

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
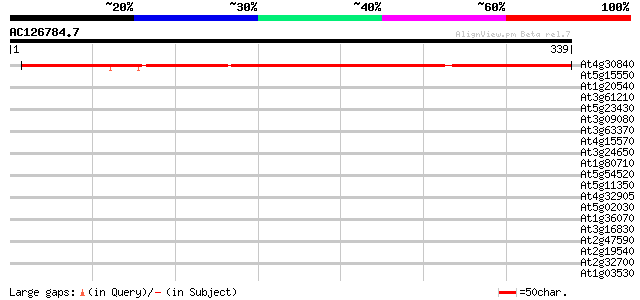
Query= AC126784.7 - phase: 0
(339 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At4g30840 unknown protein 376 e-105
At5g15550 unknown protein 35 0.045
At1g20540 unknown protein 35 0.059
At3g61210 unknown protein 34 0.100
At5g23430 unknown protein 32 0.65
At3g09080 hypothetical protein 32 0.65
At3g63370 putative protein 31 1.1
At4g15570 SEN1 like protein 30 1.4
At3g24650 abscisic acid-insensitive protein 3 30 2.5
At1g80710 unknown protein 30 2.5
At5g54520 unknown protein 29 3.2
At5g11350 unknown protein 29 3.2
At4g32905 unknown protein 29 4.2
At5g02030 putative homeodomain protein 28 5.5
At1g36070 unknown protein 28 5.5
At3g16830 unknown protein 28 7.2
At2g47590 photolyase/blue-light receptor (PHR2) 28 7.2
At2g19540 putative WD-40 repeat protein 28 7.2
At2g32700 unknown protein 28 9.3
At1g03530 unknown protein 28 9.3
>At4g30840 unknown protein
Length = 361
Score = 376 bits (966), Expect = e-105
Identities = 201/348 (57%), Positives = 233/348 (66%), Gaps = 23/348 (6%)
Query: 8 EIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLS-------F 60
++HR PQ KY+D VRWLP SA NRF A+ D D + SSIEI S PNP
Sbjct: 9 QVHRIPQSKYVDGVRWLPQASALNRFFATASYDADCDSSSIEIQSLDPNPRGNHNTNPLI 68
Query: 61 EFQSSWTSPSPISSIK-------SSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSV 113
E SSWTSPS +SS++ F K +++++TSSGSLH L D + + E
Sbjct: 69 ESLSSWTSPSRVSSLEVAGNGGGGGSF--KPMVSAATSSGSLHVLMIDLVEGAAIEEFYA 126
Query: 114 PENELHLEGKC-CIDLMDGGVECVTVGDDGRINLVT-VGDSNLNYRRLFDSGGLVSYTSV 171
E E G+ +D +GG ECVTVG+DGR+N+V V L YR++FD GLV+Y +V
Sbjct: 127 AEGERFHVGRVEGVDWREGG-ECVTVGEDGRVNVVKIVNGEGLRYRKVFDGNGLVAYRAV 185
Query: 172 KWASPVEFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLA 231
KWASP EF TGGYGFGL WDQRK G VSQ KGNW + S IVHSIDIHPSRKHTC+A
Sbjct: 186 KWASPTEFVTGGYGFGLQLWDQRKSGEAVSQLKGNWFQGKTSAIVHSIDIHPSRKHTCIA 245
Query: 232 AGSLGTVLAWDLRMQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSSTR 291
GS GTV AWDLR QQPI+LSG GA + N +SESEVWEVQYD KSN SS+R
Sbjct: 246 GGSSGTVFAWDLRWPQQPIVLSGVGASENINN----PLSESEVWEVQYDSYTKSNVSSSR 301
Query: 292 ILPAMICSEDGILGVIEQGEEPIELLAEPCAINSFDIDRHNPLVSGCS 339
ILP M CSEDGILG+IEQGEEPIELLAEPCAINSFDIDR NP CS
Sbjct: 302 ILPVMTCSEDGILGIIEQGEEPIELLAEPCAINSFDIDRQNPQDVICS 349
>At5g15550 unknown protein
Length = 433
Score = 35.4 bits (80), Expect = 0.045
Identities = 33/100 (33%), Positives = 42/100 (42%), Gaps = 12/100 (12%)
Query: 147 VTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGYGFGLHWWDQRKPG--GPVSQFK 204
V G +LN LF G ++ V S A GG L WD RKPG PV QF
Sbjct: 300 VETGKDSLN---LF-CGKALNTVDVGGESSALIAAGGSDPILRVWDPRKPGTSAPVFQFS 355
Query: 205 GNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLR 244
+S + + H S L+A G ++ WDLR
Sbjct: 356 S------HSSWISACKWHKSSWFHLLSASYDGKIMLWDLR 389
>At1g20540 unknown protein
Length = 351
Score = 35.0 bits (79), Expect = 0.059
Identities = 35/146 (23%), Positives = 56/146 (37%), Gaps = 17/146 (11%)
Query: 116 NELHLEGKCCIDLMDGGVECVTVGDDGRIN-LVTVGDSNLNYRRLFDSGGLVSYTSVKWA 174
N LE +D G + CV GR + L+++ + N+ L S S A
Sbjct: 106 NSPQLERVASLDAHVGKINCVLWWPSGRCDKLISIDEQNIFLWSLDCSKKSAEVLSKDSA 165
Query: 175 SPVEFATGGYGFGLHWWDQRKPGGPVSQFKGN---WD----KNLNS---GIVHSIDIHPS 224
+ +GG WD + + + WD K +NS V +D +P
Sbjct: 166 GMLHSLSGGA------WDPHDVNAVAATGESSVQFWDLRTMKKVNSIEHAHVRGVDYNPK 219
Query: 225 RKHTCLAAGSLGTVLAWDLRMQQQPI 250
R+H + A + WDLR + P+
Sbjct: 220 REHILVTAEDESGIHVWDLRKAKVPV 245
>At3g61210 unknown protein
Length = 261
Score = 34.3 bits (77), Expect = 0.100
Identities = 29/116 (25%), Positives = 52/116 (44%), Gaps = 14/116 (12%)
Query: 222 HPSRKHTCLAAGSLGTVLAWDLRMQQQPIILSGTGAGDGAGNTA--VQSISESEVWEVQY 279
+P+ + LA + +AWD+ GTG G A A Q + +++ E Q
Sbjct: 22 YPTIWYKVLAGRTSNHKVAWDV----------GTGNGQAAIGVAEYYQKVVATDINESQL 71
Query: 280 DRCIKSNTSSTRILPAMICSEDGILGVIEQGEEPIELLAEPCAINSFDIDRHNPLV 335
R +K + P+ + +D L + GE I+++ A++ FD+ R P+V
Sbjct: 72 QRAMKHPKVTYYHTPSSMSDDD--LVTLLGGENSIDIIIAAQALHYFDLKRFYPIV 125
>At5g23430 unknown protein
Length = 837
Score = 31.6 bits (70), Expect = 0.65
Identities = 34/110 (30%), Positives = 47/110 (41%), Gaps = 12/110 (10%)
Query: 136 VTVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASPVEFATGGYGFGLHWWDQRK 195
VT G+D ++NL +G N S G+ S T AS V A G + WD +
Sbjct: 33 VTGGEDHKVNLWAIGKPNAILSLYGHSSGIDSVTFD--ASEVLVAAGAASGTIKLWDLEE 90
Query: 196 PGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVL-AWDLR 244
V G+ + S+D HP + A+GSL T L WD+R
Sbjct: 91 -AKIVRTLTGHRSNCI------SVDFHPFGEF--FASGSLDTNLKIWDIR 131
>At3g09080 hypothetical protein
Length = 1026
Score = 31.6 bits (70), Expect = 0.65
Identities = 33/133 (24%), Positives = 52/133 (38%), Gaps = 28/133 (21%)
Query: 188 LHWWDQRKPGGP------VSQFKGNWD-KNLNSGIVHSIDIHPSRKHTCLAAGSL----- 235
L+ WD P +S G WD KNL+ G +HS P+ C+A G
Sbjct: 275 LYVWDVDDVNKPTRCSMMISHSAGIWDIKNLSCGNMHS----PTA--ACVARGCSEGVSF 328
Query: 236 ------GTVLAWDLRMQQQPIILSGTGAGDGAGNTAVQSISESEVWEVQYDRCIKSNTSS 289
GT+ WDL Q P+ + + + + ++ + ++E R + S
Sbjct: 329 TTCSEDGTIRLWDLAFQVNPLEANASSNPSESSTQGIMHLASAGIFE----RDLVETCGS 384
Query: 290 TRILPAMICSEDG 302
A+ SEDG
Sbjct: 385 KFGFRALAVSEDG 397
>At3g63370 putative protein
Length = 1017
Score = 30.8 bits (68), Expect = 1.1
Identities = 23/82 (28%), Positives = 38/82 (46%), Gaps = 12/82 (14%)
Query: 19 DAVRWLPVLSAFNRFAVLATSDFDSNLSSIEI----HSFKPNPLSFEFQSSWTSPSPISS 74
DAV W +LS++ S +L ++E+ H P P S+ S+ T+ S
Sbjct: 211 DAVLWNSILSSY--------STSGKSLETLELFREMHMTGPAPNSYTIVSALTACDGFSY 262
Query: 75 IKSSQFLQKSIIASSTSSGSLH 96
K + + S++ SST S L+
Sbjct: 263 AKLGKEIHASVLKSSTHSSELY 284
>At4g15570 SEN1 like protein
Length = 818
Score = 30.4 bits (67), Expect = 1.4
Identities = 20/53 (37%), Positives = 28/53 (52%), Gaps = 6/53 (11%)
Query: 185 GFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGT 237
GFG W K GP+++F +++NLN +ID+ SRK L G GT
Sbjct: 238 GFGDEAW---KISGPLNEF---FNENLNKSQKEAIDVGLSRKSFVLIQGPPGT 284
>At3g24650 abscisic acid-insensitive protein 3
Length = 720
Score = 29.6 bits (65), Expect = 2.5
Identities = 19/68 (27%), Positives = 30/68 (43%), Gaps = 7/68 (10%)
Query: 57 PLSFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPEN 116
P SS TSP+P+++I SS + +S++S+ S L +D D P
Sbjct: 80 PCMSSSSSSSTSPAPVNAIVSSASSSSAASSSTSSAASWAILRSDGED-------PTPNQ 132
Query: 117 ELHLEGKC 124
+ G C
Sbjct: 133 NQYASGNC 140
>At1g80710 unknown protein
Length = 516
Score = 29.6 bits (65), Expect = 2.5
Identities = 13/39 (33%), Positives = 21/39 (53%), Gaps = 2/39 (5%)
Query: 206 NWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLAWDLR 244
+W+ L+ ++SID +P H + + GT WDLR
Sbjct: 343 HWE--LHERRINSIDFNPQNPHVMATSSTDGTACLWDLR 379
>At5g54520 unknown protein
Length = 457
Score = 29.3 bits (64), Expect = 3.2
Identities = 42/184 (22%), Positives = 66/184 (35%), Gaps = 12/184 (6%)
Query: 63 QSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADSTDARLVSEVSVPENELHLEG 122
QSS S P SS + S + S H S + S +S+ H +
Sbjct: 104 QSSDLSQKPYSSPTVLGSISDSDVPRHVLSSVRHRPKGSSLQTEMPSRMSISLTG-HTKA 162
Query: 123 KCCIDLMDGGVECV-TVGDDGRINLVTVGDSNLNYRRLFDSGGLVSYTSVKWASP-VEFA 180
ID V + + G DG + + V ++ R F VKW+ +
Sbjct: 163 VTAIDWSTSHVHLLASAGLDGAVYVWNVWSNDKKKVRAFLHHN-APVKDVKWSKQGLSLL 221
Query: 181 TGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVLA 240
+ GY +D + G FK +V + HP + L+ GS G++
Sbjct: 222 SCGYDCTSRLFDVER-GVETQSFK-------EDEVVGVVKFHPDNCNVFLSGGSKGSLRL 273
Query: 241 WDLR 244
WD+R
Sbjct: 274 WDIR 277
>At5g11350 unknown protein
Length = 754
Score = 29.3 bits (64), Expect = 3.2
Identities = 32/132 (24%), Positives = 55/132 (41%), Gaps = 14/132 (10%)
Query: 44 NLSSIEIHSFKPNPLSFEFQSSWTSPSPISSIKSSQFLQKSIIASSTSSGSLHFLFADST 103
+LSS+ I +P S + T S S + ++ + I +S S
Sbjct: 510 DLSSLTISDTEPQHASSAREDLNTDRSVSSGLSETEQTPEEICSSDQDISS--------- 560
Query: 104 DARLVSEVSVPENELHLEGKCCIDLMDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSG 163
L ++V E+ L+G + + + ++G+DG L + D+N N L G
Sbjct: 561 --SLSTKVDTFVAEMKLDGLKLDEPVVFAQDEESLGEDGETFLAKLHDNNEN---LSQKG 615
Query: 164 GLVSYTSVKWAS 175
LVS +KW+S
Sbjct: 616 ELVSEVPLKWSS 627
>At4g32905 unknown protein
Length = 716
Score = 28.9 bits (63), Expect = 4.2
Identities = 20/53 (37%), Positives = 29/53 (53%), Gaps = 4/53 (7%)
Query: 268 SISESEVWE--VQYDRCIKSNTSSTRILPAMICSEDGILGVIEQGEEPIELLA 318
S+ ++ WE ++Y R IKS S+T LPA + + VI+ EEP L A
Sbjct: 120 SLYQTAWWEYVMEYSRDIKSLLSNTISLPAPLIEDP--RSVIKNAEEPTSLKA 170
>At5g02030 putative homeodomain protein
Length = 575
Score = 28.5 bits (62), Expect = 5.5
Identities = 15/40 (37%), Positives = 20/40 (49%)
Query: 25 PVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQS 64
P S+ AV F++ LSS ++ FKP PLS S
Sbjct: 82 PSSSSSTAVAVAGDHSFNAGLSSGDVLVFKPEPLSLSLSS 121
>At1g36070 unknown protein
Length = 418
Score = 28.5 bits (62), Expect = 5.5
Identities = 36/142 (25%), Positives = 54/142 (37%), Gaps = 20/142 (14%)
Query: 129 MDGGVECVTVGDDGRINLVTVGDSNLNYRRLFDSGGLVS---------YTSVKWASPVEF 179
+D V +V DG++ L VGDS SG ++S + S + +
Sbjct: 247 LDWSVNNTSVSPDGKL-LAVVGDSTECLISDSHSGKVISSLRGHKDYSFASAWHPNGLIL 305
Query: 180 ATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGTVL 239
ATG WD R P + KG N G + + P + +A + V
Sbjct: 306 ATGNQDTACRLWDIRNPSESFAVLKG------NMGAIRGLKFTPEGRFLAMAEPA-DFVH 358
Query: 240 AWDLR---MQQQPIILSGTGAG 258
+D + +Q Q I L G AG
Sbjct: 359 IFDTQSGFLQSQEIDLFGEIAG 380
>At3g16830 unknown protein
Length = 1131
Score = 28.1 bits (61), Expect = 7.2
Identities = 15/37 (40%), Positives = 23/37 (61%), Gaps = 2/37 (5%)
Query: 215 IVHSIDIHPSRKHTCLAAG-SLGTVLAWDLRMQQQPI 250
+V S+D HPS HT LA G S G V W++ +++ +
Sbjct: 349 VVISMDFHPSH-HTLLAVGCSSGEVTLWEVGSREKVV 384
>At2g47590 photolyase/blue-light receptor (PHR2)
Length = 447
Score = 28.1 bits (61), Expect = 7.2
Identities = 33/143 (23%), Positives = 62/143 (43%), Gaps = 10/143 (6%)
Query: 4 TANSEIHRFPQYKYIDAVRWLPVLSAFNRFAVLATSDFDSNLSSIEIHSFKPNPLSFEFQ 63
++N E + P+ K + L + + A L+ + F S+ +++ + PN + Q
Sbjct: 3 SSNVEENLNPETKSAEEQNPLAIFHSSLPIASLSLTLFPSSTQFLKLFAHHPNKVKIPTQ 62
Query: 64 SS---WTSPSPISSIKSSQFLQKSIIASSTSSGSLHFL---FADSTDARLVSEVSVP--E 115
+S S S +S SS+ KS IA++ L + D + A + +V
Sbjct: 63 ASSLTHLSLSSVSPFPSSRISFKSTIAANPLQSPLSIVPRRPVDPSSAAALRRAAVVWFR 122
Query: 116 NELHLEGKCCIDLMDGGVECVTV 138
N+L + C++ ECV+V
Sbjct: 123 NDLRVHDNECLN--SANDECVSV 143
>At2g19540 putative WD-40 repeat protein
Length = 469
Score = 28.1 bits (61), Expect = 7.2
Identities = 14/47 (29%), Positives = 19/47 (39%)
Query: 214 GIVHSIDIHPSRKHTCLAAGSLGTVLAWDLRMQQQPIILSGTGAGDG 260
G V+ I P H C++ G V WD+ + S T DG
Sbjct: 160 GCVNRIRAMPQNSHICVSWADSGHVQVWDMSSHLNALAESETEGKDG 206
>At2g32700 unknown protein
Length = 787
Score = 27.7 bits (60), Expect = 9.3
Identities = 16/66 (24%), Positives = 26/66 (39%), Gaps = 6/66 (9%)
Query: 178 EFATGGYGFGLHWWDQRKPGGPVSQFKGNWDKNLNSGIVHSIDIHPSRKHTCLAAGSLGT 237
+ AT + + WD PG + G+ + V SID HP + + S
Sbjct: 566 QLATSSFDKTIKIWDASDPGYFLRTISGH------AAPVMSIDFHPKKTELLCSCDSNND 619
Query: 238 VLAWDL 243
+ WD+
Sbjct: 620 IRFWDI 625
>At1g03530 unknown protein
Length = 801
Score = 27.7 bits (60), Expect = 9.3
Identities = 11/25 (44%), Positives = 13/25 (52%)
Query: 194 RKPGGPVSQFKGNWDKNLNSGIVHS 218
R PG P S + G W KN S + S
Sbjct: 530 RDPGRPTSSYSGEWTKNQGSSSLSS 554
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,218,990
Number of Sequences: 26719
Number of extensions: 376993
Number of successful extensions: 1003
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 983
Number of HSP's gapped (non-prelim): 27
length of query: 339
length of database: 11,318,596
effective HSP length: 100
effective length of query: 239
effective length of database: 8,646,696
effective search space: 2066560344
effective search space used: 2066560344
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC126784.7


