

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137602.4 - phase: 0
(130 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
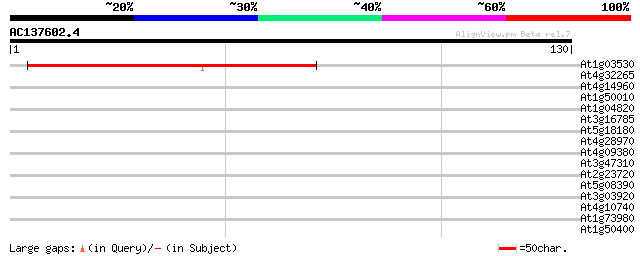
Score E
Sequences producing significant alignments: (bits) Value
At1g03530 unknown protein 83 3e-17
At4g32265 unknown protein 32 0.090
At4g14960 tubulin alpha-6 chain (TUA6) 29 0.59
At1g50010 tubulin alpha-2/alpha-4 chain, putative 29 0.59
At1g04820 tubulin alpha-2/alpha-4 chain 29 0.59
At3g16785 phospholipase D, putative, 5' partial 28 1.00
At5g18180 putative protein 27 2.9
At4g28970 putative protein 27 2.9
At4g09380 putative protein 27 3.8
At3g47310 putative protein 27 3.8
At2g23720 Mutator-like transposase 27 3.8
At5g08390 katanin p80 subunit - like protein 26 5.0
At3g03920 putative GAR1 protein 26 5.0
At4g10740 hypothetical protein 26 6.5
At1g73980 unknown protein 26 6.5
At1g50400 hypothetical protein 26 6.5
>At1g03530 unknown protein
Length = 801
Score = 83.2 bits (204), Expect = 3e-17
Identities = 39/71 (54%), Positives = 54/71 (75%), Gaps = 4/71 (5%)
Query: 5 PLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSE----KGIREGTLISFVAEFVN 60
PL EGSILWITE++T LGL+DEIFG VK PYY VR+NSE +G+ +GT +SFVA+F
Sbjct: 405 PLTEGSILWITEKRTPLGLVDEIFGPVKCPYYIVRFNSESEVPEGVCQGTPVSFVADFAQ 464
Query: 61 YVMHPVSMMKR 71
++++ + K+
Sbjct: 465 HILNIKELQKK 475
>At4g32265 unknown protein
Length = 239
Score = 32.0 bits (71), Expect = 0.090
Identities = 18/60 (30%), Positives = 31/60 (51%), Gaps = 2/60 (3%)
Query: 40 YNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCPMKQNFWMMRKRLSTRDC--KNIIKEV 97
Y +E GI++G + F+ NY M V + P +F ++ +T + KN+I+EV
Sbjct: 106 YLTEFGIKDGDQLRFIRHISNYCMLMVKHKSKTPRVSSFKQLKLFSTTPETRKKNVIREV 165
>At4g14960 tubulin alpha-6 chain (TUA6)
Length = 450
Score = 29.3 bits (64), Expect = 0.59
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 7/91 (7%)
Query: 14 ITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCP 73
+TE QT+L I + + YA ++EK E VAE N P SMM +C
Sbjct: 252 VTEFQTNLVPYPRIHFMLSS--YAPVISAEKAFHEQLS---VAEITNSAFEPASMMAKCD 306
Query: 74 MKQNFWMMRKRLSTRDC--KNIIKEVVTIKT 102
+ +M + D K++ V TIKT
Sbjct: 307 PRHGKYMACCLMYRGDVVPKDVNAAVGTIKT 337
>At1g50010 tubulin alpha-2/alpha-4 chain, putative
Length = 450
Score = 29.3 bits (64), Expect = 0.59
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 7/91 (7%)
Query: 14 ITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCP 73
+TE QT+L I + + YA ++EK E VAE N P SMM +C
Sbjct: 252 VTEFQTNLVPYPRIHFMLSS--YAPVISAEKAFHEQLS---VAEITNSAFEPASMMAKCD 306
Query: 74 MKQNFWMMRKRLSTRDC--KNIIKEVVTIKT 102
+ +M + D K++ V TIKT
Sbjct: 307 PRHGKYMACCLMYRGDVVPKDVNAAVGTIKT 337
>At1g04820 tubulin alpha-2/alpha-4 chain
Length = 450
Score = 29.3 bits (64), Expect = 0.59
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 7/91 (7%)
Query: 14 ITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCP 73
+TE QT+L I + + YA ++EK E VAE N P SMM +C
Sbjct: 252 VTEFQTNLVPYPRIHFMLSS--YAPVISAEKAFHEQLS---VAEITNSAFEPASMMAKCD 306
Query: 74 MKQNFWMMRKRLSTRDC--KNIIKEVVTIKT 102
+ +M + D K++ V TIKT
Sbjct: 307 PRHGKYMACCLMYRGDVVPKDVNAAVGTIKT 337
>At3g16785 phospholipase D, putative, 5' partial
Length = 682
Score = 28.5 bits (62), Expect = 1.00
Identities = 12/38 (31%), Positives = 20/38 (52%)
Query: 12 LWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREG 49
+W+ +T+ + ++F V N RYNS+K R G
Sbjct: 633 IWMATAKTNTMIYQDVFSCVPNDLIHSRYNSDKAYRIG 670
>At5g18180 putative protein
Length = 189
Score = 26.9 bits (58), Expect = 2.9
Identities = 11/54 (20%), Positives = 29/54 (53%), Gaps = 6/54 (11%)
Query: 16 ERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMM 69
+ +T +G +DEIFG + ++++ +REG + + ++ + + P ++
Sbjct: 78 QNKTQIGRVDEIFGPINESLFSIK------MREGIVATSYSQGDKFFISPEKLL 125
>At4g28970 putative protein
Length = 914
Score = 26.9 bits (58), Expect = 2.9
Identities = 10/26 (38%), Positives = 20/26 (76%), Gaps = 1/26 (3%)
Query: 89 DCKNII-KEVVTIKTPIERMENNIKQ 113
DC+N++ E+V I+ P+ER N++++
Sbjct: 153 DCENVVGTEIVAIERPMERPVNSVEE 178
>At4g09380 putative protein
Length = 960
Score = 26.6 bits (57), Expect = 3.8
Identities = 9/26 (34%), Positives = 20/26 (76%), Gaps = 1/26 (3%)
Query: 89 DCKNII-KEVVTIKTPIERMENNIKQ 113
DC+N++ E+V ++ P+ER N++++
Sbjct: 153 DCENVVGTEIVAVERPMERPVNSVEE 178
>At3g47310 putative protein
Length = 735
Score = 26.6 bits (57), Expect = 3.8
Identities = 9/26 (34%), Positives = 20/26 (76%), Gaps = 1/26 (3%)
Query: 89 DCKNII-KEVVTIKTPIERMENNIKQ 113
DC+N++ E+V ++ P+ER N++++
Sbjct: 102 DCENVVGTEIVAVERPMERPVNSVEE 127
>At2g23720 Mutator-like transposase
Length = 821
Score = 26.6 bits (57), Expect = 3.8
Identities = 9/26 (34%), Positives = 20/26 (76%), Gaps = 1/26 (3%)
Query: 89 DCKNII-KEVVTIKTPIERMENNIKQ 113
DC+N++ E+V ++ P+ER N++++
Sbjct: 183 DCENVVGTEIVAVERPMERPVNSVEE 208
>At5g08390 katanin p80 subunit - like protein
Length = 823
Score = 26.2 bits (56), Expect = 5.0
Identities = 11/42 (26%), Positives = 20/42 (47%)
Query: 58 FVNYVMHPVSMMKRCPMKQNFWMMRKRLSTRDCKNIIKEVVT 99
F+N P+ + ++ FW++ K L R CK + + T
Sbjct: 45 FLNLHSFPIDLNLVSVLRLAFWVLLKNLDGRRCKGFVSAMNT 86
>At3g03920 putative GAR1 protein
Length = 202
Score = 26.2 bits (56), Expect = 5.0
Identities = 8/24 (33%), Positives = 16/24 (66%)
Query: 16 ERQTSLGLIDEIFGQVKNPYYAVR 39
E +T +G +DEIFG + ++++
Sbjct: 91 ENKTQIGKVDEIFGPINESLFSIK 114
>At4g10740 hypothetical protein
Length = 427
Score = 25.8 bits (55), Expect = 6.5
Identities = 10/44 (22%), Positives = 21/44 (47%)
Query: 36 YAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCPMKQNFW 79
YA+RY + G R ++ F+ +F ++ +P + + W
Sbjct: 155 YALRYETNSGNRSPKILRFIDDFHHHPENPALRYETYDFDTDLW 198
>At1g73980 unknown protein
Length = 643
Score = 25.8 bits (55), Expect = 6.5
Identities = 12/38 (31%), Positives = 23/38 (59%)
Query: 92 NIIKEVVTIKTPIERMENNIKQIPLKDGSIPTMPVALG 129
N+++E V KT IER++++ ++ T+P+ LG
Sbjct: 591 NLLREYVGEKTRIERLDSSRTNSTTQNLESSTVPILLG 628
>At1g50400 hypothetical protein
Length = 310
Score = 25.8 bits (55), Expect = 6.5
Identities = 13/32 (40%), Positives = 18/32 (55%), Gaps = 1/32 (3%)
Query: 36 YAVRYNSEKGIREGTLISFVAEFVNYVMHPVS 67
YA RY ++K + G + S +NYV H VS
Sbjct: 201 YAARYETDKTVASGQIASTGVAVMNYV-HKVS 231
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.322 0.138 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,819,023
Number of Sequences: 26719
Number of extensions: 102690
Number of successful extensions: 229
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 221
Number of HSP's gapped (non-prelim): 16
length of query: 130
length of database: 11,318,596
effective HSP length: 88
effective length of query: 42
effective length of database: 8,967,324
effective search space: 376627608
effective search space used: 376627608
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC137602.4


