

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.23 + phase: 0
(150 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
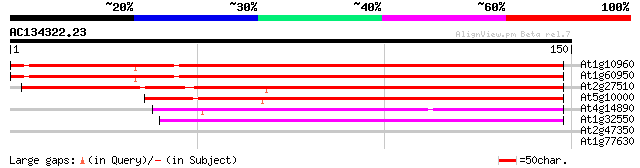
Score E
Sequences producing significant alignments: (bits) Value
At1g10960 ferredoxin precusor isolog 191 1e-49
At1g60950 ferrodoxin precursor 191 2e-49
At2g27510 putative ferredoxin 143 3e-35
At5g10000 putative protein 115 9e-27
At4g14890 ferredoxin 81 2e-16
At1g32550 ferredoxin, putative 76 6e-15
At2g47350 unknown protein 28 2.4
At1g77630 unknown protein with predicted GPI-anchor 27 5.4
>At1g10960 ferredoxin precusor isolog
Length = 148
Score = 191 bits (486), Expect = 1e-49
Identities = 100/149 (67%), Positives = 115/149 (77%), Gaps = 3/149 (2%)
Query: 1 MATTPALYGTAVSTSFLRRQPMPVSISTTTKA-FPSGFGLKSKTGKRGDLAVAMATYKVK 59
MA+T AL VSTSFLRRQ P+S+ + A S FGLKS T RG AMATYKVK
Sbjct: 1 MAST-ALSSAIVSTSFLRRQQTPISLRSLPFANTQSLFGLKSSTA-RGGRVTAMATYKVK 58
Query: 60 LVTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDD 119
+TPEG QE +C DVY+LD AEE G+DLPYSCRAGSCSSCAGKV +G++DQSD SFLDD
Sbjct: 59 FITPEGEQEVECEEDVYVLDAAEEAGLDLPYSCRAGSCSSCAGKVVSGSIDQSDQSFLDD 118
Query: 120 DQIEEGWVLTCVAYPTSDVTIETHKEEEL 148
+Q+ EG+VLTCVAYPTSDV IETHKEE +
Sbjct: 119 EQMSEGYVLTCVAYPTSDVVIETHKEEAI 147
>At1g60950 ferrodoxin precursor
Length = 148
Score = 191 bits (484), Expect = 2e-49
Identities = 100/149 (67%), Positives = 114/149 (76%), Gaps = 3/149 (2%)
Query: 1 MATTPALYGTAVSTSFLRRQPMPVSISTTTKA-FPSGFGLKSKTGKRGDLAVAMATYKVK 59
MA+T AL V TSF+RR P P+S+ + A S FGLKS T RG AMATYKVK
Sbjct: 1 MAST-ALSSAIVGTSFIRRSPAPISLRSLPSANTQSLFGLKSGTA-RGGRVTAMATYKVK 58
Query: 60 LVTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDD 119
+TPEG E +C DVY+LD AEE GIDLPYSCRAGSCSSCAGKV +G+VDQSD SFLDD
Sbjct: 59 FITPEGELEVECDDDVYVLDAAEEAGIDLPYSCRAGSCSSCAGKVVSGSVDQSDQSFLDD 118
Query: 120 DQIEEGWVLTCVAYPTSDVTIETHKEEEL 148
+QI EG+VLTC AYPTSDVTIETHKEE++
Sbjct: 119 EQIGEGFVLTCAAYPTSDVTIETHKEEDI 147
>At2g27510 putative ferredoxin
Length = 155
Score = 143 bits (361), Expect = 3e-35
Identities = 74/146 (50%), Positives = 97/146 (65%), Gaps = 4/146 (2%)
Query: 4 TPALYGTAVSTSFLRRQPMPVSISTTTKAFPSGFGLKSKTGKRGDLAVAMATYKVKLVTP 63
T A+ + + + + +S+ +T + S FGLK G A A YKVKL+ P
Sbjct: 12 TKAVLRSQTTNKLITNKSYNLSVGSTKRVSRS-FGLKCSANSGG--ATMSAVYKVKLLGP 68
Query: 64 EGTQ-EFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDGSFLDDDQI 122
+G + EF+ D YILD AEE G+DLPYSCRAG+CS+CAG++ +G VDQSDGSFL+D +
Sbjct: 69 DGQEDEFEVQDDQYILDAAEEAGVDLPYSCRAGACSTCAGQIVSGNVDQSDGSFLEDSHL 128
Query: 123 EEGWVLTCVAYPTSDVTIETHKEEEL 148
E+G+VLTCVAYP SD I THKE EL
Sbjct: 129 EKGYVLTCVAYPQSDCVIHTHKETEL 154
>At5g10000 putative protein
Length = 148
Score = 115 bits (288), Expect = 9e-27
Identities = 55/113 (48%), Positives = 79/113 (69%), Gaps = 2/113 (1%)
Query: 37 FGLKSKTGKRGDLAVAMATYKVKLVTPEGT-QEFDCPSDVYILDHAEEVGIDLPYSCRAG 95
FGL S G G + A + KVKL++PEG QE + D IL+ AE G++LPYSCR+G
Sbjct: 36 FGLSSSRGNFGKV-FAKESRKVKLISPEGEEQEIEGNEDCCILESAENAGLELPYSCRSG 94
Query: 96 SCSSCAGKVAAGAVDQSDGSFLDDDQIEEGWVLTCVAYPTSDVTIETHKEEEL 148
+C +C GK+ +G VDQS GSFL+++QI++G++LTC+A P D + THK+ +L
Sbjct: 95 TCGTCCGKLVSGKVDQSLGSFLEEEQIQKGYILTCIALPLEDCVVYTHKQSDL 147
>At4g14890 ferredoxin
Length = 154
Score = 80.9 bits (198), Expect = 2e-16
Identities = 45/113 (39%), Positives = 59/113 (51%), Gaps = 4/113 (3%)
Query: 39 LKSKTGKRGDLA---VAMATYKVKLVTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAG 95
L + T +R L + YKV + T E + D IL A + G+D+PY C G
Sbjct: 33 LSTTTNRRNFLTTGRIIARAYKVVVEHDGKTTELEVEPDETILSKALDSGLDVPYDCNLG 92
Query: 96 SCSSCAGKVAAGAVDQSDGSFLDDDQIEEGWVLTCVAYPTSDVTIETHKEEEL 148
C +C K+ G VDQS G L DD +E G+ L C +YPTSD I+ EEEL
Sbjct: 93 VCMTCPAKLVTGTVDQS-GGMLSDDVVERGYTLLCASYPTSDCHIKMIPEEEL 144
>At1g32550 ferredoxin, putative
Length = 181
Score = 76.3 bits (186), Expect = 6e-15
Identities = 40/108 (37%), Positives = 56/108 (51%)
Query: 41 SKTGKRGDLAVAMATYKVKLVTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSC 100
S RG L V V EF+ P D YIL AE I LP++CR G C+SC
Sbjct: 46 STPSDRGSLVVPSHKVTVHDRQRGVVHEFEVPEDQYILHSAESQNISLPFACRHGCCTSC 105
Query: 101 AGKVAAGAVDQSDGSFLDDDQIEEGWVLTCVAYPTSDVTIETHKEEEL 148
A +V +G + Q + + +G+ L CV +PTSD+ +ET E+E+
Sbjct: 106 AVRVKSGELRQPQALGISAELKSQGYALLCVGFPTSDLEVETQDEDEV 153
>At2g47350 unknown protein
Length = 486
Score = 27.7 bits (60), Expect = 2.4
Identities = 14/58 (24%), Positives = 25/58 (42%)
Query: 57 KVKLVTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSSCAGKVAAGAVDQSDG 114
K K V+ + +++ DC ++ +E+G D S + A+D SDG
Sbjct: 272 KKKAVSEQASEDMDCAEEIETASDEKEIGNDNKRESTMTSRQRALASGRSSAIDFSDG 329
>At1g77630 unknown protein with predicted GPI-anchor
Length = 423
Score = 26.6 bits (57), Expect = 5.4
Identities = 14/40 (35%), Positives = 23/40 (57%)
Query: 60 LVTPEGTQEFDCPSDVYILDHAEEVGIDLPYSCRAGSCSS 99
LV+PE Q + +D+ +LD ++ I LP +C G+ S
Sbjct: 128 LVSPEQIQVANSETDLSVLDVGTKLVIPLPCACFNGTDES 167
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.314 0.130 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,450,370
Number of Sequences: 26719
Number of extensions: 137002
Number of successful extensions: 232
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 1
Number of HSP's that attempted gapping in prelim test: 220
Number of HSP's gapped (non-prelim): 8
length of query: 150
length of database: 11,318,596
effective HSP length: 90
effective length of query: 60
effective length of database: 8,913,886
effective search space: 534833160
effective search space used: 534833160
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Medicago: description of AC134322.23


