAFKYGAKQSSPLFRICRICAELHFVGDICCPSCGQMHAGPDVGE 1261
Query: 916 EKYAIHIAMCRE---KEDSNTFPNHKVL-PSQIQSLKAAIYAIESVMPEDALVGAWRKSA 971
+A +A + + D+ +L P +I+ LK + +E+ +P + L W ++
Sbjct: 1262 LCFAEQVAQLGDNLRRGDTGFILRSSILSPLRIRLLKVQLALVEASLPPEGLEAFWTENL 1321
Query: 972 HNLWIKRLRRTSTLVELLQVLADFVGAFNDSWLFQCKFPD-----GVVEETIASFAS--- 1023
W +L +S+ +L QVL A +L F G+ E +AS +
Sbjct: 1322 RKSWGMKLLSSSSHEDLYQVLTTLEAALKRDFL-SSNFETTSELLGLQEGALASDLTCGV 1380
Query: 1024 -----MPHTSSALALWLVKLDAII 1042
+P T+ +AL L D+ I
Sbjct: 1381 NVLPWIPKTAGGVALRLFDFDSSI 1404
>At5g08630 unknown protein
Length = 723
Score = 48.1 bits (113), Expect = 3e-05
Identities = 26/80 (32%), Positives = 40/80 (49%), Gaps = 8/80 (10%)
Query: 707 LGSDRRYNRYWLFLGPCNIDDPGHRRVYFESSEDGHWEVIDTEEALCALLSVLDDRGKRE 766
LG DR YNRYW F + R++ E+S+ W +E L AL+ L+ +G+RE
Sbjct: 608 LGKDRDYNRYWWFRS--------NGRIFVENSDSEEWGYYTAKEELDALMGSLNRKGERE 659
Query: 767 ALLIESLERRQTSLCRSMSR 786
L LE +C ++ +
Sbjct: 660 LSLYTQLEIFYDRICSTLQK 679
>At4g16050 hypothetical protein
Length = 900
Score = 36.6 bits (83), Expect = 0.078
Identities = 33/117 (28%), Positives = 52/117 (44%), Gaps = 20/117 (17%)
Query: 466 VSELAKSMQIAELNLSSTTEELESLIYSTLSSDITL--------FEKISSSAYRLRMSTV 517
VSE+ K + N+SST + Y DI L ++K+ + + ST
Sbjct: 680 VSEMRKE-SMETFNVSSTVDH-----YDDSDDDIPLKIVPLSQVYQKLDDEMKKAKHSTN 733
Query: 518 -----AKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKLRRHNSRK 569
A++DD+S ++TED S D E +D + D+ N+ +R NSRK
Sbjct: 734 KRRKRAREDDESAAETEDDESADTE-DDESADTEDDESAETEDDDNMTIAQRINSRK 789
>At3g06670 unknown protein
Length = 863
Score = 35.4 bits (80), Expect = 0.17
Identities = 26/72 (36%), Positives = 37/72 (51%), Gaps = 1/72 (1%)
Query: 478 LNLSSTTEELESLIYSTLSSDITLFEKISSSAYRLRMSTVAKDDDDSQSDTE-DSGSVDD 536
L SS E + +++ STLS + +K A S VAK +DS+S E +S S DD
Sbjct: 762 LGKSSKRENVFAVLCSTLSHAVLTGKKSPGPAGSAARSIVAKGAEDSKSSEENNSSSSDD 821
Query: 537 ELNDSDTCSSGD 548
E + D SS +
Sbjct: 822 ENHKDDGVSSSE 833
>At3g15120 chaperone-like ATPase
Length = 1954
Score = 35.0 bits (79), Expect = 0.23
Identities = 39/149 (26%), Positives = 60/149 (40%), Gaps = 7/149 (4%)
Query: 113 MKAPVERHGMGKGLATNPNCKMKAPVKRHGMGKGLATNPNCNMKAPVKRHGMGKGLAANP 172
M V G G G+ + N +M V G G G+ N M+ V G G G+ +
Sbjct: 329 MSVIVSESGNGTGIREDENKEMDVIVSESGNGTGILEGENKKMEVMVSGSGNGTGIRED- 387
Query: 173 NSNMKAPVK-RHGMGKGLMTIWRATNHDARDLPISFGSVDKDVHLT-SNTKTPISVNRSQ 230
+S+ A VK R G + A+ L + ++ V T SN KT
Sbjct: 388 DSDFAAKVKNREGDTLHPELLGEASTEINESLKQNDDIGEQGVSRTPSNNKT----KEHN 443
Query: 231 KAVTTNGESNQYATQNQLPIEKCELALDS 259
+ + GES + + + E C+ A+DS
Sbjct: 444 EFLDRGGESVEMPDELPIQNETCKKAVDS 472
Score = 31.6 bits (70), Expect = 2.5
Identities = 17/57 (29%), Positives = 23/57 (39%)
Query: 133 KMKAPVKRHGMGKGLATNPNCNMKAPVKRHGMGKGLAANPNSNMKAPVKRHGMGKGL 189
+M V G G G+ + N M V G G G+ N M+ V G G G+
Sbjct: 328 EMSVIVSESGNGTGIREDENKEMDVIVSESGNGTGILEGENKKMEVMVSGSGNGTGI 384
>At4g32420 cyclophilin (AtCYP95)
Length = 837
Score = 34.7 bits (78), Expect = 0.29
Identities = 21/59 (35%), Positives = 31/59 (51%), Gaps = 3/59 (5%)
Query: 518 AKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKLRRHNSRKAKHNKLK 576
+ + +S SD+E S D +DSD S F S S H +K R+ +S+K KH + K
Sbjct: 205 SSSESESSSDSETDSSESDSESDSDL--SSPSFLSSSSHER-QKKRKRSSKKDKHRRSK 260
>At1g32750 unknown protein
Length = 1919
Score = 33.9 bits (76), Expect = 0.50
Identities = 42/188 (22%), Positives = 78/188 (41%), Gaps = 22/188 (11%)
Query: 503 EKISSSAYRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKL 562
EK R +S A D D+++S+ E + +D D + ++ G G SNI K
Sbjct: 1186 EKCQEIWDRQLLSLSAFDGDENESENEANSDLDSFAGDLENLLDAEEGGEGE-ESNISKN 1244
Query: 563 RRHNSRKA-----KHNKLKVYTEIDESHAGEVWLLGLMDSEYSDLKIEEKLNALAALTG- 616
+ + K + ++++ EI++ L L+ + K ++K+ + G
Sbjct: 1245 DKLDGVKGLKMRRRPSQVETDEEIEDEATEYAELCRLLMQDEDQKKKKKKMKGVGEGMGS 1304
Query: 617 --------LLSSGSSIRM-----KDPVKVTADCSSSIQLRGSGAKIKRSVNPIEQMQCTK 663
L SG +R K P+ + D +S + S K R+V+ I + K
Sbjct: 1305 YPPPRPNIALQSGEPVRKANAMDKKPIAIQPD--ASFLVNESTIKDNRNVDSIIKTPKGK 1362
Query: 664 EVHMNSHA 671
+V NS++
Sbjct: 1363 QVKENSNS 1370
>At3g61170 putative protein
Length = 783
Score = 33.5 bits (75), Expect = 0.66
Identities = 28/101 (27%), Positives = 48/101 (46%), Gaps = 17/101 (16%)
Query: 553 GSIHSNIRKLRRHNSRKAKHNKLKVYTEIDES---------HAGEVWLLGLMDSEYSDLK 603
G +HS + + RRH +++Y+++DE A + L +D E +L
Sbjct: 642 GKVHSFMSEDRRHP------RMVEIYSKVDEMMLLIKEAGYFADMSFALHDLDKEGKELG 695
Query: 604 IEEKLNALAALTGLL--SSGSSIRMKDPVKVTADCSSSIQL 642
+ LA GLL SG+ IR+ ++V DC S+++L
Sbjct: 696 LAYHSEKLAVAFGLLVVPSGAPIRIIKNLRVCGDCHSAMKL 736
>At1g71850 hypothetical protein
Length = 470
Score = 33.5 bits (75), Expect = 0.66
Identities = 18/55 (32%), Positives = 27/55 (48%), Gaps = 2/55 (3%)
Query: 506 SSSAYRLRMSTVAKDDD--DSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSN 558
S + Y M+TV KD+ S S ED G V+ E+ D+D + DD + +
Sbjct: 351 SRNRYMKLMNTVKKDNKAVSSSSKKEDKGKVEGEVCDTDAKAENDDISGSDVEDD 405
>At1g63980 unknown protein
Length = 391
Score = 33.5 bits (75), Expect = 0.66
Identities = 21/74 (28%), Positives = 33/74 (44%), Gaps = 2/74 (2%)
Query: 503 EKISSSAYRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKL 562
+KI+ Y + ++ DDD D +D +DE D D + DD I S++
Sbjct: 249 KKIAGVRYEGKKTSFDNSDDD--DDDDDDDDEEDEEEDEDESEADDDDKDSVIESSLPAK 306
Query: 563 RRHNSRKAKHNKLK 576
R+H+ KLK
Sbjct: 307 RKHDEIIEPKIKLK 320
>At4g31630 putative protein
Length = 512
Score = 33.1 bits (74), Expect = 0.86
Identities = 20/58 (34%), Positives = 25/58 (42%)
Query: 514 MSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKLRRHNSRKAK 571
+S+ DDDD + D DD +D D S DDF S I S R + S K
Sbjct: 110 ISSSTSDDDDDERTVFDDDEDDDVGDDDDNSISEDDFCSKKISSKKRARKETESSSDK 167
>At3g25100 Cdc45-like protein
Length = 596
Score = 33.1 bits (74), Expect = 0.86
Identities = 37/155 (23%), Positives = 68/155 (43%), Gaps = 28/155 (18%)
Query: 504 KISSSAYRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKLR 563
K+++ +++LR+ ++ D+ + D E+ DD+ +D D S G G +KL+
Sbjct: 154 KLANESFQLRVEDAGEESDEEEEDEEEDEEDDDD-DDGDRPSKRRKMGDGV--KVFKKLK 210
Query: 564 RHNSRKAK-HNKLKVYTEIDESHAGE------VWLL-----------GLMDSEY--SDLK 603
R + H K + SH +WL L D Y + ++
Sbjct: 211 RDYYKMGTFHGKPSGCLLFELSHMLRKNTNELLWLACVSLTDQFVHERLTDERYQAAVME 270
Query: 604 IEEKLNALAALTGLLSSGSSIRMKDPVKVTA-DCS 637
+E+ +N+ +G + +S+ +KD KV A DCS
Sbjct: 271 LEQHINS----SGNIDKITSVTLKDGTKVRAPDCS 301
>At3g16750 hypothetical protein
Length = 194
Score = 33.1 bits (74), Expect = 0.86
Identities = 15/28 (53%), Positives = 18/28 (63%)
Query: 522 DDSQSDTEDSGSVDDELNDSDTCSSGDD 549
DD S +D GS DDE + SD S+GDD
Sbjct: 158 DDDGSSNDDDGSSDDESHSSDDESAGDD 185
>At1g58220 MYB-family transcription factor-like protein
Length = 834
Score = 33.1 bits (74), Expect = 0.86
Identities = 33/125 (26%), Positives = 55/125 (43%), Gaps = 12/125 (9%)
Query: 144 GKGLATNPNCNMKAPVKRHGMGKGLAANPNSNMKAPVKRHGMGKGLMTIWRATNHD--AR 201
G GLA++ AP KR K +A + + A VKRHG G + + A
Sbjct: 182 GNGLASS-----LAPRKRR---KKWSAEEDEELIAAVKRHGEGSWALISKEEFEGERTAS 233
Query: 202 DLPISFGSVDKDVHLTSNTKTPISVNRSQKAVTTNGESNQYATQNQLPIEKCELALDSSI 261
L +G++ + TSNT T + R++ + N + A N+LP +K + + +
Sbjct: 234 QLSQRWGAIRRRTD-TSNTSTQTGLQRTEAQMAAN-RALSLAVGNRLPSKKLAVGMTPML 291
Query: 262 SDAGV 266
S +
Sbjct: 292 SSGTI 296
>At4g31520 putative protein
Length = 698
Score = 32.7 bits (73), Expect = 1.1
Identities = 53/223 (23%), Positives = 91/223 (40%), Gaps = 38/223 (17%)
Query: 452 LFKILSERGNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDITL--FEKISSSA 509
L + +RG G +A+ + E+N+ S ++ L+ + ++ L + I
Sbjct: 406 LLLVKKDRGRPG---GPIARPKKYGEVNVFSNVPNVD-LLQESDDDEVALPGSDDIEQEL 461
Query: 510 YRLRMSTVAKDDDDSQSDTED----SGSVDDELNDSDTCSSG----DDFGSGSIHSNIRK 561
+ +D D ++TED SG ++E NDSD + +D G S+ +
Sbjct: 462 ITEDEAEEDSNDGDDMNNTEDDTLVSGDEEEEKNDSDEAETDWENEEDEGEASVEGS--- 518
Query: 562 LRRHNSRKAKHNKLKVYTEIDESHAGEVWLLGLMDSEYSDLK----IEEKLNALAALTGL 617
N KAK K K+ + D S L D+ LK E + + A G+
Sbjct: 519 ---GNREKAKGKKRKL-VDFDAS-------LLAADTSLRALKRCAEAEREQTSFAERDGI 567
Query: 618 LSSGSSIRMKDPVKVTADCSSSIQLRG-----SGAKIKRSVNP 655
LS+ ++K+ VK D ++ +G S K+ VNP
Sbjct: 568 LSNEDFRKIKE-VKGKKDAKLALARKGLKVPDSDKLSKKLVNP 609
>At3g48830 putative protein
Length = 881
Score = 32.7 bits (73), Expect = 1.1
Identities = 25/79 (31%), Positives = 40/79 (49%), Gaps = 5/79 (6%)
Query: 225 SVNRSQKAVTTNGESNQYATQNQLPI--EKCELALDSSISDAGVDQISMLIDDEELELRE 282
S N ++KA+ ++ +Y N+L + CE D + Q S+ I++EE +
Sbjct: 613 SANEAKKALDM--KNGEYLCGNKLILGMASCENIAQPKYEDY-IQQESLPIEEEETPPKI 669
Query: 283 IQEGSNLLICSDQLAANGM 301
+Q G+N L CS Q AA M
Sbjct: 670 VQSGANTLGCSIQAAAFSM 688
>At4g02740 hypothetical protein
Length = 645
Score = 32.3 bits (72), Expect = 1.5
Identities = 17/58 (29%), Positives = 27/58 (46%), Gaps = 6/58 (10%)
Query: 502 FEKISSSAY------RLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSG 553
F K S Y LR+ + + +DD + +DS D+++D D + DD G G
Sbjct: 311 FGKFSGKRYGVCDEEELRLIEMMEAEDDEVDEEDDSDDDTDDVSDEDESENDDDMGMG 368
>At3g42670 putative protein
Length = 1256
Score = 32.3 bits (72), Expect = 1.5
Identities = 13/29 (44%), Positives = 17/29 (57%)
Query: 510 YRLRMSTVAKDDDDSQSDTEDSGSVDDEL 538
YR + V+ DDDD + D ED DD+L
Sbjct: 302 YRYNIWNVSSDDDDEEEDCEDDKDTDDDL 330
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.132 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,535,554
Number of Sequences: 26719
Number of extensions: 1100790
Number of successful extensions: 4324
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 27
Number of HSP's successfully gapped in prelim test: 34
Number of HSP's that attempted gapping in prelim test: 4003
Number of HSP's gapped (non-prelim): 215
length of query: 1060
length of database: 11,318,596
effective HSP length: 110
effective length of query: 950
effective length of database: 8,379,506
effective search space: 7960530700
effective search space used: 7960530700
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 65 (29.6 bits)
Medicago: description of AC140023.16


