

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136472.7 + phase: 0
(123 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
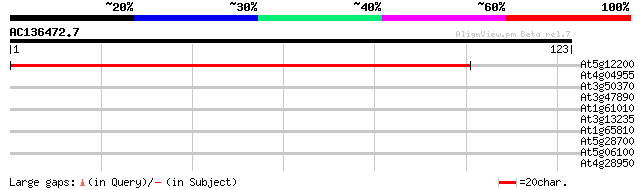
Score E
Sequences producing significant alignments: (bits) Value
At5g12200 dihydropyrimidinase like protein 174 1e-44
At4g04955 Dihydroorotase like protein 32 0.078
At3g50370 putative protein 28 1.1
At3g47890 putative protein 27 1.9
At1g61010 putative cleavage and polyadenylation specificity factor 27 2.5
At3g13235 unknown protein 26 5.6
At1g65810 hypothetical protein 26 5.6
At5g28700 putative protein 25 7.3
At5g06100 transcription factor MYB33 - like protein 25 7.3
At4g28950 rac GTP binding protein Arac7 25 9.5
>At5g12200 dihydropyrimidinase like protein
Length = 531
Score = 174 bits (441), Expect = 1e-44
Identities = 81/101 (80%), Positives = 90/101 (88%)
Query: 1 MSIDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA 60
MS+DAM+E+AKARKSGQ+VIGEPV+SGL LDD WLW PDF A+KYVMSPPIR GH KA
Sbjct: 284 MSVDAMDEIAKARKSGQKVIGEPVVSGLILDDHWLWDPDFTIASKYVMSPPIRPVGHGKA 343
Query: 61 LQAALSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNG 101
LQ ALSTG+LQLVGTDHC FNSTQKA G+DDFR+IPNGVNG
Sbjct: 344 LQDALSTGILQLVGTDHCTFNSTQKALGLDDFRRIPNGVNG 384
>At4g04955 Dihydroorotase like protein
Length = 506
Score = 32.0 bits (71), Expect = 0.078
Identities = 27/107 (25%), Positives = 49/107 (45%), Gaps = 8/107 (7%)
Query: 5 AMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAA 64
+++ + +A+ G V E LA + P+ DT ++ SPPIR + + L A
Sbjct: 299 SLDLIKEAKGKGDSVTVETCPHYLAFSAEEI--PEGDT--RFKCSPPIRDAANREKLWEA 354
Query: 65 LSTGVLQLVGTDHCVFNSTQKAFGIDDFRKIPNGVNGDHSNLRYLRP 111
L G + ++ +DH K +F K G+ S+L+++ P
Sbjct: 355 LMEGDIDMLSSDHSPTKPELKLMSDGNFLKAWGGI----SSLQFVLP 397
>At3g50370 putative protein
Length = 2152
Score = 28.1 bits (61), Expect = 1.1
Identities = 14/32 (43%), Positives = 19/32 (58%)
Query: 81 NSTQKAFGIDDFRKIPNGVNGDHSNLRYLRPK 112
N QK +DF K P GV+G S+L +RP+
Sbjct: 166 NQKQKQAAGEDFSKEPRGVSGMSSSLVDMRPQ 197
>At3g47890 putative protein
Length = 1528
Score = 27.3 bits (59), Expect = 1.9
Identities = 18/64 (28%), Positives = 28/64 (43%), Gaps = 5/64 (7%)
Query: 40 FDTAAKYVMSPPIRKQGHDKALQAALSTGVLQLVG----TDHCVFNSTQKAFGIDDFRKI 95
FD A PI ++ +K L L + + G T+HCV + T K + +
Sbjct: 1037 FDDADLQEQEKPINQEKWNKQLDD-LEGAKVNINGVFPSTNHCVISDTAKVLDVISQEVV 1095
Query: 96 PNGV 99
PNG+
Sbjct: 1096 PNGI 1099
>At1g61010 putative cleavage and polyadenylation specificity factor
Length = 693
Score = 26.9 bits (58), Expect = 2.5
Identities = 21/76 (27%), Positives = 27/76 (34%), Gaps = 14/76 (18%)
Query: 28 LALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQAALSTGVL--------QLVGTDHCV 79
L LD+ W HPD Y SP +K A T +L Q ++ V
Sbjct: 261 LILDEYWANHPDLHNIPIYYASPLAKK------CMAVYQTYILSMNDRIRNQFANSNPFV 314
Query: 80 FNSTQKAFGIDDFRKI 95
F IDDF +
Sbjct: 315 FKHISPLNSIDDFNDV 330
>At3g13235 unknown protein
Length = 414
Score = 25.8 bits (55), Expect = 5.6
Identities = 18/63 (28%), Positives = 33/63 (51%), Gaps = 4/63 (6%)
Query: 3 IDAMEEVAKARKSGQRVIGEPVLSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKALQ 62
++ +++V +AR + V+ LAL L+ FD A+ + IR++G D+ +
Sbjct: 131 LNKLQDVLRARHRQRSVLQRQKEEELAL----LYADPFDVEAQRKIEAAIRQKGIDENWE 186
Query: 63 AAL 65
AAL
Sbjct: 187 AAL 189
>At1g65810 hypothetical protein
Length = 1050
Score = 25.8 bits (55), Expect = 5.6
Identities = 20/63 (31%), Positives = 31/63 (48%), Gaps = 2/63 (3%)
Query: 25 LSGLALDDSWLWHPDFDTAAKYVMSPPIRKQGHDKA--LQAALSTGVLQLVGTDHCVFNS 82
L L LD+++L ++A+ MS PI+ D+A L+ S LQL G H +
Sbjct: 526 LQKLCLDNAYLLFCTASSSARLHMSSPIQLLVIDEAAQLKECESAIPLQLRGLQHAILIG 585
Query: 83 TQK 85
+K
Sbjct: 586 DEK 588
>At5g28700 putative protein
Length = 292
Score = 25.4 bits (54), Expect = 7.3
Identities = 10/21 (47%), Positives = 12/21 (56%)
Query: 20 IGEPVLSGLALDDSWLWHPDF 40
IGE L +A D W+WH F
Sbjct: 104 IGESTLLVVASQDLWIWHAIF 124
>At5g06100 transcription factor MYB33 - like protein
Length = 520
Score = 25.4 bits (54), Expect = 7.3
Identities = 14/46 (30%), Positives = 24/46 (51%), Gaps = 3/46 (6%)
Query: 40 FDTAAKYVMSPPIRKQGHDKALQAALSTGVLQLVGTDHCVFNSTQK 85
FDT Y+ SPP G + L + TG+L ++ + + N++ K
Sbjct: 343 FDT---YIQSPPPPTGGEESDLYSNFDTGLLDMLLLEAKIRNNSTK 385
>At4g28950 rac GTP binding protein Arac7
Length = 209
Score = 25.0 bits (53), Expect = 9.5
Identities = 10/16 (62%), Positives = 12/16 (74%)
Query: 40 FDTAAKYVMSPPIRKQ 55
FDTA K V+ PP RK+
Sbjct: 168 FDTAIKVVLQPPRRKE 183
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.320 0.137 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,795,582
Number of Sequences: 26719
Number of extensions: 105021
Number of successful extensions: 240
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 232
Number of HSP's gapped (non-prelim): 11
length of query: 123
length of database: 11,318,596
effective HSP length: 87
effective length of query: 36
effective length of database: 8,994,043
effective search space: 323785548
effective search space used: 323785548
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Medicago: description of AC136472.7


