

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139344.4 - phase: 0 /pseudo
(826 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
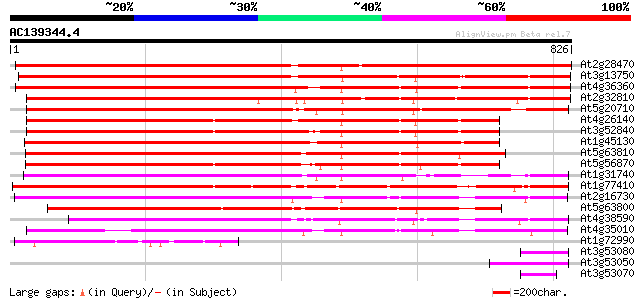
Score E
Sequences producing significant alignments: (bits) Value
At2g28470 beta-galactosidase like protein 1214 0.0
At3g13750 galactosidase, putative 907 0.0
At4g36360 beta-galactosidase like protein 904 0.0
At2g32810 putative beta-galactosidase (BGAL9) 882 0.0
At5g20710 beta-galactosidase like protein 813 0.0
At4g26140 putative beta-galactosidase 806 0.0
At3g52840 beta-galactosidase precursor - like protein 800 0.0
At1g45130 beta-galactosidase like protein 798 0.0
At5g63810 beta-galactosidase (emb|CAB64746.1) 780 0.0
At5g56870 beta-galactosidase (emb|CAB64740.1) 777 0.0
At1g31740 putative protein 723 0.0
At1g77410 unknown protein 719 0.0
At2g16730 putative beta-galactosidase 646 0.0
At5g63800 beta-galactosidase like protein 610 e-174
At4g38590 galactosidase like protein 540 e-153
At4g35010 beta-galactosidase - like protein 536 e-152
At1g72990 unknown protein 161 2e-39
At3g53080 unknown protein 66 9e-11
At3g53050 putative protein 52 1e-06
At3g53070 putative protein 44 4e-04
>At2g28470 beta-galactosidase like protein
Length = 839
Score = 1214 bits (3142), Expect = 0.0
Identities = 591/830 (71%), Positives = 671/830 (80%), Gaps = 24/830 (2%)
Query: 9 VLLWFLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGG 68
++L + V V A+ +NVTYDHRALVIDGKR+VL+SGSIHYPRSTP+MWP+LIQKSKDGG
Sbjct: 9 MILLLILVIVVAATAANVTYDHRALVIDGKRKVLISGSIHYPRSTPEMWPELIQKSKDGG 68
Query: 69 IDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFP 128
+DVIETYVFW+ HEP + +YNFEGR DLV FVK A AGLYVHLRIGPYVCAEWNYGGFP
Sbjct: 69 LDVIETYVFWSGHEPEKNKYNFEGRYDLVKFVKLAAKAGLYVHLRIGPYVCAEWNYGGFP 128
Query: 129 LWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDT 188
+WLHF+ GIKFRT+NEPFK EM+RFT KIVD+MKQE LYASQGGPIILSQIENEYGNID+
Sbjct: 129 VWLHFVPGIKFRTDNEPFKEEMQRFTTKIVDLMKQEKLYASQGGPIILSQIENEYGNIDS 188
Query: 189 HDARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKM 248
AAKSYI W+ASMA SLDTGVPW MCQQ +APDP+INTCN FYCDQFTPNS+NKPKM
Sbjct: 189 AYGAAAKSYIKWSASMALSLDTGVPWNMCQQTDAPDPMINTCNGFYCDQFTPNSNNKPKM 248
Query: 249 WTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFIS 308
WTENWSGWFL FG PYRPVEDLAFAVARF+QRGGTFQNYYMYHGGTNF RT+GGP IS
Sbjct: 249 WTENWSGWFLGFGDPSPYRPVEDLAFAVARFYQRGGTFQNYYMYHGGTNFDRTSGGPLIS 308
Query: 309 TSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKT-G 367
TSYDYDAPIDEYG +RQPKWGHL+DLHKAIKLCE+ALIA+DPTITS G NLE AVYKT
Sbjct: 309 TSYDYDAPIDEYGLLRQPKWGHLRDLHKAIKLCEDALIATDPTITSLGSNLEAAVYKTES 368
Query: 368 AVCSAFLANIG-MSDATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATE 426
C+AFLAN+ SDATVTFNG SY+LP WSVSILPDCKNV NTAKV
Sbjct: 369 GSCAAFLANVDTKSDATVTFNGKSYNLPAWSVSILPDCKNVAFNTAKVK---------FN 419
Query: 427 SLKEKVDSLDSSSSG--WSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVY 484
S+ + D S+ G WS+I EP+GIS DAF K GLLEQINTTAD+SDYLWYSL
Sbjct: 420 SISKTPDGGSSAELGSQWSYIKEPIGISKADAFLKPGLLEQINTTADKSDYLWYSLRTDI 479
Query: 485 ED-----NAGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTI 539
+ + G + VLHIESLG ++AF+NGKLAGS G K+++DIPI LVTG NTI
Sbjct: 480 KGDETFLDEGSKAVLHIESLGQVVYAFINGKLAGS---GHGKQKISLDIPINLVTGTNTI 536
Query: 540 DLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVGLSS 599
DLLS+TVGL NYGAF+D VGAGITGPV LK K GSS+DL SQQWTYQVGL+GE GL++
Sbjct: 537 DLLSVTVGLANYGAFFDLVGAGITGPVTLKSAKGGSSIDLASQQWTYQVGLKGEDTGLAT 596
Query: 600 GNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTY 659
+ +W S+S LP QPL WYKT F APSGS PVAIDFTG GKG AWVNGQSIGRYWPT
Sbjct: 597 VDSSEWVSKSPLPTKQPLIWYKTTFDAPSGSEPVAIDFTGTGKGIAWVNGQSIGRYWPTS 656
Query: 660 ISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTK 719
I+ N GCT+SC+YRG+Y A+KCLKNCGKPSQTLYHVPR+WLKP N VLFEE GGDPT+
Sbjct: 657 IAGNGGCTESCDYRGSYRANKCLKNCGKPSQTLYHVPRSWLKPSGNILVLFEEMGGDPTQ 716
Query: 720 ISFGTKQIES-VCSHVTESHPPPVDTWNSNAE--SERKVGPVLSLECPYPNQAISSIKFA 776
ISF TKQ S +C V++SHPPPVDTW S+++ + + PVLSL+CP Q I SIKFA
Sbjct: 717 ISFATKQTGSNLCLTVSQSHPPPVDTWTSDSKISNRNRTRPVLSLKCPISTQVIFSIKFA 776
Query: 777 SFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTFGNPCR 826
SFGTP+GTCG++ G C+S+R+LS+VQKACIG SCN+ VS FG PCR
Sbjct: 777 SFGTPKGTCGSFTQGHCNSSRSLSLVQKACIGLRSCNVEVSTRVFGEPCR 826
>At3g13750 galactosidase, putative
Length = 847
Score = 907 bits (2343), Expect = 0.0
Identities = 446/825 (54%), Positives = 569/825 (68%), Gaps = 24/825 (2%)
Query: 13 FLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVI 72
FL ++ S +V+YD RA+ I+GKRR+L+SGSIHYPRSTP+MWPDLI+K+K+GG+DVI
Sbjct: 21 FLLGFLVCSVSGSVSYDSRAITINGKRRILISGSIHYPRSTPEMWPDLIRKAKEGGLDVI 80
Query: 73 ETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLH 132
+TYVFWN HEP G+Y FEG DLV FVK V +GLY+HLRIGPYVCAEWN+GGFP+WL
Sbjct: 81 QTYVFWNGHEPSPGKYYFEGNYDLVKFVKLVQQSGLYLHLRIGPYVCAEWNFGGFPVWLK 140
Query: 133 FIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDAR 192
+I GI FRT+N PFKA+M+RFT KIV+MMK E L+ SQGGPIILSQIENEYG ++
Sbjct: 141 YIPGISFRTDNGPFKAQMQRFTTKIVNMMKAERLFESQGGPIILSQIENEYGPMEYELGA 200
Query: 193 AAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTEN 252
+SY +WAA MA L TGVPW+MC+Q +APDPIIN CN FYCD F+PN KPKMWTE
Sbjct: 201 PGRSYTNWAAKMAVGLGTGVPWVMCKQDDAPDPIINACNGFYCDYFSPNKAYKPKMWTEA 260
Query: 253 WSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYD 312
W+GWF FGG VPYRP ED+AF+VARF Q+GG+F NYYMYHGGTNFGRT GGPFI+TSYD
Sbjct: 261 WTGWFTKFGGPVPYRPAEDMAFSVARFIQKGGSFINYYMYHGGTNFGRTAGGPFIATSYD 320
Query: 313 YDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKT-GAVCS 371
YDAP+DEYG RQPKWGHLKDLH+AIKLCE AL++ +PT G E VYK+ CS
Sbjct: 321 YDAPLDEYGLERQPKWGHLKDLHRAIKLCEPALVSGEPTRMPLGNYQEAHVYKSKSGACS 380
Query: 372 AFLANIG-MSDATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKE 430
AFLAN S A V+F N Y+LP WS+SILPDCKN V NTA+V A S +
Sbjct: 381 AFLANYNPKSYAKVSFGNNHYNLPPWSISILPDCKNTVYNTARVG--------AQTSRMK 432
Query: 431 KVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVYEDNA-- 488
V W +E ++FT GL+EQINTT D SDYLWY + + N
Sbjct: 433 MVRVPVHGGLSWQAYNEDPSTYIDESFTMVGLVEQINTTRDTSDYLWYMTDVKVDANEGF 492
Query: 489 ---GDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLT 545
GD P L + S GHA+H F+NG+L+GS GS + K+ + L G N I +LS+
Sbjct: 493 LRNGDLPTLTVLSAGHAMHVFINGQLSGSAYGSLDSPKLTFRKGVNLRAGFNKIAILSIA 552
Query: 546 VGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFV---GLSSGNV 602
VGL N G ++T AG+ GPV L GL NG DL+ Q+WTY+VGL+GE + LS +
Sbjct: 553 VGLPNVGPHFETWNAGVLGPVSLNGL-NGGRRDLSWQKWTYKVGLKGESLSLHSLSGSSS 611
Query: 603 GQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISP 662
+W + + QPLTWYKT F AP+G +P+A+D MGKG+ W+NGQS+GR+WP Y +
Sbjct: 612 VEWAEGAFVAQKQPLTWYKTTFSAPAGDSPLAVDMGSMGKGQIWINGQSLGRHWPAYKAV 671
Query: 663 NSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTKISF 722
S C++ C+Y GT+ KCL+NCG+ SQ YHVPR+WLKP N V+FEE GGDP I+
Sbjct: 672 GS-CSE-CSYTGTFREDKCLRNCGEASQRWYHVPRSWLKPSGNLLVVFEEWGGDPNGITL 729
Query: 723 GTKQIESVCSHVTESHPPPVD-TWNSNAESERKVGPVLSLECPYPNQAISSIKFASFGTP 781
++++SVC+ + E V+ +++ + + + P L+C P Q I+++KFASFGTP
Sbjct: 730 VRREVDSVCADIYEWQSTLVNYQLHASGKVNKPLHPKAHLQCG-PGQKITTVKFASFGTP 788
Query: 782 RGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTF-GNPC 825
GTCG+Y GSC ++ + K C+G + C++ V+ F G+PC
Sbjct: 789 EGTCGSYRQGSCHAHHSYDAFNKLCVGQNWCSVTVAPEMFGGDPC 833
>At4g36360 beta-galactosidase like protein
Length = 856
Score = 904 bits (2335), Expect = 0.0
Identities = 454/842 (53%), Positives = 574/842 (67%), Gaps = 47/842 (5%)
Query: 9 VLLWF-LGVYV-PASFCS-NVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSK 65
++LWF LG + F VTYD +AL+I+G+RR+L SGSIHYPRSTP MW DLIQK+K
Sbjct: 13 LILWFCLGFLILGVGFVQCGVTYDRKALLINGQRRILFSGSIHYPRSTPDMWEDLIQKAK 72
Query: 66 DGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYG 125
DGGIDVIETYVFWNLHEP G+Y+FEGR DLV FVK + AGLY HLRIGPYVCAEWN+G
Sbjct: 73 DGGIDVIETYVFWNLHEPSPGKYDFEGRNDLVRFVKTIHKAGLYAHLRIGPYVCAEWNFG 132
Query: 126 GFPLWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGN 185
GFP+WL ++ GI FRT+NEPFK MK FT +IV++MK ENL+ SQGGPIILSQIENEYG
Sbjct: 133 GFPVWLKYVPGISFRTDNEPFKRAMKGFTERIVELMKSENLFESQGGPIILSQIENEYGR 192
Query: 186 IDTHDARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNK 245
+Y+ WAA MA + +TGVPW+MC++ +APDP+INTCN FYCD F PN K
Sbjct: 193 QGQLLGAEGHNYMTWAAKMAIATETGVPWVMCKEDDAPDPVINTCNGFYCDSFAPNKPYK 252
Query: 246 PKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGP 305
P +WTE WSGWF FGG + +RPV+DLAF VARF Q+GG+F NYYMYHGGTNFGRT GGP
Sbjct: 253 PLIWTEAWSGWFTEFGGPMHHRPVQDLAFGVARFIQKGGSFVNYYMYHGGTNFGRTAGGP 312
Query: 306 FISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYK 365
F++TSYDYDAPIDEYG IRQPK+GHLK+LH+AIK+CE+AL+++DP +TS G + VY
Sbjct: 313 FVTTSYDYDAPIDEYGLIRQPKYGHLKELHRAIKMCEKALVSADPVVTSIGNKQQAHVYS 372
Query: 366 T-GAVCSAFLANIGM-SDATVTFNGNSYHLPGWSVSILPDCKNVVLNTAK--VNTASM-- 419
CSAFLAN S A V FN Y+LP WS+SILPDC+N V NTAK V T+ M
Sbjct: 373 AESGDCSAFLANYDTESAARVLFNNVHYNLPPWSISILPDCRNAVFNTAKVGVQTSQMEM 432
Query: 420 ----ISSFATESLKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDY 475
+F ES E + SLD SS+ FT GLLEQIN T D SDY
Sbjct: 433 LPTDTKNFQWESYLEDLSSLDDSST----------------FTTHGLLEQINVTRDTSDY 476
Query: 476 LWYSLSIVYED-----NAGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPI 530
LWY S+ D + G+ P L I+S GHA+H FVNG+L+GS G+ N + I
Sbjct: 477 LWYMTSVDIGDSESFLHGGELPTLIIQSTGHAVHIFVNGQLSGSAFGTRQNRRFTYQGKI 536
Query: 531 TLVTGKNTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGL 590
L +G N I LLS+ VGL N G +++ GI GPV L GL G +DL+ Q+WTYQVGL
Sbjct: 537 NLHSGTNRIALLSVAVGLPNVGGHFESWNTGILGPVALHGLSQG-KMDLSWQKWTYQVGL 595
Query: 591 QGEFVGL----SSGNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAW 646
+GE + L ++ ++G ++ + QPLTW+KT F AP G+ P+A+D GMGKG+ W
Sbjct: 596 KGEAMNLAFPTNTPSIGWMDASLTVQKPQPLTWHKTYFDAPEGNEPLALDMEGMGKGQIW 655
Query: 647 VNGQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNT 706
VNG+SIGRYW + +G C+Y GTY +KC CG+P+Q YHVPRAWLKP N
Sbjct: 656 VNGESIGRYWTAFA---TGDCSHCSYTGTYKPNKCQTGCGQPTQRWYHVPRAWLKPSQNL 712
Query: 707 FVLFEESGGDPTKISFGTKQIESVCSHVTESHPPPVDTW--NSNAESERKVGPVLSLECP 764
V+FEE GG+P+ +S + + VC+ V+E H P + W S + + P + L+C
Sbjct: 713 LVIFEELGGNPSTVSLVKRSVSGVCAEVSEYH-PNIKNWQIESYGKGQTFHRPKVHLKCS 771
Query: 765 YPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTFG-N 823
P QAI+SIKFASFGTP GTCG+Y G C + + +I+++ C+G + C + +S + FG +
Sbjct: 772 -PGQAIASIKFASFGTPLGTCGSYQQGECHAATSYAILERKCVGKARCAVTISNSNFGKD 830
Query: 824 PC 825
PC
Sbjct: 831 PC 832
>At2g32810 putative beta-galactosidase (BGAL9)
Length = 887
Score = 882 bits (2279), Expect = 0.0
Identities = 443/834 (53%), Positives = 559/834 (66%), Gaps = 40/834 (4%)
Query: 25 NVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPV 84
NV+YDHRAL+I GKRR+L+S IHYPR+TP+MW DLI KSK+GG DV++TYVFWN HEPV
Sbjct: 37 NVSYDHRALIIAGKRRMLVSAGIHYPRATPEMWSDLIAKSKEGGADVVQTYVFWNGHEPV 96
Query: 85 RGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNE 144
+GQYNFEGR DLV FVK + ++GLY+HLRIGPYVCAEWN+GGFP+WL I GI+FRT+NE
Sbjct: 97 KGQYNFEGRYDLVKFVKLIGSSGLYLHLRIGPYVCAEWNFGGFPVWLRDIPGIEFRTDNE 156
Query: 145 PFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAASM 204
PFK EM++F KIVD+M++ L+ QGGPII+ QIENEYG+++ + K Y+ WAASM
Sbjct: 157 PFKKEMQKFVTKIVDLMREAKLFCWQGGPIIMLQIENEYGDVEKSYGQKGKDYVKWAASM 216
Query: 205 ATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAFGGAV 264
A L GVPW+MC+Q +AP+ II+ CN +YCD F PNS KP +WTE+W GW+ +GG++
Sbjct: 217 ALGLGAGVPWVMCKQTDAPENIIDACNGYYCDGFKPNSRTKPVLWTEDWDGWYTKWGGSL 276
Query: 265 PYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDIR 324
P+RP EDLAFAVARF+QRGG+FQNYYMY GGTNFGRT+GGPF TSYDYDAP+DEYG
Sbjct: 277 PHRPAEDLAFAVARFYQRGGSFQNYYMYFGGTNFGRTSGGPFYITSYDYDAPLDEYGLRS 336
Query: 325 QPKWGHLKDLHKAIKLCEEALIASD-PTITSPGPNLETAVY-----KTGAVCSAFLANIG 378
+PKWGHLKDLH AIKLCE AL+A+D P G E +Y G VC+AFLANI
Sbjct: 337 EPKWGHLKDLHAAIKLCEPALVAADAPQYRKLGSKQEAHIYHGDGETGGKVCAAFLANID 396
Query: 379 -MSDATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMIS-------SFATESLKE 430
A V FNG SY LP WSVSILPDC++V NTAKV + + S + S+ +
Sbjct: 397 EHKSAHVKFNGQSYTLPPWSVSILPDCRHVAFNTAKVGAQTSVKTVESARPSLGSMSILQ 456
Query: 431 KV---DSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSI-VYED 486
KV D++ S W + EP+GI + FT GLLE +N T DRSDYLW+ I V ED
Sbjct: 457 KVVRQDNVSYISKSWMALKEPIGIWGENNFTFQGLLEHLNVTKDRSDYLWHKTRISVSED 516
Query: 487 NA------GDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTID 540
+ G + I+S+ L FVN +LAGS G V P+ + G N +
Sbjct: 517 DISFWKKNGPNSTVSIDSMRDVLRVFVNKQLAGSIVGH----WVKAVQPVRFIQGNNDLL 572
Query: 541 LLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGE---FVGL 597
LL+ TVGLQNYGAF + GAG G L G KNG +DL+ WTYQVGL+GE +
Sbjct: 573 LLTQTVGLQNYGAFLEKDGAGFRGKAKLTGFKNG-DLDLSKSSWTYQVGLKGEADKIYTV 631
Query: 598 SSGNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWP 657
+W++ + WYKT F P+G++PV ++ MG+G+AWVNGQ IGRYW
Sbjct: 632 EHNEKAEWSTLETDASPSIFMWYKTYFDPPAGTDPVVLNLESMGRGQAWVNGQHIGRYW- 690
Query: 658 TYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDP 717
IS GC +C+YRG Y++ KC NCGKP+QT YHVPR+WLKP SN VLFEE+GG+P
Sbjct: 691 NIISQKDGCDRTCDYRGAYNSDKCTTNCGKPTQTRYHVPRSWLKPSSNLLVLFEETGGNP 750
Query: 718 TKISFGTKQIESVCSHVTESHPPPVDTWN-----SNAESERKVGPVLSLECPYPNQAISS 772
KIS T +C V+ESH PP+ W+ + S V P + L C ISS
Sbjct: 751 FKISVKTVTAGILCGQVSESHYPPLRKWSTPDYINGTMSINSVAPEVHLHCE-DGHVISS 809
Query: 773 IKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTF-GNPC 825
I+FAS+GTPRG+C ++ G C ++ +LSIV +AC G +SC I VS F +PC
Sbjct: 810 IEFASYGTPRGSCDGFSIGKCHASNSLSIVSEACKGRNSCFIEVSNTAFISDPC 863
>At5g20710 beta-galactosidase like protein
Length = 824
Score = 813 bits (2101), Expect = 0.0
Identities = 402/813 (49%), Positives = 517/813 (63%), Gaps = 46/813 (5%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPVR 85
V++D RA+ I+GKRR+L+SGSIHYPRST MWPDLI K+KDGG+D IETYVFWN HEP R
Sbjct: 26 VSHDERAITINGKRRILLSGSIHYPRSTADMWPDLINKAKDGGLDAIETYVFWNAHEPKR 85
Query: 86 GQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEP 145
+Y+F G D+V F+K + AGLY LRIGPYVCAEWNYGGFP+WLH + +KFRT N
Sbjct: 86 REYDFSGNLDVVRFIKTIQDAGLYSVLRIGPYVCAEWNYGGFPVWLHNMPNMKFRTVNPS 145
Query: 146 FKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAASMA 205
F EM+ FT KIV MMK+E L+ASQGGPIIL+QIENEYGN+ + K+YIDW A+MA
Sbjct: 146 FMNEMQNFTTKIVKMMKEEKLFASQGGPIILAQIENEYGNVISSYGAEGKAYIDWCANMA 205
Query: 206 TSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAFGGAVP 265
SLD GVPW+MCQQ NAP P++ TCN FYCDQ+ P + + PKMWTENW+GWF +GG P
Sbjct: 206 NSLDIGVPWLMCQQPNAPQPMLETCNGFYCDQYEPTNPSTPKMWTENWTGWFKNWGGKHP 265
Query: 266 YRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDIRQ 325
YR EDLAF+VARFFQ GGTFQNYYMYHGGTNFGR GGP+I+TSYDY AP+DE+G++ Q
Sbjct: 266 YRTAEDLAFSVARFFQTGGTFQNYYMYHGGTNFGRVAGGPYITTSYDYHAPLDEFGNLNQ 325
Query: 326 PKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTGAVCSAFLANI-GMSDATV 384
PKWGHLK LH +K E++L + + G +++ +Y T S F+ N+ +DA V
Sbjct: 326 PKWGHLKQLHTVLKSMEKSLTYGNISRIDLGNSIKATIYTTKEGSSCFIGNVNATADALV 385
Query: 385 TFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKEKVDSLDSSSSGWSW 444
F G YH+P WSVS+LPDC NTAKVNT +S TE DS W+W
Sbjct: 386 NFKGKDYHVPAWSVSVLPDCDKEAYNTAKVNTQ---TSIMTE------DSSKPERLEWTW 436
Query: 445 ISEPVG---ISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVYEDNA---GDQPVLHIES 498
E + GL++Q + T D SDYLWY + + L + S
Sbjct: 437 RPESAQKMILKGSGDLIAKGLVDQKDVTNDASDYLWYMTRLHLDKKDPLWSRNMTLRVHS 496
Query: 499 LGHALHAFVNGKLAGSKAGSSGNAKVNVDIPIT-LVTGKNTIDLLSLTVGLQNYGAFYDT 557
H LHA+VNGK G++ G + + LV G N I LLS++VGLQNYG F+++
Sbjct: 497 NAHVLHAYVNGKYVGNQFVKDGKFDYRFERKVNHLVHGTNHISLLSVSVGLQNYGPFFES 556
Query: 558 VGAGITGPVILKGLKNGSSV--DLTSQQWTYQVGLQG---EFVGLSSGNVGQWNSQSNLP 612
GI GPV L G K ++ DL+ QW Y++GL G + + S +W + LP
Sbjct: 557 GPTGINGPVSLVGYKGEETIEKDLSQHQWDYKIGLNGYNDKLFSIKSVGHQKW-ANEKLP 615
Query: 613 ANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSCNY 672
+ LTWYK F AP G PV +D G+GKGEAW+NGQSIGRYWP++ S + GC D C+Y
Sbjct: 616 TGRMLTWYKAKFKAPLGKEPVIVDLNGLGKGEAWINGQSIGRYWPSFNSSDDGCKDECDY 675
Query: 673 RGTYSASKCLKNCGKPSQTLYHVPRAWLKPDS-NTFVLFEESGGDPTKISFGTKQIESVC 731
RG Y + KC CGKP+Q YHVPR++L NT LFEE GG+P+ ++F T + +VC
Sbjct: 676 RGAYGSDKCAFMCGKPTQRWYHVPRSFLNASGHNTITLFEEMGGNPSMVNFKTVVVGTVC 735
Query: 732 SHVTESHPPPVDTWNSNAESERKVGPVLSLECPYPNQAISSIKFASFGTPRGTCGNYNHG 791
+ E + +E N+ IS++KFASFG P G CG++ G
Sbjct: 736 ARAHEHN---------------------KVELSCHNRPISAVKFASFGNPLGHCGSFAVG 774
Query: 792 SCSSNR-ALSIVQKACIGSSSCNIGVSINTFGN 823
+C ++ A V K C+G +C + VS +TFG+
Sbjct: 775 TCQGDKDAAKTVAKECVGKLNCTVNVSSDTFGS 807
>At4g26140 putative beta-galactosidase
Length = 729
Score = 806 bits (2081), Expect = 0.0
Identities = 391/705 (55%), Positives = 494/705 (69%), Gaps = 20/705 (2%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPVR 85
VTYD +A++I+G+RR+L+SGSIHYPRSTP+MWPDLIQK+KDGG+DVI+TYVFWN HEP
Sbjct: 29 VTYDRKAVIINGQRRILLSGSIHYPRSTPEMWPDLIQKAKDGGLDVIQTYVFWNGHEPSP 88
Query: 86 GQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEP 145
GQY FE R DLV F+K V AGLYVHLRIGPYVCAEWN+GGFP+WL ++ G+ FRT+NEP
Sbjct: 89 GQYYFEDRYDLVKFIKVVQQAGLYVHLRIGPYVCAEWNFGGFPVWLKYVPGMVFRTDNEP 148
Query: 146 FKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAASMA 205
FKA M++FT KIV MMK+E L+ +QGGPIILSQIENEYG I+ K+Y W A MA
Sbjct: 149 FKAAMQKFTEKIVRMMKEEKLFETQGGPIILSQIENEYGPIEWEIGAPGKAYTKWVAEMA 208
Query: 206 TSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAFGGAVP 265
L TGVPWIMC+Q +AP+ IINTCN FYC+ F PNSDNKPKMWTENW+GWF FGGAVP
Sbjct: 209 QGLSTGVPWIMCKQDDAPNSIINTCNGFYCENFKPNSDNKPKMWTENWTGWFTEFGGAVP 268
Query: 266 YRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDIRQ 325
YRP ED+A +VARF Q GG+F NYYMYHGGTNF R T G FI+TSYDYDAP+DEYG R+
Sbjct: 269 YRPAEDIALSVARFIQNGGSFINYYMYHGGTNFDR-TAGEFIATSYDYDAPLDEYGLPRE 327
Query: 326 PKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTGAVCSAFLANIGMSDAT-V 384
PK+ HLK LHK IKLCE AL+++DPT+TS G E V+K+ + C+AFL+N S A V
Sbjct: 328 PKYSHLKRLHKVIKLCEPALVSADPTVTSLGDKQEAHVFKSKSSCAAFLSNYNTSSAARV 387
Query: 385 TFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKEKVDSLDSSSSGWSW 444
F G++Y LP WSVSILPDCK NTAKV T S+ K+ ++ S S+
Sbjct: 388 LFGGSTYDLPPWSVSILPDCKTEYYNTAKVQV-------RTSSIHMKMVPTNTPFSWGSY 440
Query: 445 ISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVYEDN----AGDQPVLHIESLG 500
E + F++ GL+EQI+ T D++DY WY I + G+ P+L I S G
Sbjct: 441 NEEIPSANDNGTFSQDGLVEQISITRDKTDYFWYLTDITISPDEKFLTGEDPLLTIGSAG 500
Query: 501 HALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYGAFYDTVGA 560
HALH FVNG+LAG+ GS K+ I L G N + LLS GL N G Y+T
Sbjct: 501 HALHVFVNGQLAGTAYGSLEKPKLTFSQKIKLHAGVNKLALLSTAAGLPNVGVHYETWNT 560
Query: 561 GITGPVILKGLKNGSSVDLTSQQWTY-QVGLQGEFVG---LSSGNVGQWNSQSNLPANQP 616
G+ GPV L G+ +G + D+T +W+Y Q+G +GE + L+ + +W S + QP
Sbjct: 561 GVLGPVTLNGVNSG-TWDMTKWKWSYKQIGTKGEALSVHTLAGSSTVEWKEGSLVAKKQP 619
Query: 617 LTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSCNYRGTY 676
LTWYK+ F +P+G+ P+A+D MGKG+ W+NGQ+IGR+WP Y + G + C+Y GT+
Sbjct: 620 LTWYKSTFDSPTGNEPLALDMNTMGKGQMWINGQNIGRHWPAYTA--RGKCERCSYAGTF 677
Query: 677 SASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTKIS 721
+ KCL NCG+ SQ YHVPR+WLKP +N ++ EE GG+P IS
Sbjct: 678 TEKKCLSNCGEASQRWYHVPRSWLKPTNNLVIVLEEWGGEPNGIS 722
>At3g52840 beta-galactosidase precursor - like protein
Length = 727
Score = 800 bits (2066), Expect = 0.0
Identities = 392/705 (55%), Positives = 490/705 (68%), Gaps = 22/705 (3%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPVR 85
VTYDH+AL+I+G+RR+L+SGSIHYPRSTP+MWPDLI+K+K+GG+DVI+TYVFWN HEP
Sbjct: 29 VTYDHKALIINGQRRILISGSIHYPRSTPEMWPDLIKKAKEGGLDVIQTYVFWNGHEPSP 88
Query: 86 GQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEP 145
G Y F+ R DLV F K V AGLY+ LRIGPYVCAEWN+GGFP+WL ++ G+ FRT+NEP
Sbjct: 89 GNYYFQDRYDLVKFTKLVHQAGLYLDLRIGPYVCAEWNFGGFPVWLKYVPGMVFRTDNEP 148
Query: 146 FKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAASMA 205
FK M++FT KIVDMMK+E L+ +QGGPIILSQIENEYG + A K+Y W A MA
Sbjct: 149 FKIAMQKFTKKIVDMMKEEKLFETQGGPIILSQIENEYGPMQWEMGAAGKAYSKWTAEMA 208
Query: 206 TSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAFGGAVP 265
L TGVPWIM +Q +AP PII+TCN FYC+ F PNSDNKPK+WTENW+GWF FGGA+P
Sbjct: 209 LGLSTGVPWIMSKQEDAPYPIIDTCNGFYCEGFKPNSDNKPKLWTENWTGWFTEFGGAIP 268
Query: 266 YRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDIRQ 325
RPVED+AF+VARF Q GG+F NYYMY+GGTNF R T G FI+TSYDYDAPIDEYG +R+
Sbjct: 269 NRPVEDIAFSVARFIQNGGSFMNYYMYYGGTNFDR-TAGVFIATSYDYDAPIDEYGLLRE 327
Query: 326 PKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTGAVCSAFLANIGMSDAT-V 384
PK+ HLK+LHK IKLCE AL++ DPTITS G E V+K+ C+AFL+N S A V
Sbjct: 328 PKYSHLKELHKVIKLCEPALVSVDPTITSLGDKQEIHVFKSKTSCAAFLSNYDTSSAARV 387
Query: 385 TFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKEKVDSLDSSSSGWSW 444
F G Y LP WSVSILPDCK NTAK+ +++ S K +S + S
Sbjct: 388 MFRGFPYDLPPWSVSILPDCKTEYYNTAKIRAPTILMKMIPTSTKFSWESYNEGSPS--- 444
Query: 445 ISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVYEDN-----AGDQPVLHIESL 499
S G F K GL+EQI+ T D++DY WY I + GD P+L I S
Sbjct: 445 -SNEAG-----TFVKDGLVEQISMTRDKTDYFWYFTDITIGSDESFLKTGDNPLLTIFSA 498
Query: 500 GHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYGAFYDTVG 559
GHALH FVNG LAG+ G+ N+K+ I L G N + LLS VGL N G Y+T
Sbjct: 499 GHALHVFVNGLLAGTSYGALSNSKLTFSQNIKLSVGINKLALLSTAVGLPNAGVHYETWN 558
Query: 560 AGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVG---LSSGNVGQWNSQSNLPANQP 616
GI GPV LKG+ +G + D++ +W+Y++GL+GE + L+ + +W + + QP
Sbjct: 559 TGILGPVTLKGVNSG-TWDMSKWKWSYKIGLRGEAMSLHTLAGSSAVKWWIKGFVVKKQP 617
Query: 617 LTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSCNYRGTY 676
LTWYK++F P G+ P+A+D MGKG+ WVNG +IGR+WP Y + G CNY G Y
Sbjct: 618 LTWYKSSFDTPRGNEPLALDMNTMGKGQVWVNGHNIGRHWPAYTA--RGNCGRCNYAGIY 675
Query: 677 SASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTKIS 721
+ KCL +CG+PSQ YHVPR+WLKP N V+FEE GGDP+ IS
Sbjct: 676 NEKKCLSHCGEPSQRWYHVPRSWLKPFGNLLVIFEEWGGDPSGIS 720
>At1g45130 beta-galactosidase like protein
Length = 732
Score = 798 bits (2061), Expect = 0.0
Identities = 397/711 (55%), Positives = 483/711 (67%), Gaps = 25/711 (3%)
Query: 23 CSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHE 82
CS+VTYD +A+VI+G RR+L+SGSIHYPRSTP+MW DLI+K+KDGG+DVI+TYVFWN HE
Sbjct: 28 CSSVTYDKKAIVINGHRRILLSGSIHYPRSTPEMWEDLIKKAKDGGLDVIDTYVFWNGHE 87
Query: 83 PVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTN 142
P G YNFEGR DLV F+K + GLYVHLRIGPYVCAEWN+GGFP+WL ++ GI FRT+
Sbjct: 88 PSPGTYNFEGRYDLVRFIKTIQEVGLYVHLRIGPYVCAEWNFGGFPVWLKYVDGISFRTD 147
Query: 143 NEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAA 202
N PFK+ M+ FT KIV MMK+ +ASQGGPIILSQIENE+ A SY++WAA
Sbjct: 148 NGPFKSAMQGFTEKIVQMMKEHRFFASQGGPIILSQIENEFEPDLKGLGPAGHSYVNWAA 207
Query: 203 SMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAFGG 262
MA L+TGVPW+MC++ +APDPIINTCN FYCD FTPN KP MWTE WSGWF FGG
Sbjct: 208 KMAVGLNTGVPWVMCKEDDAPDPIINTCNGFYCDYFTPNKPYKPTMWTEAWSGWFTEFGG 267
Query: 263 AVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGD 322
VP RPVEDLAF VARF Q+GG++ NYYMYHGGTNFGRT GGPFI+TSYDYDAPIDEYG
Sbjct: 268 TVPKRPVEDLAFGVARFIQKGGSYINYYMYHGGTNFGRTAGGPFITTSYDYDAPIDEYGL 327
Query: 323 IRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTG-AVCSAFLANIGM-S 380
+++PK+ HLK LH+AIK CE AL++SDP +T G E V+ G C AFL N M +
Sbjct: 328 VQEPKYSHLKQLHQAIKQCEAALVSSDPHVTKLGNYEEAHVFTAGKGSCVAFLTNYHMNA 387
Query: 381 DATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNT-ASMISSFATESLKEKVDSLDSSS 439
A V FN Y LP WS+SILPDC+NVV NTA V S + + S+ V D
Sbjct: 388 PAKVVFNNRHYTLPAWSISILPDCRNVVFNTATVAAKTSHVQMVPSGSILYSVARYDEDI 447
Query: 440 SGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVYEDN-----AGDQPVL 494
+ + T GLLEQ+N T D +DYLWY+ S+ + + G P L
Sbjct: 448 ATY---------GNRGTITARGLLEQVNVTRDTTDYLWYTTSVDIKASESFLRGGKWPTL 498
Query: 495 HIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYGAF 554
++S GHA+H FVNG GS G+ N K + + L G N I LLS+ VGL N G
Sbjct: 499 TVDSAGHAVHVFVNGHFYGSAFGTRENRKFSFSSQVNLRGGANKIALLSVAVGLPNVGPH 558
Query: 555 YDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVGLSS---GNVGQWNSQSNL 611
++T GI G V+L GL G+ DL+ Q+WTYQ GL+GE + L S + W S
Sbjct: 559 FETWATGIVGSVVLHGLDEGNK-DLSWQKWTYQAGLRGESMNLVSPTEDSSVDWIKGSLA 617
Query: 612 PAN-QPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSC 670
N QPLTWYK F AP G+ P+A+D MGKG+AW+NGQSIGRYW + + G SC
Sbjct: 618 KQNKQPLTWYKAYFDAPRGNEPLALDLKSMGKGQAWINGQSIGRYWMAFAKGDCG---SC 674
Query: 671 NYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTKIS 721
NY GTY +KC CG+P+Q YHVPR+WLKP N VLFEE GGD +K+S
Sbjct: 675 NYAGTYRQNKCQSGCGEPTQRWYHVPRSWLKPKGNLLVLFEELGGDISKVS 725
>At5g63810 beta-galactosidase (emb|CAB64746.1)
Length = 741
Score = 780 bits (2014), Expect = 0.0
Identities = 378/720 (52%), Positives = 484/720 (66%), Gaps = 21/720 (2%)
Query: 24 SNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEP 83
+NV+YDHR+L I +R++++S +IHYPRS P MWP L+Q +K+GG + IE+YVFWN HEP
Sbjct: 30 ANVSYDHRSLTIGNRRQLIISAAIHYPRSVPAMWPSLVQTAKEGGCNAIESYVFWNGHEP 89
Query: 84 VRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNN 143
G+Y F GR ++V F+K V AG+++ LRIGP+V AEWNYGG P+WLH++ G FR +N
Sbjct: 90 SPGKYYFGGRYNIVKFIKIVQQAGMHMILRIGPFVAAEWNYGGVPVWLHYVPGTVFRADN 149
Query: 144 EPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAAS 203
EP+K M+ FT IV+++KQE L+A QGGPIILSQ+ENEYG + K Y W+AS
Sbjct: 150 EPWKHYMESFTTYIVNLLKQEKLFAPQGGPIILSQVENEYGYYEKDYGEGGKRYAQWSAS 209
Query: 204 MATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAFGGA 263
MA S + GVPW+MCQQ +AP +I+TCN FYCDQFTPN+ +KPK+WTENW GWF FGG
Sbjct: 210 MAVSQNIGVPWMMCQQWDAPPTVISTCNGFYCDQFTPNTPDKPKIWTENWPGWFKTFGGR 269
Query: 264 VPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDI 323
P+RP ED+A++VARFF +GG+ NYYMYHGGTNFGRT+GGPFI+TSYDY+APIDEYG
Sbjct: 270 DPHRPAEDVAYSVARFFGKGGSVHNYYMYHGGTNFGRTSGGPFITTSYDYEAPIDEYGLP 329
Query: 324 RQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVY-KTGAVCSAFLANI-GMSD 381
R PKWGHLKDLHKAI L E LI+ + + G +LE VY + C+AFL+N+ +D
Sbjct: 330 RLPKWGHLKDLHKAIMLSENLLISGEHQNFTLGHSLEADVYTDSSGTCAAFLSNLDDKND 389
Query: 382 ATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKEKVDSLDSSSSG 441
V F SYHLP WSVSILPDCK V NTAKV + S E LK SS
Sbjct: 390 KAVMFRNTSYHLPAWSVSILPDCKTEVFNTAKVTSKSSKVEMLPEDLK------SSSGLK 443
Query: 442 WSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVYEDN-----AGDQPVLHI 496
W SE GI F K+ L++ INTT D +DYLWY+ SI +N G PVL I
Sbjct: 444 WEVFSEKPGIWGAADFVKNELVDHINTTKDTTDYLWYTTSITVSENEAFLKKGSSPVLFI 503
Query: 497 ESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYGAFYD 556
ES GH LH F+N + G+ G+ + + P+ L G+N IDLLS+TVGL N G+FY+
Sbjct: 504 ESKGHTLHVFINKEYLGTATGNGTHVPFKLKKPVALKAGENNIDLLSMTVGLANAGSFYE 563
Query: 557 TVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVGL-SSGNVG--QWNSQSNLPA 613
VGAG+T V +KG G +++LT+ +W+Y++G++GE + L GN G +W + P
Sbjct: 564 WVGAGLTS-VSIKGFNKG-TLNLTNSKWSYKLGVEGEHLELFKPGNSGAVKWTVTTKPPK 621
Query: 614 NQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYI---SPNSGCTDSC 670
QPLTWYK PSGS PV +D MGKG AW+NG+ IGRYWP SPN C C
Sbjct: 622 KQPLTWYKVVIEPPSGSEPVGLDMISMGKGMAWLNGEEIGRYWPRIARKNSPNDECVKEC 681
Query: 671 NYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTKISFGTKQIESV 730
+YRG + KCL CG+PSQ YHVPR+W K N V+FEE GG+P KI +++ V
Sbjct: 682 DYRGKFMPDKCLTGCGEPSQRWYHVPRSWFKSSGNELVIFEEKGGNPMKIKLSKRKVSVV 741
>At5g56870 beta-galactosidase (emb|CAB64740.1)
Length = 724
Score = 777 bits (2006), Expect = 0.0
Identities = 385/712 (54%), Positives = 490/712 (68%), Gaps = 33/712 (4%)
Query: 24 SNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEP 83
++V+YD +A++I+G+RR+L+SGSIHYPRSTP+MWP LIQK+K+GG+DVIETYVFWN HEP
Sbjct: 27 ASVSYDRKAVIINGQRRILLSGSIHYPRSTPEMWPGLIQKAKEGGLDVIETYVFWNGHEP 86
Query: 84 VRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNN 143
GQY F R DLV F+K V AGLYV+LRIGPYVCAEWN+GGFP+WL F+ G+ FRT+N
Sbjct: 87 SPGQYYFGDRYDLVKFIKLVHQAGLYVNLRIGPYVCAEWNFGGFPVWLKFVPGMAFRTDN 146
Query: 144 EPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAAS 203
EPFKA MK+FT KIV MMK E L+ +QGGPIIL+QIENEYG ++ K+Y W A
Sbjct: 147 EPFKAAMKKFTEKIVWMMKAEKLFQTQGGPIILAQIENEYGPVEWEIGAPGKAYTKWVAQ 206
Query: 204 MATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAFGGA 263
MA L TGVPWIMC+Q +AP PII+TCN +YC+ F PNS NKPKMWTENW+GW+ FGGA
Sbjct: 207 MALGLSTGVPWIMCKQEDAPGPIIDTCNGYYCEDFKPNSINKPKMWTENWTGWYTDFGGA 266
Query: 264 VPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDI 323
VPYRPVED+A++VARF Q+GG+ NYYMYHGGTNF R T G F+++SYDYDAP+DEYG
Sbjct: 267 VPYRPVEDIAYSVARFIQKGGSLVNYYMYHGGTNFDR-TAGEFMASSYDYDAPLDEYGLP 325
Query: 324 RQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTGAVCSAFLANIG-MSDA 382
R+PK+ HLK LHKAIKL E AL+++D T+TS G E V+ + + C+AFL+N S A
Sbjct: 326 REPKYSHLKALHKAIKLSEPALLSADATVTSLGAKQEAYVFWSKSSCAAFLSNKDENSAA 385
Query: 383 TVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKEKVDSLDSSSSGW 442
V F G Y LP WSVSILPDCK V NTAKVN S+ + K +
Sbjct: 386 RVLFRGFPYDLPPWSVSILPDCKTEVYNTAKVNAPSVHRNMVPTGTK------------F 433
Query: 443 SWISEPVGISTPDA-----FTKSGLLEQINTTADRSDYLWYSLSIVYED-----NAGDQP 492
SW S +TP A F ++GL+EQI+ T D+SDY WY I GD P
Sbjct: 434 SWGS--FNEATPTANEAGTFARNGLVEQISMTWDKSDYFWYITDITIGSGETFLKTGDSP 491
Query: 493 VLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYG 552
+L + S GHALH FVNG+L+G+ G + K+ I L G N I LLS+ VGL N G
Sbjct: 492 LLTVMSAGHALHVFVNGQLSGTAYGGLDHPKLTFSQKIKLHAGVNKIALLSVAVGLPNVG 551
Query: 553 AFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGEFVGLSSGNVG---QWNSQS 609
++ G+ GPV LKG+ +G + D++ +W+Y++G++GE + L + +W S
Sbjct: 552 THFEQWNKGVLGPVTLKGVNSG-TWDMSKWKWSYKIGVKGEALSLHTNTESSGVRWTQGS 610
Query: 610 NLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDS 669
+ QPLTWYK+ F P+G+ P+A+D MGKG+ W+NG++IGR+WP Y + G
Sbjct: 611 FVAKKQPLTWYKSTFATPAGNEPLALDMNTMGKGQVWINGRNIGRHWPAYKA--QGSCGR 668
Query: 670 CNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTKIS 721
CNY GT+ A KCL NCG+ SQ YHVPR+WLK N V+FEE GGDP IS
Sbjct: 669 CNYAGTFDAKKCLSNCGEASQRWYHVPRSWLK-SQNLIVVFEELGGDPNGIS 719
>At1g31740 putative protein
Length = 757
Score = 723 bits (1867), Expect = 0.0
Identities = 379/814 (46%), Positives = 492/814 (59%), Gaps = 96/814 (11%)
Query: 21 SFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNL 80
++ + V++D RA+ IDG RRVL+SGSIHYPRST +MWPDLI+K K+G +D IETYVFWN
Sbjct: 11 AYATIVSHDGRAITIDGHRRVLLSGSIHYPRSTTEMWPDLIKKGKEGSLDAIETYVFWNA 70
Query: 81 HEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFR 140
HEP R QY+F G DL+ F+K + G+Y LRIGPYVCAEWNYGGFP+WLH + G++FR
Sbjct: 71 HEPTRRQYDFSGNLDLIRFLKTIQNEGMYGVLRIGPYVCAEWNYGGFPVWLHNMPGMEFR 130
Query: 141 TNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDW 200
T N F EM+ FT IV+M+K+E L+ASQGGPIIL+QIENEYGN+ A K+YI W
Sbjct: 131 TTNTAFMNEMQNFTTMIVEMVKKEKLFASQGGPIILAQIENEYGNVIGSYGEAGKAYIQW 190
Query: 201 AASMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAF 260
A+MA SLD GVPWIMCQQ +AP P++NTCN +YCD F+PN+ N PKMWTENW+GW+ +
Sbjct: 191 CANMANSLDVGVPWIMCQQDDAPQPMLNTCNGYYCDNFSPNNPNTPKMWTENWTGWYKNW 250
Query: 261 GGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEY 320
GG P+R ED+AFAVARFFQ+ GTFQNYYMYHGGTNF RT GGP+I+T+YDYDAP+DE+
Sbjct: 251 GGKDPHRTTEDVAFAVARFFQKEGTFQNYYMYHGGTNFDRTAGGPYITTTYDYDAPLDEF 310
Query: 321 GDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTGAVCSAFLANIG-M 379
G++ QPK+GHLK LH + E+ L + + G + VY+T S F+ N+
Sbjct: 311 GNLNQPKYGHLKQLHDVLHAMEKTLTYGNISTVDFGNLVTATVYQTEEGSSCFIGNVNET 370
Query: 380 SDATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNT-ASMISSFATESLKEKVDSLDSS 438
SDA + F G SY +P WSVSILPDCK NTAK+NT S++ A E+ E S
Sbjct: 371 SDAKINFQGTSYDVPAWSVSILPDCKTETYNTAKINTQTSVMVKKANEAENE------PS 424
Query: 439 SSGWSWISEPVG---ISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVYEDN---AGDQP 492
+ WSW E + + T L +Q + D SDYLWY ++ ++ G
Sbjct: 425 TLKWSWRPENIDSVLLKGKGESTMRQLFDQKVVSNDESDYLWYMTTVNLKEQDPVLGKNM 484
Query: 493 VLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYG 552
L I S H LHAFVNG+ G+ +G + G N I LLS+TVGL NYG
Sbjct: 485 SLRINSTAHVLHAFVNGQHIGNYRVENGKFHYVFEQDAKFNPGANVITLLSITVGLPNYG 544
Query: 553 AFYDTVGAGITGPVILKGLKNGSSV---DLTSQQWTYQVGLQGEFVGLSSGNVGQWNSQS 609
AF++ AGITGPV + G +NG DL++ +W+Y+ GL G L S
Sbjct: 545 AFFENFSAGITGPVFIIG-RNGDETIVKDLSTHKWSYKTGLSGFENQLFS---------- 593
Query: 610 NLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDS 669
+ P TW AP GS PV +D G+GKG AW+NG +IGRYWP ++S G
Sbjct: 594 ---SESPSTW-----SAPLGSEPVVVDLLGLGKGTAWINGNNIGRYWPAFLSDIDG---- 641
Query: 670 CNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGGDPTKISFGTKQIES 729
NT VLFEE GG+P+ ++F T + S
Sbjct: 642 ----------------------------------DNTLVLFEEIGGNPSLVNFQTIGVGS 667
Query: 730 VCSHVTESHPPPVDTWNSNAESERKVGPVLSLECPYPNQAISSIKFASFGTPRGTCGNYN 789
VC++V E + VL L C + IS+IKFASFG P G CG++
Sbjct: 668 VCANVYEKN-------------------VLELSC--NGKPISAIKFASFGNPGGDCGSFE 706
Query: 790 HGSC-SSNRALSIVQKACIGSSSCNIGVSINTFG 822
G+C +SN A +I+ + C+G C+I VS + FG
Sbjct: 707 KGTCEASNNAAAILTQECVGKEKCSIDVSEDKFG 740
>At1g77410 unknown protein
Length = 815
Score = 719 bits (1856), Expect = 0.0
Identities = 391/833 (46%), Positives = 508/833 (60%), Gaps = 55/833 (6%)
Query: 5 QIVFVLLWFLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKS 64
Q V L + V V A +NVTYD R+L+IDG+ ++L SGSIHY RSTPQMWP LI K+
Sbjct: 5 QYSLVFLVLMAVIV-AGDVANVTYDGRSLIIDGEHKILFSGSIHYTRSTPQMWPSLIAKA 63
Query: 65 KDGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNY 124
K GGIDV++TYVFWN+HEP +GQ++F G D+V F+K V GLYV LRIGP++ EW+Y
Sbjct: 64 KSGGIDVVDTYVFWNVHEPQQGQFDFSGSRDIVKFIKEVKNHGLYVCLRIGPFIQGEWSY 123
Query: 125 GGFPLWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYG 184
GG P WLH + GI FRT+NEPFK MKR+ IV +MK ENLYASQGGPIILSQIENEYG
Sbjct: 124 GGLPFWLHNVQGIVFRTDNEPFKYHMKRYAKMIVKLMKSENLYASQGGPIILSQIENEYG 183
Query: 185 NIDTHDARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYCDQF--TPNS 242
+ + KSY+ W A +A LDTGVPW+MC+Q +APDP++N CN C + PNS
Sbjct: 184 MVGRAFRQEGKSYVKWTAKLAVELDTGVPWVMCKQDDAPDPLVNACNGRQCGETFKGPNS 243
Query: 243 DNKPKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTT 302
NKP +WTENW+ ++ +G R ED+AF VA F + G+F NYYMYHGGTNFGR
Sbjct: 244 PNKPAIWTENWTSFYQTYGEEPLIRSAEDIAFHVALFIAKNGSFVNYYMYHGGTNFGR-N 302
Query: 303 GGPFISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETA 362
F+ TSY AP+DEYG +RQPKWGHLK+LH A+KLCEE L++ T S G L+TA
Sbjct: 303 ASQFVITSYYDQAPLDEYGLLRQPKWGHLKELHAAVKLCEEPLLSGLQTTISLG-KLQTA 361
Query: 363 VY--KTGAVCSAFLANIGMSDATVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMI 420
K +C+A L N ++TV F +SY L SVS+LPDCKNV NTAKVN
Sbjct: 362 FVFGKKANLCAAILVNQDKCESTVQFRNSSYRLSPKSVSVLPDCKNVAFNTAKVN----- 416
Query: 421 SSFATESLKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSL 480
+ + T + K + + SS W +E V + + LLE +NTT D SDYLW +
Sbjct: 417 AQYNTRTRKARQNL--SSPQMWEEFTETVPSFSETSIRSESLLEHMNTTQDTSDYLWQTT 474
Query: 481 SIVYEDNAGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTID 540
++ + G VL + LGHALHAFVNG+ GS G+ + ++ ++L G N +
Sbjct: 475 R--FQQSEGAPSVLKVNHLGHALHAFVNGRFIGSMHGTFKAHRFLLEKNMSLNNGTNNLA 532
Query: 541 LLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGE--FVGLS 598
LLS+ VGL N GA + + G +K + + W YQVGL+GE V
Sbjct: 533 LLSVMVGLPNSGAHLE---RRVVGSRSVKIWNGRYQLYFNNYSWGYQVGLKGEKFHVYTE 589
Query: 599 SGNVGQWNSQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPT 658
G+ Q +QPLTWYK +F P G +PVA++ MGKGEAWVNGQSIGRYW +
Sbjct: 590 DGSAKVQWKQYRDSKSQPLTWYKASFDTPEGEDPVALNLGSMGKGEAWVNGQSIGRYWVS 649
Query: 659 YISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEES-GGDP 717
+ TY G PSQ YH+PR++LKP+SN V+ EE G+P
Sbjct: 650 F--------------HTYK--------GNPSQIWYHIPRSFLKPNSNLLVILEEEREGNP 687
Query: 718 TKISFGTKQIESVCSHVTESHPPPV--------DTWNSNAESERKVGPVLSLECPYPNQA 769
I+ T + VC HV+ ++P PV + N +RK P + L+CP +
Sbjct: 688 LGITIDTVSVTEVCGHVSNTNPHPVISPRKKGLNRKNLTYRYDRK--PKVQLQCP-TGRK 744
Query: 770 ISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTFG 822
IS I FASFGTP G+CG+Y+ GSC S +L++VQKAC+ S C++ V TFG
Sbjct: 745 ISKILFASFGTPNGSCGSYSIGSCHSPNSLAVVQKACLKKSRCSVPVWSKTFG 797
>At2g16730 putative beta-galactosidase
Length = 832
Score = 646 bits (1667), Expect = 0.0
Identities = 349/840 (41%), Positives = 499/840 (58%), Gaps = 70/840 (8%)
Query: 8 FVLLWFLGVYVPASFCSNVTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDG 67
++LL L + + S ++TYD +L+I+G R +L SGSIHYPRSTP+MWP++I+++K G
Sbjct: 10 WLLLAVLVILLSFSGALSITYDGTSLIINGNRELLYSGSIHYPRSTPEMWPNIIKRAKQG 69
Query: 68 GIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGF 127
G++ I+TYVFWN+HEP +G++NF GR DLV F+K + GLYV LR+GP++ AEW +GG
Sbjct: 70 GLNTIQTYVFWNVHEPEQGKFNFSGRADLVKFIKLIEKNGLYVTLRLGPFIQAEWTHGGL 129
Query: 128 PLWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNID 187
P WL + GI FRT+NEPFK +R+ ++DMMK+E L+ASQGGPIIL QIENEY +
Sbjct: 130 PYWLREVPGIFFRTDNEPFKEHTERYVKVVLDMMKEEKLFASQGGPIILGQIENEYSAVQ 189
Query: 188 THDARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYC-DQFT-PNSDNK 245
+YI WA+ + S+D G+PW+MC+Q +APDP+IN CN +C D F PN DNK
Sbjct: 190 RAYKEDGLNYIKWASKLVHSMDLGIPWVMCKQNDAPDPMINACNGRHCGDTFPGPNKDNK 249
Query: 246 PKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGP 305
P +WTENW+ F FG R VED+A++VARFF + GT NYYMYHGGTNFGRT+
Sbjct: 250 PSLWTENWTTQFRVFGDPPAQRSVEDIAYSVARFFSKNGTHVNYYMYHGGTNFGRTS-AH 308
Query: 306 FISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYK 365
+++T Y DAP+DE+G R+PK+GHLK LH A+ LC++AL+ P + P E Y+
Sbjct: 309 YVTTRYYDDAPLDEFGLEREPKYGHLKHLHNALNLCKKALLWGQPRVEKPSNETEIRYYE 368
Query: 366 TGA--VCSAFLANIGMSDA-TVTFNGNSYHLPGWSVSILPDCKNVVLNTAKV-----NTA 417
VC+AFLAN A + F G Y +P S+SILPDCK VV NT ++ +
Sbjct: 369 QPGTKVCAAFLANNNTEAAEKIKFRGKEYLIPHRSISILPDCKTVVYNTGEIISHHTSRN 428
Query: 418 SMISSFATESLKEKV--DSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDY 475
M S A ++ KV +S+ S G S+I +E T D SDY
Sbjct: 429 FMKSKKANKNFDFKVFTESVPSKIKGDSFIP----------------VELYGLTKDESDY 472
Query: 476 LWYSLSIVYEDN-----AGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPI 530
WY+ S +DN G +P L I SLGHALH ++NG+ G+ GS P+
Sbjct: 473 GWYTTSFKIDDNDLSKKKGGKPNLRIASLGHALHVWLNGEYLGNGHGSHEEKSFVFQKPV 532
Query: 531 TLVTGKNTIDLLSLTVGLQNYGAFYDTVGAGITGP--VILKGLKNGSSVDLTSQ-QWTYQ 587
TL G+N + +L + G + G++ + TGP V + GL +G ++DLT + +W +
Sbjct: 533 TLKEGENHLTMLGVLTGFPDSGSYME---HRYTGPRSVSILGLGSG-TLDLTEENKWGNK 588
Query: 588 VGLQGEFVGLSSGNVGQWNSQSNLPANQP-LTWYKTNFVAPSGSNPVAIDFTGMGKGEAW 646
VG++GE +G+ + + +P +TWY+T F AP + AI GMGKG W
Sbjct: 589 VGMEGERLGIHAEEGLKKVKWEKASGKEPGMTWYQTYFDAPESQSAAAIRMNGMGKGLIW 648
Query: 647 VNGQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNT 706
VNG+ +GRYW +++SP G+P+Q YH+PR++LKP N
Sbjct: 649 VNGEGVGRYWMSFLSP----------------------LGQPTQIEYHIPRSFLKPKKNL 686
Query: 707 FVLFEESGG-DPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAESERKVGP----VLSL 761
V+FEE P I F ++VCS++ E++ P V W + + + +L
Sbjct: 687 LVIFEEEPNVKPELIDFVIVNRDTVCSYIGENYTPSVRHWTRKNDQVQAITDDVHLTANL 746
Query: 762 ECPYPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTF 821
+C + IS+++FASFG P GTCGN+ GSC++ + +V+K C+G + C I V+ +TF
Sbjct: 747 KCS-GTKKISAVEFASFGNPNGTCGNFTLGSCNAPVSKKVVEKYCLGKAECVIPVNKSTF 805
>At5g63800 beta-galactosidase like protein
Length = 657
Score = 610 bits (1572), Expect = e-174
Identities = 318/676 (47%), Positives = 417/676 (61%), Gaps = 40/676 (5%)
Query: 56 MWPDLIQKSKDGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIG 115
MWP LI+K+K+GGIDVI+TYVFWNLHEP GQY+F GR DLV F+K + + GLYV LRIG
Sbjct: 1 MWPSLIKKTKEGGIDVIQTYVFWNLHEPKLGQYDFSGRNDLVKFIKEIRSQGLYVCLRIG 60
Query: 116 PYVCAEWNYGGFPLWLHFIAGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPII 175
P++ AEWNYGG P WL + G+ +RT+NEPFK M++FTAKIVD+MK E LYASQGGPII
Sbjct: 61 PFIEAEWNYGGLPFWLRDVPGMVYRTDNEPFKFHMQKFTAKIVDLMKSEGLYASQGGPII 120
Query: 176 LSQIENEYGNIDTHDARAAKSYIDWAASMATSLDTGVPWIMCQQANAPDPIINTCNSFYC 235
LSQIENEY N++ SYI WA MA L TGVPWIMC+ +APDP+INTCN C
Sbjct: 121 LSQIENEYANVEGAFHEKGASYIKWAGQMAVGLKTGVPWIMCKSPDAPDPVINTCNGMKC 180
Query: 236 DQF--TPNSDNKPKMWTENWSGWFLAFGGAVPYRPVEDLAFAVARFFQRGGTFQNYYMYH 293
+ PNS NKPKMWTE+W+ +F +G R ED+AF A F + G++ NYYMYH
Sbjct: 181 GETFPGPNSPNKPKMWTEDWTSFFQVYGKEPYIRSAEDIAFHAALFVAKNGSYINYYMYH 240
Query: 294 GGTNFGRTTGGPFISTSYDYDAPIDEYGDIRQPKWGHLKDLHKAIKLCEEALIASDPTIT 353
GGTNFGRT+ FI+ YD AP+DEYG +RQPK+GHLK+LH AIK L+ TI
Sbjct: 241 GGTNFGRTSSSYFITGYYD-QAPLDEYGLLRQPKYGHLKELHAAIKSSANPLLQGKQTIL 299
Query: 354 SPGPNLETAVYK-TGAVCSAFLANIGMSDATVTFNGNSYHLPGWSVSILPDCKNVVLNTA 412
S GP + V++ C AFL N + + F N+Y L S+ IL +CKN++ TA
Sbjct: 300 SLGPMQQAYVFEDANNGCVAFLVNNDAKASQIQFRNNAYSLSPKSIGILQNCKNLIYETA 359
Query: 413 KVNTASMISSFATESLKEKVDSLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADR 472
KVN +++ T ++ + + W+ E + + + LLE N T D+
Sbjct: 360 KVNV--KMNTRVTTPVQ-----VFNVPDNWNLFRETIPAFPGTSLKTNALLEHTNLTKDK 412
Query: 473 SDYLWYSLSIVYEDNAGDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITL 532
+DYLWY+ S D+ P ++ ES GH +H FVN LAGS GS V + P++L
Sbjct: 413 TDYLWYTSSFKL-DSPCTNPSIYTESSGHVVHVFVNNALAGSGHGSRDIRVVKLQAPVSL 471
Query: 533 VTGKNTIDLLSLTVGLQNYGAFYDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQG 592
+ G+N I +LS VGL + GA+ + G+T I G +DL+ QW Y VGL G
Sbjct: 472 INGQNNISILSGMVGLPDSGAYMERRSYGLTKVQISCG--GTKPIDLSRSQWGYSVGLLG 529
Query: 593 EFVGL---SSGNVGQWN-SQSNLPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVN 648
E V L + N +W+ +++ L N+PL WYKT F P+G PV + + MGKGE WVN
Sbjct: 530 EKVRLYQWKNLNRVKWSMNKAGLIKNRPLAWYKTTFDGPNGDGPVGLHMSSMGKGEIWVN 589
Query: 649 GQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFV 708
G+SIGRYW ++++P G+PSQ++YH+PRA+LKP N V
Sbjct: 590 GESIGRYWVSFLTP----------------------AGQPSQSIYHIPRAFLKPSGNLLV 627
Query: 709 LFEESGGDPTKISFGT 724
+FEE GGDP IS T
Sbjct: 628 VFEEEGGDPLGISLNT 643
>At4g38590 galactosidase like protein
Length = 1036
Score = 540 bits (1390), Expect = e-153
Identities = 302/760 (39%), Positives = 431/760 (55%), Gaps = 63/760 (8%)
Query: 87 QYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEPF 146
QY+F+GR DLV F+K + GLYV LR+GP++ AEWN+GG P WL + + FRTNNEPF
Sbjct: 80 QYDFKGRFDLVKFIKLIHEKGLYVTLRLGPFIQAEWNHGGLPYWLREVPDVYFRTNNEPF 139
Query: 147 KAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAASMAT 206
K +R+ KI+ MMK+E L+ASQGGPIIL QIENEY + + YI WAA++
Sbjct: 140 KEHTERYVRKILGMMKEEKLFASQGGPIILGQIENEYNAVQLAYKENGEKYIKWAANLVE 199
Query: 207 SLDTGVPWIMCQQANAPDPIINTCNSFYC-DQFT-PNSDNKPKMWTENWSGWFLAFGGAV 264
S++ G+PW+MC+Q +AP +IN CN +C D F PN +KP +WTENW+ F FG
Sbjct: 200 SMNLGIPWVMCKQNDAPGNLINACNGRHCGDTFPGPNRHDKPSLWTENWTTQFRVFGDPP 259
Query: 265 PYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDIR 324
R VED+AF+VAR+F + G+ NYYMYHGGTNFGRT+ F++T Y DAP+DE+G +
Sbjct: 260 TQRTVEDIAFSVARYFSKNGSHVNYYMYHGGTNFGRTS-AHFVTTRYYDDAPLDEFGLEK 318
Query: 325 QPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTGA--VCSAFLANIGMSDA 382
PK+GHLK +H+A++LC++AL + GP+ E Y+ VC+AFL+N D
Sbjct: 319 APKYGHLKHVHRALRLCKKALFWGQLRAQTLGPDTEVRYYEQPGTKVCAAFLSNNNTRDT 378
Query: 383 -TVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKEKVDSLDSSSSG 441
T+ F G Y LP S+SILPDCK VV NTA++ A S ++ V S + +S G
Sbjct: 379 NTIKFKGQDYVLPSRSISILPDCKTVVYNTAQI--------VAQHSWRDFVKS-EKTSKG 429
Query: 442 WSWISEPVGISTPDAFTKSGLL--EQINTTADRSDYLWYSLSIV-----YEDNAGDQPVL 494
+ E + P L+ E T D++DY WY+ S+ + D G + +L
Sbjct: 430 LKF--EMFSENIPSLLDGDSLIPGELYYLTKDKTDYAWYTTSVKIDEDDFPDQKGLKTIL 487
Query: 495 HIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGLQNYGAF 554
+ SLGHAL +VNG+ AG G P+ TG N I +L + GL + G++
Sbjct: 488 RVASLGHALIVYVNGEYAGKAHGRHEMKSFEFAKPVNFKTGDNRISILGVLTGLPDSGSY 547
Query: 555 YDTVGAGITGPVILKGLKNGSSVDLTSQQWTYQVGLQGE----FVGLSSGNVGQWNSQSN 610
+ AG I+ GLK+G+ + +W + GL+GE + S V +W
Sbjct: 548 MEHRFAGPRAISII-GLKSGTRDLTENNEWGHLAGLEGEKKEVYTEEGSKKV-KWEKDGK 605
Query: 611 LPANQPLTWYKTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSC 670
+PLTWYKT F P G N VAI MGKG WVNG +GRYW +++SP
Sbjct: 606 ---RKPLTWYKTYFETPEGVNAVAIRMKAMGKGLIWVNGIGVGRYWMSFLSP-------- 654
Query: 671 NYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPD--SNTFVLFEESGGDPTK-ISFGTKQI 727
G+P+QT YH+PR+++K + N V+ EE G + I F
Sbjct: 655 --------------LGEPTQTEYHIPRSFMKGEKKKNMLVILEEEPGVKLESIDFVLVNR 700
Query: 728 ESVCSHVTESHPPPVDTWNSN----AESERKVGPVLSLECPYPNQAISSIKFASFGTPRG 783
+++CS+V E +P V +W + + + CP P + + ++FASFG P G
Sbjct: 701 DTICSNVGEDYPVSVKSWKREGPKIVSRSKDMRLKAVMRCP-PEKQMVEVQFASFGDPTG 759
Query: 784 TCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTFGN 823
TCGN+ G CS++++ +V+K C+G + C+I V+ TFG+
Sbjct: 760 TCGNFTMGKCSASKSKEVVEKECLGRNYCSIVVARETFGD 799
>At4g35010 beta-galactosidase - like protein
Length = 831
Score = 536 bits (1381), Expect = e-152
Identities = 306/829 (36%), Positives = 446/829 (52%), Gaps = 111/829 (13%)
Query: 26 VTYDHRALVIDGKRRVLMSGSIHYPRSTPQMWPDLIQKSKDGGIDVIETYVFWNLHEPVR 85
VTYD +L+IDGKR +L SGSIHYPRSTP+MWP +I+++K GG++ I+TYVFWN+HEP +
Sbjct: 54 VTYDGTSLIIDGKRELLYSGSIHYPRSTPEMWPSIIKRAKQGGLNTIQTYVFWNVHEPQQ 113
Query: 86 GQYNFEGRGDLVGFVKAVAAAGLYVHLRIGPYVCAEWNYGGFPLWLHFIAGIKFRTNNEP 145
G++NF GR DLV F+K + G+YV LR+GP++ AEW +G + H +R
Sbjct: 114 GKFNFSGRADLVKFIKLIQKNGMYVTLRLGPFIQAEWTHGYITRYDHKNIAGAYR----- 168
Query: 146 FKAEMKRFTAKIVDMMKQENLYASQGGPIILSQIENEYGNIDTHDARAAKSYIDWAASMA 205
+IENEY + + +YI WA+++
Sbjct: 169 --------------------------------KIENEYSAVQRAYKQDGLNYIKWASNLV 196
Query: 206 TSLDTGVPWIMCQQANAPDPIINTCNSFYC-DQFT-PNSDNKPKMWTENWSGWFLAFGGA 263
S+ G+PW+MC+Q +APDP+IN CN +C D F PN +NKP +WTENW+ F FG
Sbjct: 197 DSMKLGIPWVMCKQNDAPDPMINACNGRHCGDTFPGPNRENKPSLWTENWTTQFRVFGDP 256
Query: 264 VPYRPVEDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFISTSYDYDAPIDEYGDI 323
R VED+A++VARFF + GT NYYMYHGGTNFGRT+ +++T Y DAP+DEYG
Sbjct: 257 PTQRSVEDIAYSVARFFSKNGTHVNYYMYHGGTNFGRTS-AHYVTTRYYDDAPLDEYGLE 315
Query: 324 RQPKWGHLKDLHKAIKLCEEALIASDPTITSPGPNLETAVYKTGA--VCSAFLANIGMSD 381
++PK+GHLK LH A+ LC++ L+ P PG + E Y+ C+AFLAN
Sbjct: 316 KEPKYGHLKHLHNALNLCKKPLLWGQPKTEKPGKDTEIRYYEQPGTKTCAAFLANNNTEA 375
Query: 382 A-TVTFNGNSYHLPGWSVSILPDCKNVVLNTAKVNTASMISSFATESLKEK-------VD 433
A T+ F G Y + S+SILPDCK VV NTA++ + +F K +
Sbjct: 376 AETIKFKGREYVIAPRSISILPDCKTVVYNTAQIVSQHTSRNFMKSKKANKKFDFKVFTE 435
Query: 434 SLDSSSSGWSWISEPVGISTPDAFTKSGLLEQINTTADRSDYLWYSLSIVYEDN-----A 488
+L S G S+I +E T D++DY WY+ S N
Sbjct: 436 TLPSKLEGNSYIP----------------VELYGLTKDKTDYGWYTTSFKVHKNHLPTKK 479
Query: 489 GDQPVLHIESLGHALHAFVNGKLAGSKAGSSGNAKVNVDIPITLVTGKNTIDLLSLTVGL 548
G + + I SLGHALHA++NG+ GS GS +TL G+N + +L + G
Sbjct: 480 GVKTFVRIASLGHALHAWLNGEYLGSGHGSHEEKSFVFQKQVTLKAGENHLVMLGVLTGF 539
Query: 549 QNYGAFYDTVGAGITGPVILKGLKNGSSVDLT-SQQWTYQVGLQGEFVGLSSGNVGQWNS 607
+ G++ + G G IL GL +G ++DLT S +W ++G++GE +G+ + +
Sbjct: 540 PDSGSYMEHRYTGPRGISIL-GLTSG-TLDLTESSKWGNKIGMEGEKLGIHTEEGLKKVE 597
Query: 608 QSNLPANQP-LTWY----------KTNFVAPSGSNPVAIDFTGMGKGEAWVNGQSIGRYW 656
P LTWY +T F AP + I GMGKG WVNG+ +GRYW
Sbjct: 598 WKKFTGKAPGLTWYQKFSKECETLQTYFDAPESVSAATIRMHGMGKGLIWVNGEGVGRYW 657
Query: 657 PTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYHVPRAWLKPDSNTFVLFEESGG- 715
+++SP G+P+Q YH+PR++LKP N V+FEE
Sbjct: 658 QSFLSP----------------------LGQPTQIEYHIPRSFLKPKKNLLVIFEEEPNV 695
Query: 716 DPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAESERKVGPVLSLECPYP---NQAISS 772
P + F ++VCS+V E++ P V W + + + +SL + I++
Sbjct: 696 KPELMDFAIVNRDTVCSYVGENYTPSVRHWTRKKDQVQAITDNVSLTATLKCSGTKKIAA 755
Query: 773 IKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTF 821
++FASFG P G CGN+ G+C++ + +++K C+G + C I V+ +TF
Sbjct: 756 VEFASFGNPIGVCGNFTLGTCNAPVSKQVIEKHCLGKAECVIPVNKSTF 804
>At1g72990 unknown protein
Length = 697
Score = 161 bits (407), Expect = 2e-39
Identities = 125/376 (33%), Positives = 172/376 (45%), Gaps = 61/376 (16%)
Query: 7 VFVLLWFLGVYVPASFCSNVTYDHRALVI-----------DGKRRVLMSGSIHYPRSTPQ 55
VF LL L Y P S S + D + + DG R ++ G +HY R P+
Sbjct: 34 VFALLPSLS-YTPQSLPSAIPQDEKMISRKFYIKDDNFWKDGNRFQIIGGDLHYFRVLPE 92
Query: 56 MWPDLIQKSKDGGIDVIETYVFWNLHEPVRGQYNFEGRGDLVGFVKAVAAAGLYVHLRIG 115
W D + ++ G++ I+ YV WNLHEP G+ FEG GDLV F+K V LR G
Sbjct: 93 YWEDRLLRANALGLNTIQVYVPWNLHEPKPGKMVFEGIGDLVSFLKLCEKLDFLVMLRAG 152
Query: 116 PYVCAEWNYGGFPLWLHFI-AGIKFRTNNEPFKAEMKRFTAKIVDMMKQENLYASQGGPI 174
PY+C EW+ GGFP WL + ++ RT++ + ++R+ V + K L S GGP+
Sbjct: 153 PYICGEWDLGGFPAWLLAVKPRLQLRTSDPVYLKLVERWWD--VLLPKVFPLLYSNGGPV 210
Query: 175 ILSQIENEYGNIDTHDARAAKSYIDWAASMA------------------TSLDTG-VPWI 215
I+ QIENEYG+ K+Y+ SMA +LD G VP
Sbjct: 211 IMVQIENEYGSYGND-----KAYLRKLVSMARGHLGDDIIVYTTDGGTKETLDKGTVPVA 265
Query: 216 MCQQA------NAPDPIINTCNSFYCDQFTPNSDNKPKMWTENWSGWFLAFGGAVPYRPV 269
A + P PI F P + +E ++GW +G +
Sbjct: 266 DVYSAVDFSTGDDPWPIFKLQKKFNA------PGRSPPLSSEFYTGWLTHWGEKITKTDA 319
Query: 270 EDLAFAVARFFQRGGTFQNYYMYHGGTNFGRTTGGPFIS---------TSYDYDAPIDEY 320
E A ++ + R G+ YM HGGTNFG G S TSYDYDAPI E
Sbjct: 320 EFTAASLEKILSRNGS-AVLYMVHGGTNFGFYNGANTGSEESDYKPDLTSYDYDAPIKES 378
Query: 321 GDIRQPKWGHLKDLHK 336
GDI PK+ L+ + K
Sbjct: 379 GDIDNPKFQALQRVIK 394
Score = 37.4 bits (85), Expect = 0.035
Identities = 26/77 (33%), Positives = 33/77 (42%), Gaps = 28/77 (36%)
Query: 635 IDFTGMGKGEAWVNGQSIGRYWPTYISPNSGCTDSCNYRGTYSASKCLKNCGKPSQTLYH 694
+ F G GKG A+VN +IGRYWP+ + P CN +
Sbjct: 619 LSFNGWGKGVAFVNEFNIGRYWPS-VGP------QCN---------------------LY 650
Query: 695 VPRAWLKPDSNTFVLFE 711
VP LK NT V+FE
Sbjct: 651 VPAPLLKRGKNTLVVFE 667
>At3g53080 unknown protein
Length = 155
Score = 65.9 bits (159), Expect = 9e-11
Identities = 26/71 (36%), Positives = 41/71 (57%)
Query: 752 ERKVGPVLSLECPYPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSS 811
E +GP+ + C P I+ I FA +G P GTCG++ G+C + + IV+K C+G
Sbjct: 66 EPDLGPLTRISCNEPGYVITKINFADYGNPTGTCGHFRRGNCGARATMRIVKKNCLGKEK 125
Query: 812 CNIGVSINTFG 822
C++ V+ FG
Sbjct: 126 CHLLVTDEMFG 136
>At3g53050 putative protein
Length = 142
Score = 52.0 bits (123), Expect = 1e-06
Identities = 31/117 (26%), Positives = 50/117 (42%), Gaps = 1/117 (0%)
Query: 707 FVLFEESGGDPTKISFGTKQIESVCSHVTESHPPPVDTWNSNAESERKVGPVLSLECPYP 766
F+ F S +SF + S+ S + S + E K G ++L+
Sbjct: 19 FLSFVFSFASKIDVSFHGDEKRSLSSSINSSLSESAEYTMCATHKEAKDGKPIALDFDCE 78
Query: 767 N-QAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQKACIGSSSCNIGVSINTFG 822
IS I +A +G G+CG + G+C ++ L+IV K C+ C + V FG
Sbjct: 79 QGYVISKITYADYGQSTGSCGKFKRGNCGASNTLNIVNKKCLRKEKCKLFVPDKIFG 135
>At3g53070 putative protein
Length = 105
Score = 43.9 bits (102), Expect = 4e-04
Identities = 19/52 (36%), Positives = 26/52 (49%)
Query: 753 RKVGPVLSLECPYPNQAISSIKFASFGTPRGTCGNYNHGSCSSNRALSIVQK 804
R PVL +C S I FA +G G CGN+ G+C + L +V+K
Sbjct: 47 RSRAPVLMFDCKEKGYVFSKINFADYGHSSGDCGNFRRGTCGAPDTLRLVKK 98
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.135 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,673,016
Number of Sequences: 26719
Number of extensions: 968138
Number of successful extensions: 2120
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1958
Number of HSP's gapped (non-prelim): 35
length of query: 826
length of database: 11,318,596
effective HSP length: 108
effective length of query: 718
effective length of database: 8,432,944
effective search space: 6054853792
effective search space used: 6054853792
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC139344.4


