

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135467.11 - phase: 0
(807 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
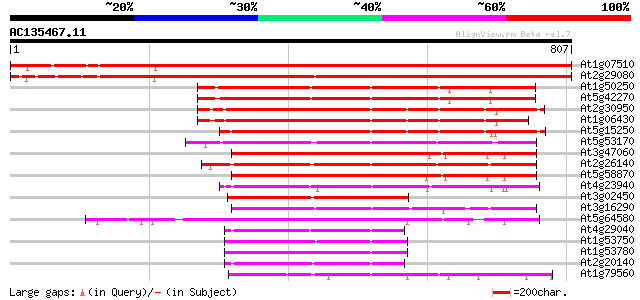
Score E
Sequences producing significant alignments: (bits) Value
At1g07510 unknown protein 1178 0.0
At2g29080 AAA-type like ATPase 1140 0.0
At1g50250 putative chloroplast FtsH protease 435 e-122
At5g42270 cell division protein FtsH 427 e-119
At2g30950 zinc dependent protease (VAR2) 404 e-113
At1g06430 cell division protease FtsH, putative 399 e-111
At5g15250 FtsH-like protein Pftf precursor - like 391 e-109
At5g53170 cell division protein FtsH protease-like 367 e-101
At3g47060 FtsH metalloprotease - like protein 362 e-100
At2g26140 FtsH like protease 357 1e-98
At5g58870 cell division protein - like 356 3e-98
At4g23940 cell division protein - like 264 1e-70
At3g02450 cell division protein FtsH-like protein 259 3e-69
At3g16290 putative FtsH-like metalloprotease 258 1e-68
At5g64580 unknown protein 248 7e-66
At4g29040 26S proteasome AAA-ATPase subunit RPT2a 195 7e-50
At1g53750 26S proteasome ATPase subunit 195 7e-50
At1g53780 hypothetical protein 195 1e-49
At2g20140 26S proteasome subunit 4 193 3e-49
At1g79560 FtsH like cell division protein 192 5e-49
>At1g07510 unknown protein
Length = 813
Score = 1178 bits (3047), Expect = 0.0
Identities = 608/819 (74%), Positives = 679/819 (82%), Gaps = 18/819 (2%)
Query: 1 MIFSRIGRSLSRSSRVKNLLHGET------RLGTLYGVSRTNVFVDDVEKGLGFVRGYVS 54
MIFS++G SL+RSSR K ++G G L V+ V+ GLGF+R + +
Sbjct: 1 MIFSKLGSSLARSSRSKGFVYGGGVRSAVFNQGRLRAPQNLEAAVNQVDGGLGFLRRHFA 60
Query: 55 SAIARNNGFGSNLYDFKSIAANRMLHRMFSSESPKKKNYEKFYPKEKKEVPKGEEKKSES 114
S AR G D AN L R FSS++PKKKNYE +YPK+ K+ PK E+K SES
Sbjct: 61 SFAARK---GLEAGDLSRAFANPRLRRFFSSQTPKKKNYENYYPKDSKKAPKNEQK-SES 116
Query: 115 KDESKSNTEDGGSFHEAFIKQFQNYLTPLLVVGLFLSSLSLGPRDQQQISFQEFKNKLLE 174
+D SK N + +AF ++QN L PL+ + L LS+ SLG R+QQQISFQEFKNKLLE
Sbjct: 117 RDGSKKNENENAG--DAFSNEYQNMLIPLMAIALILSTFSLGSREQQQISFQEFKNKLLE 174
Query: 175 PGLVDHIVVSNKSVAKIYVRNSPLNQADSEV-QGT---LPAKGSGGQYKYIINIGSVESF 230
GLVDHI VSNK VAK+YVR+SP +Q EV QG +PAKG GGQYKY NIGSVESF
Sbjct: 175 AGLVDHIDVSNKEVAKVYVRSSPKSQTTEEVVQGPGNGVPAKGRGGQYKYYFNIGSVESF 234
Query: 231 EEKLEEAQEALGVDSHNFVPVTYSSEMVWYQELMRFAPTLLLLGTLWFMGRKMQGGFGVG 290
EEKLEEAQEA+GV+SH+FVPVTY SE +WYQEL+RFAPTLLL+ TL F R+MQGG G
Sbjct: 235 EEKLEEAQEAIGVNSHDFVPVTYVSETIWYQELLRFAPTLLLVATLIFGARRMQGGLGGL 294
Query: 291 GGSTGKGSRGIFNIGKAHVTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEE 350
GG GK RGIFNIGKA +T+ DKN+KNK+YFKDVAGCEEAKQEIMEFVHFL+NPKKYE+
Sbjct: 295 GGPGGKAGRGIFNIGKAQITRADKNSKNKIYFKDVAGCEEAKQEIMEFVHFLQNPKKYED 354
Query: 351 LGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQE 410
LGAKIPKGALLVGPPGTGKTLLAKATAGES VPFLSISGSDFMEMFVGVGPSRVRNLFQE
Sbjct: 355 LGAKIPKGALLVGPPGTGKTLLAKATAGESAVPFLSISGSDFMEMFVGVGPSRVRNLFQE 414
Query: 411 ARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTN 470
ARQCAPSI+FIDEIDAIGR RGRGGFSG NDERESTLNQLLVEMDGFGTTAGVVVLAGTN
Sbjct: 415 ARQCAPSIIFIDEIDAIGRARGRGGFSGGNDERESTLNQLLVEMDGFGTTAGVVVLAGTN 474
Query: 471 RADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGF 530
R DILD ALLRPGRFDR I+ID PDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGF
Sbjct: 475 RPDILDKALLRPGRFDRQITIDKPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGF 534
Query: 531 AGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEA 590
AGADIANVCNEAALIAAR + + VTM HF++AIDR+IGGLEKKNRVISK ERRTVAYHE+
Sbjct: 535 AGADIANVCNEAALIAARHEGATVTMAHFDSAIDRVIGGLEKKNRVISKLERRTVAYHES 594
Query: 591 GHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSENLLRTKEQLLDMTCMTLGGRAAE 650
GHAVAGWFLEH EPLLKVTIVPRGTAALGFAQYVP+ENLL TKEQL DMTCMTLGGRAAE
Sbjct: 595 GHAVAGWFLEHAEPLLKVTIVPRGTAALGFAQYVPNENLLMTKEQLFDMTCMTLGGRAAE 654
Query: 651 QVLIGAISTGAQNDLEKVTKMTYAQVAIYGFSEKVGLLSFPQNEDQFGKPYSGDTGNIID 710
QVLIG ISTGAQNDLEKVTKMTYAQVA+YGFS+K+GLLSFPQ ED+F KPYS TG +ID
Sbjct: 655 QVLIGRISTGAQNDLEKVTKMTYAQVAVYGFSDKIGLLSFPQREDEFSKPYSNRTGAMID 714
Query: 711 QEVRDWVNHAYERTVQLIEEHKEKLAQIAELLLEKEVLHQEDLVRILGERPFKSAEPTNY 770
+EVR+WV AY+RTV+LIEEHKE++AQIAELLLEKEVLHQ+DL ++LGERPFKS E TNY
Sbjct: 715 EEVREWVGKAYKRTVELIEEHKEQVAQIAELLLEKEVLHQDDLTKVLGERPFKSGETTNY 774
Query: 771 DRFKLGFQDEEKAA--ETTVDEAEEGSGSSPLEPEVVPT 807
DRFK GF++ EK + E+ + E G PLEP+VVPT
Sbjct: 775 DRFKSGFEESEKESQKESVPVKPVEDDGIPPLEPQVVPT 813
>At2g29080 AAA-type like ATPase
Length = 809
Score = 1140 bits (2948), Expect = 0.0
Identities = 598/820 (72%), Positives = 676/820 (81%), Gaps = 27/820 (3%)
Query: 1 MIFSRIGRSLSRSSRVKNLLHG-----ETRLGTLYGVSRTNVFVDDVEKGLGFVRGYVSS 55
+ FS++ RS+SRS K L+G RL T G+ +V ++VE GLGF+R + +S
Sbjct: 4 IFFSKLNRSISRS---KGFLYGGGVRSAARLLTSPGLEAASV--NEVEGGLGFIRRHFAS 58
Query: 56 AIARNNGFGSNLYDFKSIAANRMLHRMFSSESPKKKNYEKFYPKEKKEVPKGEEKKSESK 115
+A G +N D + AN L R FS E+PKKKNYE ++PK+K+E PK ++K +
Sbjct: 59 -LASRKGLVNN--DLIGVFANPRLRRFFSDEAPKKKNYENYFPKDKQE-PKSDQKSEHKE 114
Query: 116 DESKSNTEDGGSFHEAFIKQFQNYLTPLLVVGLFLSSLSLGPRDQQQISFQEFKNKLLEP 175
K+ E+ G + F+ +FQN L PLL + +F S+ S G +QQQISFQEFKNKLLEP
Sbjct: 115 GSEKNENENVG---DMFMNRFQNLLIPLLALAVFFSTFSFGSGEQQQISFQEFKNKLLEP 171
Query: 176 GLVDHIVVSNKSVAKIYVRNSPLNQADSEVQ----GTLPAKGSGGQYKYIINIGSVESFE 231
GLVDHI VSNKSVAK+YVR++P +Q ++V +PAK +GGQYKY NIGSV+SFE
Sbjct: 172 GLVDHIDVSNKSVAKVYVRSTPKDQQTTDVVHGNGNGIPAKRTGGQYKYYFNIGSVDSFE 231
Query: 232 EKLEEAQEALGVDSHNFVPVTYSSEMVWYQELMRFAPTLLLLGTLWFMGRKMQGGFGVGG 291
EKLEEAQEALGVD H +VPVTY SEMVWYQE MRFAPTLLLLGTL + R+MQGG GVGG
Sbjct: 232 EKLEEAQEALGVDRHEYVPVTYVSEMVWYQEFMRFAPTLLLLGTLIYGARRMQGGLGVGG 291
Query: 292 GSTGKGSRGIFNIGKAHVTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEEL 351
+ GK RGIFNIGKA +T+ DK++KNK+YFKDVAGC+EAKQEIMEFVHFLKNPKKYE+L
Sbjct: 292 -TGGKNGRGIFNIGKATITRADKHSKNKIYFKDVAGCDEAKQEIMEFVHFLKNPKKYEDL 350
Query: 352 GAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEA 411
GAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVR+LFQEA
Sbjct: 351 GAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRHLFQEA 410
Query: 412 RQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNR 471
RQ APSI+FIDEIDAIGR RGRGG G NDERESTLNQLLVEMDGFGTTAGVVVLAGTNR
Sbjct: 411 RQAAPSIIFIDEIDAIGRARGRGGLGG-NDERESTLNQLLVEMDGFGTTAGVVVLAGTNR 469
Query: 472 ADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFA 531
DILD ALLRPGRFDR I+ID PDIKGRDQIF+IYLKKIKLDHEPSYYSQRLAALTPGFA
Sbjct: 470 PDILDKALLRPGRFDRQITIDKPDIKGRDQIFKIYLKKIKLDHEPSYYSQRLAALTPGFA 529
Query: 532 GADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAG 591
GADIANVCNEAALIAAR + + VTM HFE+AIDR+IGGLEKKNRVISK ERRTVAYHE+G
Sbjct: 530 GADIANVCNEAALIAARHEGATVTMAHFESAIDRVIGGLEKKNRVISKLERRTVAYHESG 589
Query: 592 HAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSENLLRTKEQLLDMTCMTLGGRAAEQ 651
HAV GWFLEH EPLLKVTIVPRGTAALGFAQYVP+ENLL TKEQL DMTCMTLGGRAAEQ
Sbjct: 590 HAVVGWFLEHAEPLLKVTIVPRGTAALGFAQYVPNENLLMTKEQLFDMTCMTLGGRAAEQ 649
Query: 652 VLIGAISTGAQNDLEKVTKMTYAQVAIYGFSEKVGLLSFPQNED--QFGKPYSGDTGNII 709
VLIG ISTGAQNDLEKVTKMTYAQVA+YGFS+KVGLLSFP +D F KPYS TG II
Sbjct: 650 VLIGKISTGAQNDLEKVTKMTYAQVAVYGFSDKVGLLSFPPRDDGYDFSKPYSNKTGAII 709
Query: 710 DQEVRDWVNHAYERTVQLIEEHKEKLAQIAELLLEKEVLHQEDLVRILGERPFKSAEPTN 769
D+EVRDWV AYERTV+L+EEHK K+A+IAELLLEKEVLHQ+DL++ILGERPFKSAE TN
Sbjct: 710 DEEVRDWVAKAYERTVELVEEHKVKVAEIAELLLEKEVLHQDDLLKILGERPFKSAEVTN 769
Query: 770 YDRFKLGFQDEEK--AAETTVDEAEEGSGSSPLEPEVVPT 807
YDRFK GF++ EK AA TV+ + P EP+VVPT
Sbjct: 770 YDRFKSGFEETEKDSAATPTVEPVVDDGAPPPFEPQVVPT 809
>At1g50250 putative chloroplast FtsH protease
Length = 716
Score = 435 bits (1119), Expect = e-122
Identities = 235/501 (46%), Positives = 332/501 (65%), Gaps = 23/501 (4%)
Query: 270 LLLLGTLWFMGRKMQGGFGVGGGSTGKGSRGIFNIGKAHVTKVDKNTKNKVYFKDVAGCE 329
LL G L+ + R+ QGG G G G G G + G++ +K + + V F DVAG +
Sbjct: 214 LLAFGGLFLLFRRAQGGPGGGPGGLG----GPMDFGRSK-SKFQEVPETGVSFADVAGAD 268
Query: 330 EAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISG 389
+AK E+ E V FLKNP KY LGAKIPKG LLVGPPGTGKTLLA+A AGE+GVPF S +
Sbjct: 269 QAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGEAGVPFFSCAA 328
Query: 390 SDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQ 449
S+F+E+FVGVG SRVR+LF++A+ AP IVFIDEIDA+GR+RG G G NDERE T+NQ
Sbjct: 329 SEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRG-AGMGGGNDEREQTINQ 387
Query: 450 LLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKK 509
LL EMDGF +GV+VLA TNR D+LD+ALLRPGRFDR +++D PD+ GR +I Q++ +
Sbjct: 388 LLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRVKILQVHSRG 447
Query: 510 IKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGG 569
L + + ++A TPGF GAD+ N+ NEAA++AAR + +++ D A++RII G
Sbjct: 448 KALGKDVDF--DKVARRTPGFTGADLQNLMNEAAILAARRELKEISKDEISDALERIIAG 505
Query: 570 LEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSENL 629
EKKN V+S+ ++R VAYHEAGHA+ G + +P+ K++I+PRG A G + PSE
Sbjct: 506 PEKKNAVVSEEKKRLVAYHEAGHALVGALMPEYDPVAKISIIPRGQAG-GLTFFAPSEER 564
Query: 630 LR----TKEQLLDMTCMTLGGRAAEQVLIG--AISTGAQNDLEKVTKMTYAQVAIYGFSE 683
L ++ L + + LGGR AE+V+ G ++TGA ND +V+++ + +GFS+
Sbjct: 565 LESGLYSRSYLENQMAVALGGRVAEEVIFGDENVTTGASNDFMQVSRVARQMIERFGFSK 624
Query: 684 KVGLLS--------FPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEKL 735
K+G ++ F + K YS T +I+D EVR+ V AY+R ++I H + L
Sbjct: 625 KIGQVAVGGPGGNPFMGQQMSSQKDYSMATADIVDAEVRELVEKAYKRATEIITTHIDIL 684
Query: 736 AQIAELLLEKEVLHQEDLVRI 756
++A+LL+EKE + E+ + +
Sbjct: 685 HKLAQLLIEKETVDGEEFMSL 705
>At5g42270 cell division protein FtsH
Length = 704
Score = 427 bits (1097), Expect = e-119
Identities = 233/501 (46%), Positives = 329/501 (65%), Gaps = 24/501 (4%)
Query: 270 LLLLGTLWFMGRKMQGGFGVGGGSTGKGSRGIFNIGKAHVTKVDKNTKNKVYFKDVAGCE 329
LL G L+++ R QGG G GG G + G++ +K + + V F DVAG +
Sbjct: 203 LLAFGGLFYLFRGGQGGAGGPGGLGGP-----MDFGRSK-SKFQEVPETGVTFGDVAGAD 256
Query: 330 EAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISG 389
+AK E+ E V FLKNP KY LGAKIPKG LLVGPPGTGKTLLA+A AGE+GVPF S +
Sbjct: 257 QAKLELQEVVDFLKNPDKYTALGAKIPKGCLLVGPPGTGKTLLARAVAGEAGVPFFSCAA 316
Query: 390 SDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQ 449
S+F+E+FVGVG SRVR+LF++A+ AP IVFIDEIDA+GR+RG G G NDERE T+NQ
Sbjct: 317 SEFVELFVGVGASRVRDLFEKAKSKAPCIVFIDEIDAVGRQRG-AGMGGGNDEREQTINQ 375
Query: 450 LLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKK 509
LL EMDGF +GV+VLA TNR D+LD+ALLRPGRFDR +++D PD+ GR QI +++ +
Sbjct: 376 LLTEMDGFSGNSGVIVLAATNRPDVLDSALLRPGRFDRQVTVDRPDVAGRVQILKVHSRG 435
Query: 510 IKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGG 569
+ + Y +++A TPGF GAD+ N+ NEAA++AAR + +++ D A++RII G
Sbjct: 436 KAIGKDVDY--EKVARRTPGFTGADLQNLMNEAAILAARRELKEISKDEISDALERIIAG 493
Query: 570 LEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSENL 629
EKKN V+S+ ++R VAYHEAGHA+ G + +P+ K++I+PRG A G + PSE
Sbjct: 494 PEKKNAVVSEEKKRLVAYHEAGHALVGALMPEYDPVAKISIIPRGQAG-GLTFFAPSEER 552
Query: 630 LR----TKEQLLDMTCMTLGGRAAEQVLIG--AISTGAQNDLEKVTKMTYAQVAIYGFSE 683
L ++ L + + LGGR AE+V+ G ++TGA ND +V+++ V +GFS+
Sbjct: 553 LESGLYSRSYLENQMAVALGGRVAEEVIFGDENVTTGASNDFMQVSRVARQMVERFGFSK 612
Query: 684 KVGLLS--------FPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEKL 735
K+G ++ F K YS T +++D EVR+ V AY R ++I + L
Sbjct: 613 KIGQVAVGGAGGNPFLGQSMSSQKDYSMATADVVDAEVRELVEKAYVRAKEIITTQIDIL 672
Query: 736 AQIAELLLEKEVLHQEDLVRI 756
++A+LL+EKE + E+ + +
Sbjct: 673 HKLAQLLIEKETVDGEEFMSL 693
>At2g30950 zinc dependent protease (VAR2)
Length = 695
Score = 404 bits (1039), Expect = e-113
Identities = 234/510 (45%), Positives = 321/510 (62%), Gaps = 25/510 (4%)
Query: 271 LLLGTLWFMGRKMQGGFGVGGGSTGKGSRGIFNIGKAHVTKVDKNTKNKVYFKDVAGCEE 330
LL+G L+ + R+ GG G G G G+ F KA K V F DVAG +E
Sbjct: 181 LLIGGLFLLSRRSGGGMG---GPGGPGNPLQFGQSKA---KFQMEPNTGVTFDDVAGVDE 234
Query: 331 AKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGS 390
AKQ+ ME V FLK P+++ +GAKIPKG LL+GPPGTGKTLLAKA AGE+GVPF SISGS
Sbjct: 235 AKQDFMEVVEFLKKPERFTAVGAKIPKGVLLIGPPGTGKTLLAKAIAGEAGVPFFSISGS 294
Query: 391 DFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQL 450
+F+EMFVGVG SRVR+LF++A++ AP IVF+DEIDA+GR+RG G G NDERE TLNQL
Sbjct: 295 EFVEMFVGVGASRVRDLFKKAKENAPCIVFVDEIDAVGRQRGT-GIGGGNDEREQTLNQL 353
Query: 451 LVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKI 510
L EMDGF GV+V+A TNRADILD+ALLRPGRFDR +S+DVPD+KGR I +++
Sbjct: 354 LTEMDGFEGNTGVIVVAATNRADILDSALLRPGRFDRQVSVDVPDVKGRTDILKVHAGNK 413
Query: 511 KLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGL 570
K D++ S + +A TPGF+GAD+AN+ NEAA++A R + ++ + +IDRI+ G+
Sbjct: 414 KFDNDVSL--EIIAMRTPGFSGADLANLLNEAAILAGRRARTSISSKEIDDSIDRIVAGM 471
Query: 571 EKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSEN-L 629
E + + + VAYHE GHAV G + + KVT++PRG A G ++PS++
Sbjct: 472 E-GTVMTDGKSKSLVAYHEVGHAVCGTLTPGHDAVQKVTLIPRGQAR-GLTWFIPSDDPT 529
Query: 630 LRTKEQLLDMTCMTLGGRAAEQVLIG--AISTGAQNDLEKVTKMTYAQVAIYGFSEKVGL 687
L +K+QL LGGRAAE+++ G ++TGA DL+++T + V +G S+ +G
Sbjct: 530 LISKQQLFARIVGGLGGRAAEEIIFGDSEVTTGAVGDLQQITGLARQMVTTFGMSD-IGP 588
Query: 688 LSFPQNEDQFG--------KPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEKLAQIA 739
S + Q S ID V+ + AYE + I+ ++E + ++
Sbjct: 589 WSLMDSSAQSDVIMRMMARNSMSEKLAEDIDSAVKKLSDSAYEIALSHIKNNREAMDKLV 648
Query: 740 ELLLEKEVLHQEDLVRILGERPFKSAEPTN 769
E+LLEKE + ++ IL E F P N
Sbjct: 649 EVLLEKETIGGDEFRAILSE--FTEIPPEN 676
>At1g06430 cell division protease FtsH, putative
Length = 649
Score = 399 bits (1026), Expect = e-111
Identities = 228/489 (46%), Positives = 314/489 (63%), Gaps = 25/489 (5%)
Query: 270 LLLLGTLWFMGRKMQGGFGVGGGSTGKGSRGI-FNIGKAHVTKVDKNTKNKVYFKDVAGC 328
++L+G L+ + R+ GG G G G G IG++ K V F DVAG
Sbjct: 173 VILIGGLFLLSRRSSGGMG------GPGGPGFPLQIGQSKA-KFQMEPNTGVTFDDVAGV 225
Query: 329 EEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSIS 388
+EAKQ+ ME V FLK P+++ +GA+IPKG LLVGPPGTGKTLLAKA AGE+GVPF SIS
Sbjct: 226 DEAKQDFMEVVEFLKKPERFTAVGARIPKGVLLVGPPGTGKTLLAKAIAGEAGVPFFSIS 285
Query: 389 GSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLN 448
GS+F+EMFVGVG SRVR+LF++A++ AP IVF+DEIDA+GR+RG G G NDERE TLN
Sbjct: 286 GSEFVEMFVGVGASRVRDLFKKAKENAPCIVFVDEIDAVGRQRGT-GIGGGNDEREQTLN 344
Query: 449 QLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLK 508
QLL EMDGF GV+V+A TNRADILD+ALLRPGRFDR +S+DVPD+KGR I +++
Sbjct: 345 QLLTEMDGFEGNTGVIVVAATNRADILDSALLRPGRFDRQVSVDVPDVKGRTDILKVHSG 404
Query: 509 KIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIG 568
K + S + +A TPGF+GAD+AN+ NEAA++A R ++ ++ + +IDRI+
Sbjct: 405 NKKFESGVSL--EVIAMRTPGFSGADLANLLNEAAILAGRRGKTAISSKEIDDSIDRIVA 462
Query: 569 GLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSEN 628
G+E + + + VAYHE GHA+ G + + KVT++PRG A G ++PS++
Sbjct: 463 GME-GTVMTDGKSKSLVAYHEVGHAICGTLTPGHDAVQKVTLIPRGQAR-GLTWFIPSDD 520
Query: 629 -LLRTKEQLLDMTCMTLGGRAAEQVLIG--AISTGAQNDLEKVTKMTYAQVAIYGFSEKV 685
L +K+QL LGGRAAE+V+ G ++TGA +DL+++T + V +G SE +
Sbjct: 521 PTLISKQQLFARIVGGLGGRAAEEVIFGESEVTTGAVSDLQQITGLAKQMVTTFGMSE-I 579
Query: 686 GLLSFPQNEDQFG--------KPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEKLAQ 737
G S + +Q S N ID V+ + AYE + I ++E + +
Sbjct: 580 GPWSLMDSSEQSDVIMRMMARNSMSEKLANDIDTAVKTLSDKAYEIALSQIRNNREAMDK 639
Query: 738 IAELLLEKE 746
I E+LLEKE
Sbjct: 640 IVEILLEKE 648
>At5g15250 FtsH-like protein Pftf precursor - like
Length = 687
Score = 391 bits (1005), Expect = e-109
Identities = 225/482 (46%), Positives = 313/482 (64%), Gaps = 23/482 (4%)
Query: 302 FNIGKAHVTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALL 361
F +G +++ NT + F+DVAG +EAKQ+ E V FLK P+K+ LGAKIPKG LL
Sbjct: 204 FGLGSKAKFQMEPNTG--ITFEDVAGVDEAKQDFEEIVEFLKTPEKFSALGAKIPKGVLL 261
Query: 362 VGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFI 421
GPPGTGKTLLAKA AGE+GVPF S+SGS+F+EMFVGVG SR R+LF +A+ +P IVFI
Sbjct: 262 TGPPGTGKTLLAKAIAGEAGVPFFSLSGSEFIEMFVGVGASRARDLFNKAKANSPCIVFI 321
Query: 422 DEIDAIGRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLR 481
DEIDA+GR RG G G NDERE TLNQ+L EMDGF GV+V+A TNR +ILD+ALLR
Sbjct: 322 DEIDAVGRMRGT-GIGGGNDEREQTLNQILTEMDGFAGNTGVIVIAATNRPEILDSALLR 380
Query: 482 PGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNE 541
PGRFDR +S+ +PDI+GR++I +++ + KLD + S +A TPGF+GAD+AN+ NE
Sbjct: 381 PGRFDRQVSVGLPDIRGREEILKVHSRSKKLDKDVSL--SVIAMRTPGFSGADLANLMNE 438
Query: 542 AALIAARTDESQVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEH 601
AA++A R + ++T+ + +IDRI+ G+E ++I + + VAYHE GHA+ E
Sbjct: 439 AAILAGRRGKDKITLTEIDDSIDRIVAGME-GTKMIDGKSKAIVAYHEVGHAICATLTEG 497
Query: 602 CEPLLKVTIVPRGTAALGFAQYVPSEN-LLRTKEQLLDMTCMTLGGRAAEQVLIG--AIS 658
+P+ KVT+VPRG A G ++P E+ L +K+QL LGGRAAE V+ G I+
Sbjct: 498 HDPVQKVTLVPRGQAR-GLTWFLPGEDPTLVSKQQLFARIVGGLGGRAAEDVIFGEPEIT 556
Query: 659 TGAQNDLEKVTKMTYAQVAIYGFSEKVG--LLSFP---QNEDQF----GKPYSGDTGNII 709
TGA DL++VT++ V ++G SE +G L+ P QN+ S I
Sbjct: 557 TGAAGDLQQVTEIARQMVTMFGMSE-IGPWALTDPAVKQNDVVLRMLARNSMSEKLAEDI 615
Query: 710 DQEVRDWVNHAYERTVQLIEEHKEKLAQIAELLLEKEVLHQEDLVRILGERPFKSAEPTN 769
D V+ + AYE + + ++E + ++ ++LLEKE L ++ IL E + +P N
Sbjct: 616 DSCVKKIIGDAYEVAKKHVRNNREAIDKLVDVLLEKETLTGDEFRAILSE---YTDQPLN 672
Query: 770 YD 771
D
Sbjct: 673 TD 674
>At5g53170 cell division protein FtsH protease-like
Length = 806
Score = 367 bits (941), Expect = e-101
Identities = 220/513 (42%), Positives = 310/513 (59%), Gaps = 29/513 (5%)
Query: 254 SSEMVWYQELMRFAPTLLLLGTLWFMG----RKMQGGFGVGGGSTGKGSRGIFNIGKAHV 309
S++ + QEL+ + +G +W MG +K G G G G++G GS ++ +
Sbjct: 292 SNKSRFAQELVSTILFTVAVGLVWIMGAAALQKYIGSLG-GIGTSGVGSSSSYS--PKEL 348
Query: 310 TKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGK 369
K KN FKDV GC++AKQE+ E V +LKNP K+ LG K+PKG LL G PGTGK
Sbjct: 349 NKEITPEKNVKTFKDVKGCDDAKQELEEVVEYLKNPSKFTRLGGKLPKGILLTGAPGTGK 408
Query: 370 TLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIG- 428
TLLAKA AGE+GVPF +GS+F EMFVGVG RVR+LFQ A++ AP I+FIDEIDA+G
Sbjct: 409 TLLAKAIAGEAGVPFFYRAGSEFEEMFVGVGARRVRSLFQAAKKKAPCIIFIDEIDAVGS 468
Query: 429 -RKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
RK+ G + TL+QLLVEMDGF G++V+A TN DILD AL RPGRFDR
Sbjct: 469 TRKQWEG-------HTKKTLHQLLVEMDGFEQNEGIIVMAATNLPDILDPALTRPGRFDR 521
Query: 488 TISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAA 547
I + PD++GR++I ++YL+ + + + +A TPGF GAD+AN+ N AA+ AA
Sbjct: 522 HIVVPSPDVRGREEILELYLQGKPMSEDVDV--KAIARGTPGFNGADLANLVNIAAIKAA 579
Query: 548 RTDESQVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLK 607
+++ + E A DRI+ G E+K +S+ ++ AYHE+GHA+ + P+ K
Sbjct: 580 VEGAEKLSSEQLEFAKDRIVMGTERKTMFVSEDSKKLTAYHESGHAIVALNTKGAHPIHK 639
Query: 608 VTIVPRGTAALGFAQYVPS-ENLLRTKEQLLDMTCMTLGGRAAEQVLIGA--ISTGAQND 664
TI+PRG +ALG +PS + +K QLL + +GGR AE+++ G I+TGA +D
Sbjct: 640 ATIMPRG-SALGMVTQLPSNDETSVSKRQLLARLDVCMGGRVAEELIFGLDHITTGASSD 698
Query: 665 LEKVTKMTYAQVAIYGFSEKVGLLSFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERT 724
L + T++ V+ G SE +G + + S D + ID EV + AYER
Sbjct: 699 LSQATELAQYMVSSCGMSEAIGPVHIKERP-------SSDMQSRIDAEVVKLLREAYERV 751
Query: 725 VQLIEEHKEKLAQIAELLLEKEVLHQEDLVRIL 757
L++ H+++L +A LLE E L ED+ RIL
Sbjct: 752 KSLLKRHEKQLHTLANALLEYETLTAEDIKRIL 784
>At3g47060 FtsH metalloprotease - like protein
Length = 802
Score = 362 bits (930), Expect = e-100
Identities = 207/454 (45%), Positives = 292/454 (63%), Gaps = 17/454 (3%)
Query: 320 VYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGE 379
+ F DVAG +EAK+E+ E V FL+NP+KY LGA+ P+G LLVG PGTGKTLLAKA AGE
Sbjct: 322 ITFADVAGVDEAKEELEEIVEFLRNPEKYVRLGARPPRGVLLVGLPGTGKTLLAKAVAGE 381
Query: 380 SGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGS 439
+ VPF+S S S+F+E++VG+G SRVR+LF A++ APSI+FIDEIDA+ + R GS
Sbjct: 382 AEVPFISCSASEFVELYVGMGASRVRDLFARAKKEAPSIIFIDEIDAVAKSRDGKFRMGS 441
Query: 440 NDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGR 499
NDERE TLNQLL EMDGF + + V+VL TNRAD+LD AL RPGRFDR ++++ PD GR
Sbjct: 442 NDEREQTLNQLLTEMDGFDSNSAVIVLGATNRADVLDPALRRPGRFDRVVTVETPDKIGR 501
Query: 500 DQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHF 559
+ I ++++ K +L +A++T GF GAD+AN+ NEAAL+A R +++ V F
Sbjct: 502 ESILRVHVSKKELPLGDDVNLGSIASMTTGFTGADLANLVNEAALLAGRKNKTNVEKIDF 561
Query: 560 EAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHC---EPLL-KVTIVPRGT 615
A++R I G+EKK+ + E+ VA HEAGHAV G + + +P + K++I+PR
Sbjct: 562 IQAVERSIAGIEKKSARLKGNEKAVVARHEAGHAVVGTAVANLLTGQPRVEKLSILPRTG 621
Query: 616 AALGFAQYVP---SENLLRTKEQLLDMTCMTLGGRAAEQVLI-GAISTGAQNDLEKVTKM 671
ALGF Y+P + L ++LL LGGRAAE+V+ G ISTGA +D+ + T M
Sbjct: 622 GALGFT-YIPPTSEDRYLLFIDELLGRLVTLLGGRAAEEVVYSGRISTGAFDDIRRATDM 680
Query: 672 TYAQVAIYGFSEKVG-----LLSFPQNEDQFGKPYSGDTGNIID---QEVRDWVNHAYER 723
Y VA YG ++K+G LS +D G P+ D G ++D +EV + A +
Sbjct: 681 AYKAVAEYGLNQKIGPVSVATLSGGGIDDSGGSPWGRDQGKLVDLVQKEVTILLQSALDV 740
Query: 724 TVQLIEEHKEKLAQIAELLLEKEVLHQEDLVRIL 757
+ ++ + + L + L EKE + E+L + L
Sbjct: 741 ALSVVRANPDVLEGLGAQLEEKEKVEGEELQKWL 774
>At2g26140 FtsH like protease
Length = 717
Score = 357 bits (916), Expect = 1e-98
Identities = 207/489 (42%), Positives = 296/489 (60%), Gaps = 20/489 (4%)
Query: 276 LWFMGRKMQGGF----GVGGGSTGKGSRGIFNIGKAHVTKVDKNTKNKVYFKDVAGCEEA 331
LW R + GF G+G +G + + +D +TK F DV G +EA
Sbjct: 180 LWSTIRTIGVGFLLISGIGALIEDRGIGKGLGLHEEVQPSMDSSTK----FSDVKGVDEA 235
Query: 332 KQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFLSISGSD 391
K E+ E VH+L++PK++ LG K+PKG LLVGPPGTGKT+LA+A AGE+GVPF S SGS+
Sbjct: 236 KAELEEIVHYLRDPKRFTRLGGKLPKGVLLVGPPGTGKTMLARAIAGEAGVPFFSCSGSE 295
Query: 392 FMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERESTLNQLL 451
F EMFVGVG RVR+LF A++C+P I+FIDEIDAIG R + TLNQ+L
Sbjct: 296 FEEMFVGVGARRVRDLFSAAKKCSPCIIFIDEIDAIGGSRN----PKDQQYMKMTLNQML 351
Query: 452 VEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIK 511
VE+DGF G++V+A TN + LD AL+RPGRFDR I + PD++GR QI + ++ K+
Sbjct: 352 VELDGFKQNEGIIVVAATNFPESLDKALVRPGRFDRHIVVPNPDVEGRRQILESHMSKVL 411
Query: 512 LDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGLE 571
+ +A TPGF+GAD+AN+ N AAL AA VTM E A DRI+ G E
Sbjct: 412 KAEDVDL--MIIARGTPGFSGADLANLVNVAALKAAMDGSKDVTMSDLEFAKDRIMMGSE 469
Query: 572 KKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAALGFAQYVPSENLLR 631
+K+ VIS R+ A+HE GHA+ E P+ K TIVPRG ALG +P ++
Sbjct: 470 RKSAVISDESRKLTAFHEGGHALVAIHTEGALPVHKATIVPRG-MALGMVSQLPDKDETS 528
Query: 632 -TKEQLLDMTCMTLGGRAAEQVLIG--AISTGAQNDLEKVTKMTYAQVAIYGFSEKVGLL 688
+++Q+L + +GGR AE+++ G +++GA +DLE+ TK+ A V +G S++VGL+
Sbjct: 529 ISRKQMLARLDVCMGGRVAEELIFGESEVTSGASSDLEQATKLARAMVTKFGMSKEVGLV 588
Query: 689 SFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEKLAQIAELLLEKEVL 748
+ N D GK S +T +I+ EV+ + AY ++ + ++L +A LL+ E L
Sbjct: 589 A--HNYDDNGKSMSTETRLLIESEVKQLLEKAYNNAKTILTVYNKELHALANALLQHETL 646
Query: 749 HQEDLVRIL 757
+ + +L
Sbjct: 647 SGKQIKELL 655
>At5g58870 cell division protein - like
Length = 806
Score = 356 bits (914), Expect = 3e-98
Identities = 205/454 (45%), Positives = 286/454 (62%), Gaps = 17/454 (3%)
Query: 320 VYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGE 379
+ F DVAG +EAK+E+ E V FLKNP +Y LGA+ P+G LLVG PGTGKTLLAKA AGE
Sbjct: 326 ITFADVAGVDEAKEELEEIVEFLKNPDRYVRLGARPPRGVLLVGLPGTGKTLLAKAVAGE 385
Query: 380 SGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGS 439
S VPF+S S S+F+E++VG+G SRVR+LF A++ APSI+FIDEIDA+ + R S
Sbjct: 386 SDVPFISCSASEFVELYVGMGASRVRDLFARAKKEAPSIIFIDEIDAVAKSRDGKFRMVS 445
Query: 440 NDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGR 499
NDERE TLNQLL EMDGF +++ V+VL TNRAD+LD AL RPGRFDR ++++ PD GR
Sbjct: 446 NDEREQTLNQLLTEMDGFDSSSAVIVLGATNRADVLDPALRRPGRFDRVVTVESPDKVGR 505
Query: 500 DQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHF 559
+ I ++++ K +L +A++T GF GAD+AN+ NEAAL+A R + V F
Sbjct: 506 ESILKVHVSKKELPLGDDVNLASIASMTTGFTGADLANLVNEAALLAGRKSKMTVDKIDF 565
Query: 560 EAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGW----FLEHCEPLLKVTIVPRGT 615
A++R I G+EKK + E+ VA HEAGHAV G L + K++I+PR
Sbjct: 566 IHAVERSIAGIEKKTARLKGSEKAVVARHEAGHAVVGTAVASLLSGQSRVEKLSILPRSG 625
Query: 616 AALGFAQYVP---SENLLRTKEQLLDMTCMTLGGRAAEQVLI-GAISTGAQNDLEKVTKM 671
ALGF Y+P + L ++L LGGRAAE+V+ G ISTGA +D+ + T M
Sbjct: 626 GALGFT-YIPPTHEDRYLLFIDELHGRLVTLLGGRAAEEVVYSGRISTGALDDIRRATDM 684
Query: 672 TYAQVAIYGFSEKVG-----LLSFPQNEDQFGKPYSGDTGNIID---QEVRDWVNHAYER 723
Y VA YG +EK+G LS +D G P+ D G+++D +EV + + A +
Sbjct: 685 AYKAVAEYGLNEKIGPVSVATLSAGGIDDSGGSPWGRDQGHLVDLVQREVTNLLQSALDV 744
Query: 724 TVQLIEEHKEKLAQIAELLLEKEVLHQEDLVRIL 757
+ ++ + + L + L ++E + E+L + L
Sbjct: 745 ALTVVRANPDVLEGLGAQLEDEEKVEGEELQKWL 778
>At4g23940 cell division protein - like
Length = 946
Score = 264 bits (675), Expect = 1e-70
Identities = 178/500 (35%), Positives = 273/500 (54%), Gaps = 49/500 (9%)
Query: 302 FNIGKAHVTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALL 361
F+ KA +VD +T K F DVAG +EA E+ E V +LKNP ++++G K P G LL
Sbjct: 412 FSQSKAEA-RVDGSTGVK--FADVAGIDEAVDELQELVKYLKNPDLFDKMGIKPPHGVLL 468
Query: 362 VGPPGTGKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFI 421
GPPG GKTL+AKA AGE+GVPF ++GS+F+E+ VGVG +R+R+LF+ A+ PS++FI
Sbjct: 469 EGPPGCGKTLVAKAIAGEAGVPFYQMAGSEFVEVLVGVGSARIRDLFKRAKVNKPSVIFI 528
Query: 422 DEIDAIGRKRGRGGFSGSND--------ERESTLNQLLVEMDGFGTTAGVVVLAGTNRAD 473
DEIDA+ +R +G F ++D ERE+TLNQLL+E+DGF T GV+ L TNR D
Sbjct: 529 DEIDALATRR-QGIFKENSDQLYNAATQERETTLNQLLIELDGFDTGKGVIFLGATNRRD 587
Query: 474 ILDNALLRPGRFDRTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGA 533
+LD ALLRPGRFDR I + P+ KGR I +I+ K+K+ S A+ PG++GA
Sbjct: 588 LLDPALLRPGRFDRKIRVRPPNAKGRLDILKIHASKVKMSDSVDLSS--YASNLPGWSGA 645
Query: 534 DIANVCNEAALIAARTDESQVTMDHFEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHA 593
+A + EAAL+A R + + + A+DR+ G + + + + A E G A
Sbjct: 646 KLAQLVQEAALVAVRKTHNSILQSDMDDAVDRLTVGPTRIGLELGHQGQCRRATTEVGVA 705
Query: 594 VAGWFL--------EHCEPLLKVTIVPRG--TAALGFAQYVPSENLLRTKEQLLDMTCMT 643
+ L E C+ +V+I+PRG + + F + + QLL +
Sbjct: 706 ITSHLLLRYENAKIERCD---RVSIIPRGQTLSQVVFHRLDDESYMFGRLPQLLHRLQVL 762
Query: 644 LGGRAAEQVLIGAISTGAQND-LEKVTKMTYAQVAIYGFSEKVGLLSFP---QNEDQFGK 699
LGGRAAE+V+ G+ ++ A D L + + + I+ + + P + QF
Sbjct: 763 LGGRAAEEVIYGSDTSKASVDYLSDASWLARKILTIWNLENPMVIHGEPPPWRKRPQFVG 822
Query: 700 PYSGDTGNI--------------IDQEV----RDWVNHAYERTVQLIEEHKEKLAQIAEL 741
P G++ +D EV + ++ Y +TV L+ +++ L + ++
Sbjct: 823 PRLDFEGSLYDDYDLVEPPVNFNMDDEVAHRSEELISQMYNKTVSLLRQNQTALLKTVKV 882
Query: 742 LLEKEVLHQEDLVRILGERP 761
LL ++ + E + IL P
Sbjct: 883 LLNQKEISGEAIDFILDHYP 902
>At3g02450 cell division protein FtsH-like protein
Length = 622
Score = 259 bits (663), Expect = 3e-69
Identities = 134/261 (51%), Positives = 182/261 (69%), Gaps = 5/261 (1%)
Query: 314 KNTKNK-VYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLL 372
+ +KN V F DV G + AK E++E V L+ Y++LGA++P+G LLVGPPGTGKTLL
Sbjct: 324 RRSKNPTVGFDDVEGVDSAKDELVEIVSCLQGSINYKKLGARLPRGVLLVGPPGTGKTLL 383
Query: 373 AKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRG 432
A+A AGE+GVPF S+S S+F+E+FVG G +R+R+LF AR+ +PSI+FIDE+DA+G KRG
Sbjct: 384 ARAVAGEAGVPFFSVSASEFVELFVGRGAARIRDLFNAARKNSPSIIFIDELDAVGGKRG 443
Query: 433 RGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISID 492
R NDER+ TLNQLL EMDGF + V+V+A TNR + LD+AL RPGRF R + +
Sbjct: 444 R----SFNDERDQTLNQLLTEMDGFESDTKVIVIAATNRPEALDSALCRPGRFSRKVLVA 499
Query: 493 VPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDES 552
PD +GR +I I+L+ + L+ + +A+LTPGF GAD+AN+ NEAAL+AAR
Sbjct: 500 EPDQEGRRKILAIHLRDVPLEEDAFLICDLVASLTPGFVGADLANIVNEAALLAARRGGE 559
Query: 553 QVTMDHFEAAIDRIIGGLEKK 573
V + AI+R G+ K
Sbjct: 560 AVAREDIMEAIERAKFGINDK 580
>At3g16290 putative FtsH-like metalloprotease
Length = 876
Score = 258 bits (658), Expect = 1e-68
Identities = 168/443 (37%), Positives = 251/443 (55%), Gaps = 18/443 (4%)
Query: 320 VYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGE 379
V F DVAG + + E+ E V F + + Y G KIP G LL GPPG GKTLLAKA AGE
Sbjct: 407 VKFTDVAGLGKIRLELEEIVKFFTHGEMYRRRGVKIPGGILLCGPPGVGKTLLAKAVAGE 466
Query: 380 SGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGS 439
+GV F SIS S F+E++VGVG SRVR L+QEAR+ APS+VFIDE+DA+GR+RG SG
Sbjct: 467 AGVNFFSISASQFVEIYVGVGASRVRALYQEARENAPSVVFIDELDAVGRERGLIKGSG- 525
Query: 440 NDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGR 499
ER++TLNQLLV +DGF V+ +A TNR DILD AL+RPGRFDR I I P + GR
Sbjct: 526 GQERDATLNQLLVSLDGFEGRGEVITIASTNRPDILDPALVRPGRFDRKIFIPKPGLIGR 585
Query: 500 DQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDH- 558
+I Q++ +K + + Y + +A++T G GA++AN+ AA+ R +++T D
Sbjct: 586 MEILQVHARKKPMAEDLDYMA--VASMTDGMVGAELANIVEIAAINMMRDGRTELTTDDL 643
Query: 559 FEAAIDRIIGGLEKKNRVISKRERRTVAYHEAGHAVAGWFLEHCEPLLKVTIVPRGTAAL 618
+AA G L++K+R S R VA +EA AV + + +TI PR L
Sbjct: 644 LQAAQIEERGMLDRKDR--SLETWRQVAINEAAMAVVAVNFPDMKNIEFLTINPRAGREL 701
Query: 619 GFAQ----YVPSENLLRTKEQLLDMTCMTLGGRAAEQVLIGAISTGAQNDLEKVTKMTYA 674
G+ + ++ + + +++ +LD + L RAA+++ G ++ L + T
Sbjct: 702 GYVRVKMDHIKFKEGMLSRQSILDHITVQLAPRAADELWYG------EDQLSTIWAETSD 755
Query: 675 QVAIYGFSEKVGLLSFPQNEDQFGKPYSGDTGNIIDQEVRDWVNHAYERTVQLIEEHKEK 734
S +G LS + D N ID E +N YER +++ ++
Sbjct: 756 NARSAARSLVLGGLS--DKHHGLNNFWVADRINDIDVEALRILNMCYERAKEILGRNRTL 813
Query: 735 LAQIAELLLEKEVLHQEDLVRIL 757
+ ++ E L++K+ L +++ ++
Sbjct: 814 MDEVVEKLVQKKSLTKQEFFTLV 836
>At5g64580 unknown protein
Length = 855
Score = 248 bits (634), Expect = 7e-66
Identities = 204/694 (29%), Positives = 324/694 (46%), Gaps = 68/694 (9%)
Query: 109 EKKSESKDESKSNTED----GGSFHEAFIKQFQNYLTPLLVVGLFLSSLSLGPRDQQQIS 164
EK E++ E SN E+ E + ++ LL + L P +I+
Sbjct: 90 EKLRETERERLSNMEELERKANVQLERQLVMASDWSRTLLTMRGKLKGTEWDPETSHRIN 149
Query: 165 FQEFKNKLLEPGLVDHIVVSNKS-----VAKIYVRNSPLNQADSE---------VQGTLP 210
F +F KLL+ V ++ SN + Y PL + + + +P
Sbjct: 150 FSDFM-KLLDSNSVQYMEYSNYGQTISVILPYYKDGEPLGEEEDSKKEIIFRRHIVDRMP 208
Query: 211 AKGSGGQYKYIIN-IGSVESFEEKLEEAQEALGVDSHNFVPVTYSSEMVWYQELMRFAPT 269
G +K + I +VE F + A+ V T ++ +VW L F
Sbjct: 209 IDGWNDVWKKLHQQIVNVEVFNVDVVPAE----------VYTTVATFVVWSMRLALFVSL 258
Query: 270 LLLLGTLW--FMGRKMQGGFGVGGGSTGKGSR--GIFNIGKAHVTKVDKNTKNKVYFKDV 325
+ + ++ + + G + + + ++GK+ + K V F D
Sbjct: 259 YVWIDSITRPIYAKLIPCDLGTPTKKIRQPLKRQALGSLGKSRAKFISAEEKTGVTFDDF 318
Query: 326 AGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKATAGESGVPFL 385
AG E K+E+ E V LKN ++++ G PKG LL GPPGTGKTLLAKA AGE+G+PF
Sbjct: 319 AGQEYIKRELQEIVRILKNDEEFQNKGIYCPKGVLLHGPPGTGKTLLAKAIAGEAGLPFF 378
Query: 386 SISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGGFSGSNDERES 445
+ +G+DF+EMFVGV SRV++LF +R APSI+FIDEIDAIG KRG G ERE
Sbjct: 379 AANGTDFVEMFVGVAASRVKDLFASSRSYAPSIIFIDEIDAIGSKRGGPDIGGGGAEREQ 438
Query: 446 TLNQLLVEMDGFG-TTAGVVVLAGTNRADILDNALLRPGRFDRTISIDVPDIKGRDQIFQ 504
L Q+L EMDGF TT+ V+V+ TNR DILD ALLR GRFD+ I + +P GR I +
Sbjct: 439 GLLQILTEMDGFKVTTSQVLVIGATNRLDILDPALLRKGRFDKIIRVGLPSKDGRLAILK 498
Query: 505 IYL--KKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAARTDESQVTMDHFEAA 562
++ K + + E Q +A T F GA++ NV NEA ++ AR D + + A
Sbjct: 499 VHARNKFFRSEDEKEELLQEVAENTEDFTGAELQNVLNEAGILTARKDLDYIGREELLEA 558
Query: 563 IDRIIGGLE---KKNRVISKRERRTVAYHEAGHAVAGWFL-EHCEPLLKVTIVP-RGTAA 617
+ R G E + + + + + +AY EA AV +L + P+ + I R
Sbjct: 559 LKRQKGTFETGQEDSTEVPEELKLRLAYREAAVAVLACYLPDQYRPISETDINSIRSQPN 618
Query: 618 LGFAQYVPSENLLRTKEQLLDMTCMTLGGRAAEQVLIGAIS---TGAQNDLEKVTKMTYA 674
+ +++ S + K ++ R E+ + G + A++ LE
Sbjct: 619 MRYSE--TSGRVFARKSDYVNSIIRACAPRVVEEEMFGIENLCWISAKSTLEA------- 669
Query: 675 QVAIYGFSEKVGLLSFPQNEDQFGKPYSGDTGNIID------QEVRD-WVNHAYERTVQL 727
S++ L FGK Y + +++ + +RD ++ A E+ +
Sbjct: 670 -------SQRAEFLILQTGMTAFGKAYYRNQRDLVPNLVPKLEALRDEYMRFAVEKCSSI 722
Query: 728 IEEHKEKLAQIAELLLEKEVLHQEDLVRILGERP 761
++E++ L +I ++LLEK + +++ I P
Sbjct: 723 LQEYQSALEEITDVLLEKGEIKADEIWNIYNTAP 756
>At4g29040 26S proteasome AAA-ATPase subunit RPT2a
Length = 443
Score = 195 bits (496), Expect = 7e-50
Identities = 113/260 (43%), Positives = 155/260 (59%), Gaps = 6/260 (2%)
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
V KV+K + D+ G E QEI E V L +P+ YE++G K PKG +L G PGT
Sbjct: 176 VMKVEKAPLES--YADIGGLEAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGT 233
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKTLLAKA A + FL + GS+ ++ ++G GP VR LF+ A +PSIVFIDEIDA+
Sbjct: 234 GKTLLAKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAV 293
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
G KR SG E + T+ +LL ++DGF + V V+ TNR + LD ALLRPGR DR
Sbjct: 294 GTKR-YDAHSGGEREIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDR 352
Query: 488 TISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAA 547
I +PDIK R +IFQI+ K+ L + + + F+GADI +C EA L+A
Sbjct: 353 KIEFPLPDIKTRRRIFQIHTSKMTLSEDVNL--EEFVMTKDEFSGADIKAICTEAGLLAL 410
Query: 548 RTDESQVTMDHFEAAIDRII 567
R +VT F+ A ++++
Sbjct: 411 RERRMKVTHPDFKKAKEKVM 430
>At1g53750 26S proteasome ATPase subunit
Length = 426
Score = 195 bits (496), Expect = 7e-50
Identities = 108/265 (40%), Positives = 155/265 (57%), Gaps = 4/265 (1%)
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
VT + K V + DV GC+E +++ E V + +P+K+ +LG PKG L GPPGT
Sbjct: 154 VTMMTVEEKPDVTYNDVGGCKEQIEKMREVVELPMLHPEKFVKLGIDPPKGVLCYGPPGT 213
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKTLLA+A A + F+ + GS+ ++ +VG G VR LFQ AR IVF DE+DAI
Sbjct: 214 GKTLLARAVANRTDACFIRVIGSELVQKYVGEGARMVRELFQMARSKKACIVFFDEVDAI 273
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
G R G G N E + T+ +++ ++DGF + VL TNR D LD ALLRPGR DR
Sbjct: 274 GGARFDDGVGGDN-EVQRTMLEIVNQLDGFDARGNIKVLMATNRPDTLDPALLRPGRLDR 332
Query: 488 TISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAA 547
+ +PD++ R QIF+I+ + + + + + + LA L P GADI +VC EA + A
Sbjct: 333 KVEFGLPDLESRTQIFKIHTRTMNCERDIRF--ELLARLCPNSTGADIRSVCTEAGMYAI 390
Query: 548 RTDESQVTMDHFEAAIDRIIGGLEK 572
R VT F A++++I G +K
Sbjct: 391 RARRKTVTEKDFLDAVNKVIKGYQK 415
>At1g53780 hypothetical protein
Length = 464
Score = 195 bits (495), Expect = 1e-49
Identities = 106/265 (40%), Positives = 157/265 (59%), Gaps = 4/265 (1%)
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
VT + K + D+ GC+E ++I E V + +P+K+ LG PKG L GPPG+
Sbjct: 191 VTMMTVEEKPDATYSDIGGCKEQIEKIREVVELPMLHPEKFVRLGIDPPKGVLCYGPPGS 250
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKTL+A+A A +G F+ + GS+ ++ ++G G VR LFQ AR I+F DEIDAI
Sbjct: 251 GKTLVARAVANRTGACFIRVVGSELVQKYIGEGARMVRELFQMARSKKACILFFDEIDAI 310
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
G R G GS++E + T+ ++L ++DGF + VL TNR DILD ALLRPGR DR
Sbjct: 311 GGARFDDGV-GSDNEVQRTMLEILYQLDGFDARGNIKVLMATNRPDILDPALLRPGRLDR 369
Query: 488 TISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAA 547
+ +PD++GR QIF+I+ + + + + + + LA L P GADI +VC EA + A
Sbjct: 370 KVEFCLPDLEGRTQIFKIHTRTMSCERDIRF--ELLAGLCPNSTGADIRSVCIEAGMYAI 427
Query: 548 RTDESQVTMDHFEAAIDRIIGGLEK 572
VT F A+++++ G +K
Sbjct: 428 GARRKSVTEKDFLDAVNKVVKGYQK 452
>At2g20140 26S proteasome subunit 4
Length = 443
Score = 193 bits (491), Expect = 3e-49
Identities = 113/260 (43%), Positives = 155/260 (59%), Gaps = 6/260 (2%)
Query: 309 VTKVDKNTKNKVYFKDVAGCEEAKQEIMEFVHF-LKNPKKYEELGAKIPKGALLVGPPGT 367
V KV+K + D+ G E QEI E V L +P+ YE++G K PKG +L G PGT
Sbjct: 176 VMKVEKAPLES--YADIGGLEAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGT 233
Query: 368 GKTLLAKATAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAI 427
GKTLLAKA A + FL + GS+ ++ ++G GP VR LF+ A +PSIVFIDEIDA+
Sbjct: 234 GKTLLAKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAV 293
Query: 428 GRKRGRGGFSGSNDERESTLNQLLVEMDGFGTTAGVVVLAGTNRADILDNALLRPGRFDR 487
G KR SG E + T+ +LL ++DGF + V V+ TNR + LD ALLRPGR DR
Sbjct: 294 GTKRYDAN-SGGEREIQRTMLELLNQLDGFDSRGDVKVILATNRIESLDPALLRPGRIDR 352
Query: 488 TISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIAA 547
I +PDIK R +IFQI+ K+ L + + + F+GADI +C EA L+A
Sbjct: 353 KIEFPLPDIKTRRRIFQIHTSKMTLAEDVNL--EEFVMTKDEFSGADIKAICTEAGLLAL 410
Query: 548 RTDESQVTMDHFEAAIDRII 567
R +VT F+ A ++++
Sbjct: 411 RERRMKVTHVDFKKAKEKVM 430
>At1g79560 FtsH like cell division protein
Length = 619
Score = 192 bits (489), Expect = 5e-49
Identities = 146/509 (28%), Positives = 244/509 (47%), Gaps = 52/509 (10%)
Query: 316 TKNKVYFKDVAGCEEAKQEIMEFVHFLKNPKKYEELGAKIPKGALLVGPPGTGKTLLAKA 375
++ K +K+V + + E + ++ NP +Y E +G LL GPPGTGKTL A+
Sbjct: 97 SETKSMYKEVVLGGDVWDLLDELMIYMGNPMQYYEKDVAFVRGVLLSGPPGTGKTLFART 156
Query: 376 TAGESGVPFLSISGSDFMEMFVGVGPSRVRNLFQEARQCAPSIVFIDEIDAIGRKRGRGG 435
A ESG+PF+ SG++F + G +++ +F AR+ AP+ VF+DEIDAI + R
Sbjct: 157 LAKESGLPFVFASGAEFTDSEKS-GAAKINEMFSIARRNAPAFVFVDEIDAIAGRHAR-- 213
Query: 436 FSGSNDERESTLNQLLVEMDG---------FGTTAGVVVLAGTNRADILDNALLRPGRFD 486
+ R +T L+ ++DG F V+ + TNR D LD +R GR D
Sbjct: 214 ---KDPRRRATFEALIAQLDGEKEKTGIDRFSLRQAVIFICATNRPDELDLEFVRSGRID 270
Query: 487 RTISIDVPDIKGRDQIFQIYLKKIKLDHEPSYYSQRLAALTPGFAGADIANVCNEAALIA 546
R + I +PD K R QIF ++ L + + +L T GF+GADI N+ NEAA+++
Sbjct: 271 RRLYIGLPDAKQRVQIFGVHSAGKNLAEDIDF--GKLVFRTVGFSGADIRNLVNEAAIMS 328
Query: 547 ARTDESQVTMDHFEAAIDR-IIGGL---------EKKNRVISKRERRTVAYHEAGHAVAG 596
R S + +D+ ++ G+ +K + +S ++R +A HEAGH V
Sbjct: 329 VRKGRSYIYQQDIVDVLDKQLLEGMGVLLTEEEQQKCEQSVSYEKKRLLAVHEAGHIVLA 388
Query: 597 WFLEHCEPLLKVTIVPRGTAALGFAQYVPSENLL----RTKEQLLDMTCMTLGGRAAEQV 652
+ ++P G + + P E+++ T + + GGR AE+V
Sbjct: 389 HLFPRFDWHAFSQLLPGGKET-AVSVFYPREDMVDQGYTTFGYMKMQMVVAHGGRCAERV 447
Query: 653 LIG-AISTGAQNDLEKVTKMT--------YAQVAIYGFSEKVGLLSFPQNEDQFGKPYSG 703
+ G ++ G ++DLEK+TK+ A++ + +K+G++ P N D Y
Sbjct: 448 VFGDNVTDGGKDDLEKITKIAREMVISPQSARLGLTQLVKKIGMVDLPDNPDGELIKYRW 507
Query: 704 DTGNIIDQEVRDWVNHAYERTVQLIEEHKEKLAQIAEL-------LLEKEVLHQEDLVRI 756
D +++ E+ V+ + R + E E+LA A L+ +E+L + + +
Sbjct: 508 DHPHVMPAEMSVEVSELFTRELTRYIEETEELAMNALRANRHILDLITRELLEKSRITGL 567
Query: 757 LGERPFKSAEPTNYDRFKLGFQ----DEE 781
E K P ++ F FQ DEE
Sbjct: 568 EVEEKMKDLSPLMFEDFVKPFQINPDDEE 596
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.316 0.135 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,862,068
Number of Sequences: 26719
Number of extensions: 809444
Number of successful extensions: 4171
Number of sequences better than 10.0: 154
Number of HSP's better than 10.0 without gapping: 111
Number of HSP's successfully gapped in prelim test: 43
Number of HSP's that attempted gapping in prelim test: 3687
Number of HSP's gapped (non-prelim): 287
length of query: 807
length of database: 11,318,596
effective HSP length: 107
effective length of query: 700
effective length of database: 8,459,663
effective search space: 5921764100
effective search space used: 5921764100
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135467.11


