

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139709.6 + phase: 0
(138 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
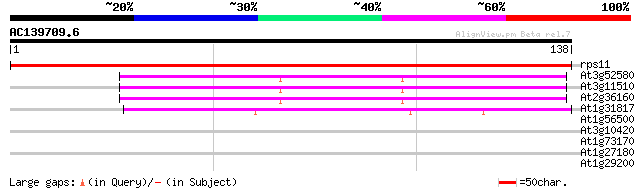
rps11 -chloroplast genome- ribosomal protein S11 234 8e-63
At3g52580 ribosomal protein S14 like protein 67 4e-12
At3g11510 40S ribosomal protein s14 like 66 5e-12
At2g36160 40S ribosomal protein S14 66 5e-12
At1g31817 30S ribosomal like protein S11 60 4e-10
At1g56500 unknown protein 28 1.2
At3g10420 unknown protein 27 4.4
At1g73170 unknown protein 27 4.4
At1g27180 disease resistance protein, putative 26 7.5
At1g29200 hypothetical protein 25 9.8
>rps11 -chloroplast genome- ribosomal protein S11
Length = 138
Score = 234 bits (598), Expect = 8e-63
Identities = 118/138 (85%), Positives = 126/138 (90%)
Query: 1 MAKSIPKIGSRKNGRIGSRKHPRKIPKGVIYVQASFNNTIVTVTDVRGRVISWSSAGSCG 60
MAK I +IGSRKN R GSRK+ R+IPKGVI+VQASFNNTIVTVTDVRGRVISWSSAG+CG
Sbjct: 1 MAKPILRIGSRKNTRSGSRKNVRRIPKGVIHVQASFNNTIVTVTDVRGRVISWSSAGTCG 60
Query: 61 FKGTRRGTPFAAQTAAANAIRTVVDQGMQRAVVIIKGPGLGRDAALRAIARSGILLRFIR 120
F+GTRRGTPFAAQTAA NAIR VVDQGMQRA V IKGPGLGRDAALRAI RSGILL F+R
Sbjct: 61 FRGTRRGTPFAAQTAAGNAIRAVVDQGMQRAEVRIKGPGLGRDAALRAIRRSGILLSFVR 120
Query: 121 DVTPIPHNGCRAPKKRRV 138
DVTP+PHNGCR PKKRRV
Sbjct: 121 DVTPMPHNGCRPPKKRRV 138
>At3g52580 ribosomal protein S14 like protein
Length = 150
Score = 66.6 bits (161), Expect = 4e-12
Identities = 45/119 (37%), Positives = 62/119 (51%), Gaps = 9/119 (7%)
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV++V ASFN+T + VTD+ GR G K R +P+AA AA + + +
Sbjct: 28 GVVHVFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVAQRCKEL 87
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ V + K PG G +ALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 88 GITAIHVKLRATGGNKTKTPGPGAQSALRALARSGMKIGRIEDVTPIPTDSTRRKGGRR 146
>At3g11510 40S ribosomal protein s14 like
Length = 150
Score = 66.2 bits (160), Expect = 5e-12
Identities = 44/119 (36%), Positives = 62/119 (51%), Gaps = 9/119 (7%)
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV+++ ASFN+T + VTD+ GR G K R +P+AA AA + + +
Sbjct: 28 GVVHIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVAQRCKEL 87
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ V + K PG G +ALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 88 GITAMHVKLRATGGNKTKTPGPGAQSALRALARSGMKIGRIEDVTPIPTDSTRRKGGRR 146
>At2g36160 40S ribosomal protein S14
Length = 150
Score = 66.2 bits (160), Expect = 5e-12
Identities = 44/119 (36%), Positives = 62/119 (51%), Gaps = 9/119 (7%)
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV+++ ASFN+T + VTD+ GR G K R +P+AA AA + + +
Sbjct: 28 GVVHIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVAQRCKEL 87
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ V + K PG G +ALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 88 GITAMHVKLRATGGNKTKTPGPGAQSALRALARSGMKIGRIEDVTPIPTDSTRRKGGRR 146
>At1g31817 30S ribosomal like protein S11
Length = 314
Score = 60.1 bits (144), Expect = 4e-10
Identities = 41/120 (34%), Positives = 58/120 (48%), Gaps = 10/120 (8%)
Query: 29 VIYVQASFNNTIVTVTDVRGRVISWSSAGSC-GFKGTRRGTPFAAQTAAANAIRTVVDQG 87
+I+++ NNT VTVTD +G V +++GS KG R+ T + A A N R G
Sbjct: 195 IIHIKMLRNNTFVTVTDSKGNVKCKATSGSLPDLKGGRKMTNYTADATAENIGRRAKAMG 254
Query: 88 MQRAVVIIKG-PGLGRDAALRAIARSGIL--------LRFIRDVTPIPHNGCRAPKKRRV 138
++ VV + G G+ R G + +I D T HNGCR P+KRRV
Sbjct: 255 LKSVVVKVNGFTHFGKKKKAIIAFRDGFTNSRSDQNPIVYIEDTTRKAHNGCRLPRKRRV 314
>At1g56500 unknown protein
Length = 1055
Score = 28.5 bits (62), Expect = 1.2
Identities = 13/35 (37%), Positives = 20/35 (57%)
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFK 62
G IY+ S+N+ I + V RV++ + G GFK
Sbjct: 814 GQIYLTDSYNHKIKKLDPVTKRVVTLAGTGKAGFK 848
>At3g10420 unknown protein
Length = 684
Score = 26.6 bits (57), Expect = 4.4
Identities = 12/38 (31%), Positives = 24/38 (62%), Gaps = 2/38 (5%)
Query: 74 TAAANAIRTVVDQGMQRAVVIIKGPGLGRDAALRAIAR 111
+ +A IR +++ G ++++I PG+G+ +R IAR
Sbjct: 200 SGSAEIIRDLIEGG--GSILVIGSPGVGKTTLIREIAR 235
>At1g73170 unknown protein
Length = 666
Score = 26.6 bits (57), Expect = 4.4
Identities = 12/36 (33%), Positives = 23/36 (63%), Gaps = 2/36 (5%)
Query: 76 AANAIRTVVDQGMQRAVVIIKGPGLGRDAALRAIAR 111
+AN +R +V G ++++I PG+G+ +R +AR
Sbjct: 186 SANLLRDLVQDG--NSLLLIGPPGVGKTTMIREVAR 219
>At1g27180 disease resistance protein, putative
Length = 1556
Score = 25.8 bits (55), Expect = 7.5
Identities = 17/48 (35%), Positives = 25/48 (51%), Gaps = 1/48 (2%)
Query: 84 VDQGMQRAVVIIKGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCR 131
+D + V I+KG GL +AALR + + LL + D T H+ R
Sbjct: 622 MDITKEEVVDILKGCGLNAEAALRVLIQKS-LLTILTDDTLWMHDQIR 668
>At1g29200 hypothetical protein
Length = 698
Score = 25.4 bits (54), Expect = 9.8
Identities = 11/37 (29%), Positives = 18/37 (47%)
Query: 59 CGFKGTRRGTPFAAQTAAANAIRTVVDQGMQRAVVII 95
CG T+R PF + A+ A + D+G + V +
Sbjct: 18 CGSFPTKRKIPFTSHAVASAARKLEADEGYDKKVATV 54
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.323 0.138 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,851,428
Number of Sequences: 26719
Number of extensions: 99561
Number of successful extensions: 204
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 192
Number of HSP's gapped (non-prelim): 10
length of query: 138
length of database: 11,318,596
effective HSP length: 89
effective length of query: 49
effective length of database: 8,940,605
effective search space: 438089645
effective search space used: 438089645
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 54 (25.4 bits)
Medicago: description of AC139709.6


