

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC126008.1 - phase: 0 /pseudo
(790 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
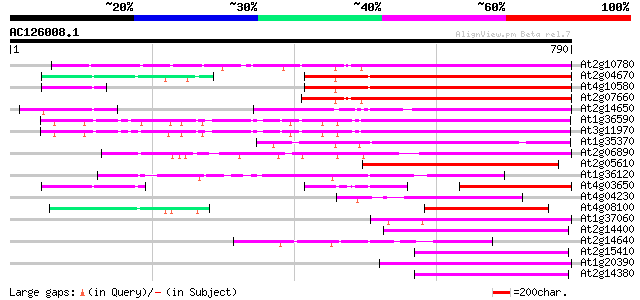
Score E
Sequences producing significant alignments: (bits) Value
At2g10780 pseudogene 474 e-133
At2g04670 putative retroelement pol polyprotein 402 e-112
At4g10580 putative reverse-transcriptase -like protein 399 e-111
At2g07660 putative retroelement pol polyprotein 397 e-110
At2g14650 putative retroelement pol polyprotein 369 e-102
At1g36590 hypothetical protein 334 1e-91
At3g11970 hypothetical protein 333 2e-91
At1g35370 hypothetical protein 291 1e-78
At2g06890 putative retroelement integrase 283 2e-76
At2g05610 putative retroelement pol polyprotein 281 8e-76
At1g36120 putative reverse transcriptase gb|AAD22339.1 263 3e-70
At4g03650 putative reverse transcriptase 207 2e-53
At4g04230 putative transposon protein 189 4e-48
At4g08100 putative polyprotein 184 2e-46
At1g37060 Athila retroelment ORF 1, putative 166 5e-41
At2g14400 putative retroelement pol polyprotein 159 4e-39
At2g14640 putative retroelement pol polyprotein 159 6e-39
At2g15410 putative retroelement pol polyprotein 157 3e-38
At1g20390 hypothetical protein 155 6e-38
At2g14380 putative retroelement pol polyprotein 148 1e-35
>At2g10780 pseudogene
Length = 1611
Score = 474 bits (1219), Expect = e-133
Identities = 293/762 (38%), Positives = 418/762 (54%), Gaps = 54/762 (7%)
Query: 59 FKGRFDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFAEEAEYWWINAKGRLELGGDV 118
F G DP A W ++R F++ +D + +A H +A WW+ + R G +V
Sbjct: 182 FMGSTDPIVADEWRSRLKRNFKSTRCPEDYQRDIAVHFLEGDAHNWWLTVEKRR--GDEV 239
Query: 119 VTWAKFKVEFLRKYFPEDLRTKKEVEFLNLKQGIMSVAEYAAKFEELSRFCPY-INAEDA 177
++A F+ EF +KYFP + + E +L+L QG +V EY +F L R+ + E A
Sbjct: 240 RSFADFEDEFNKKYFPPEAWDRLECAYLDLVQGNRTVREYDEEFNRLRRYVGRELEEEQA 299
Query: 178 MVSKCVKFESGLRPVIYQYMCVQEIRDFDTLVHKCRMFDDAGKAKSNYYKVQNEKKGKGH 237
+ + ++ GLR I + V+ LV + M ++ G + Y N +K
Sbjct: 300 QLRRFIR---GLRIEIRNHCLVRTFNSVSELVERAAMIEE-GIEEERYL---NREKAPIR 352
Query: 238 GVGKPYSKDKGKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLN-CHKCGKAGHKAV 296
DK +K + ++ +C CG +++ C + C +CG H
Sbjct: 353 NNQSTKPADKKRKFDKVDNTKSDAKTGECVTCGK--NHSGTCWKAIGACGRCGSKDHAIQ 410
Query: 297 DC--------RVVAREI-TCYNCGEKGHISTKPKKAAGKVFALNAKEVEQPDNLIRGMCF 347
C +V+ E TC+ CG+ GH+ + K + K+ Q DN RG
Sbjct: 411 SCPKMEPGQSKVLGEETRTCFYCGKTGHLKRECPKLTAE------KQAGQRDN--RGGNG 462
Query: 348 INSTPLIAIIDTGATHSFISASCVERLGLVVTPLLR------GMVIDTPASGSGTTSFMC 401
+ P + + A+ + + GMV T +
Sbjct: 463 LPPPPKRQAVASRVYELSEEANDAGNFRAITGGFRKEPNTDYGMVRAAGGQAMYPTGLVR 522
Query: 402 AKCPVNFGDVDFKLDLVCLPLKNMDVIFGMDWLLAFGVSINCLTKSVTFSKPVEEL---- 457
V G V+ DL+ +PLK DVI GMDWL + I+C + F + L
Sbjct: 523 GISVVVNG-VNMPADLIIVPLKKHDVILGMDWLGKYKAHIDCHRGRIQFERDEGMLKFQG 581
Query: 458 -----GREFLTAEQVKKSLDGEACVFMMFASLKVSGEKGASV----LPVVQEFPEVFPED 508
G ++A Q ++ L G+ C + + + E GAS +P+V EF +VF
Sbjct: 582 IRTTSGSLVISAIQAERML-GKGCE--AYLATITTKEVGASAELKDIPIVNEFSDVFAA- 637
Query: 509 ITELPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGA 568
++ +P +R F I+L PGT+PIS APYRM+ +E+ +LKKQLEELL+K FIRPS SP GA
Sbjct: 638 VSGVPPDRSDPFTIELEPGTTPISKAPYRMAPAEMAKLKKQLEELLDKGFIRPSSSPWGA 697
Query: 569 PVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQI 628
PVL VKKKDGS RLC+DYR LNK+T+KN+YPLPRID+ MDQL GA+ FSKIDL SGYHQI
Sbjct: 698 PVLFVKKKDGSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQI 757
Query: 629 RVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILV 688
++ D+ KTAFRTRY H+E+ VMPFG++NAP FM+ MN +F +LD FV++FI+DILV
Sbjct: 758 PIEPTDVRKTAFRTRYDHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFINDILV 817
Query: 689 YSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAV 748
YSKS E H EHLR VL+ L+E++L AKLSKC FW + V FLGHVIS G+SVDP K+ ++
Sbjct: 818 YSKSWEAHQEHLRAVLERLREHELFAKLSKCSFWQRSVGFLGHVISDQGVSVDPEKIRSI 877
Query: 749 LQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLALPLTQLTQK 790
+W P++ EI++ GL GYYRRF+ F+ +A PLT+LT K
Sbjct: 878 KEWPRPRNATEIRSFLGLAGYYRRFVMSFASMAQPLTRLTGK 919
>At2g04670 putative retroelement pol polyprotein
Length = 1411
Score = 402 bits (1034), Expect = e-112
Identities = 202/385 (52%), Positives = 268/385 (69%), Gaps = 11/385 (2%)
Query: 416 DLVCLPLKNMDVIFGMDWLLAFGVSINCLTKSVTFSKPVEEL---------GREFLTAEQ 466
DL+ P++ DVI GMDWL + V ++C V+F +P L G ++A Q
Sbjct: 394 DLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGRLVYQGVRPTSGSLVISAVQ 453
Query: 467 VKKSLDGEACVFMMFASLKVS-GEKGASVLPVVQEFPEVFPEDITELPSEREVEFAIDLV 525
KK ++ +++ S+ S G+ S + V+QEF +VF + + LP R F I+L
Sbjct: 454 AKKMIEKGCEAYLVTISMPESLGQVAVSDIRVIQEFEDVF-QSLQGLPPSRSDPFTIELE 512
Query: 526 PGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVD 585
PGT+P+S APYRM+ +E+ ELKKQLE+LL K FIRPS SP GAPVL VKKKDGS RLC+D
Sbjct: 513 PGTAPLSKAPYRMAPAEMTELKKQLEDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCID 572
Query: 586 YRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYG 645
YR LN +T+KN+YPLPRID+ +DQL GA FSKIDL SGYHQI + D+ KTAFRTRYG
Sbjct: 573 YRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRYG 632
Query: 646 HYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQ 705
H+E+ VMPF ++NAP FM MN +F +LD FV++FIDDILVYSKS EEH HLR V++
Sbjct: 633 HFEFVVMPFALTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEHEVHLRRVME 692
Query: 706 VLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWG 765
L+E +L AKLSKC FW +E+ FLGH++S G+SVDP K++A+ W P + EI++
Sbjct: 693 KLREQKLFAKLSKCSFWQREIGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFLR 752
Query: 766 LDGYYRRFIEGFSKLALPLTQLTQK 790
L GYYRRF++GF+ +A P+T+LT K
Sbjct: 753 LTGYYRRFVKGFASMAQPMTKLTGK 777
Score = 72.8 bits (177), Expect = 7e-13
Identities = 64/250 (25%), Positives = 99/250 (39%), Gaps = 19/250 (7%)
Query: 46 RRLDRFLRNNPPTFKGRFDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFAEEAEYWW 105
R +++ R F PE A +W ++R F + + +V LA H +A WW
Sbjct: 125 RMMEQLQRIGTWYFSSGTSPEEADSWRSRVQRNFGSSRCPAEYRVDLAVHFLEGDAHLWW 184
Query: 106 INAKGRLELGGDVVTWAKFKVEFLRKYFPEDLRTKKEVEFLNLKQGIMSVAEYAAKFEEL 165
+ R ++WA F EF KYFP++ + E FL L QG SV EY +F L
Sbjct: 185 RSVTARRRQAD--MSWADFVAEFNAKYFPQEALDRMEARFLELTQGEWSVREYDREFNRL 242
Query: 166 SRFCPYINAEDAMVSKCVKFESGLRPVIYQYMCVQEIRDFDTLVHKCRMFDD------AG 219
+ +D ++ +F GLRP + V + LV ++ G
Sbjct: 243 LAYAGRGMEDDQ--AQMRRFLRGLRPDLRVQCRVSQYATKAALVETAAEVEEDLQRQVVG 300
Query: 220 KAKSNYYKVQNEKKGKGHGVGKPYSKDKGK---KRESGGGSRPSLADVKCYRCGTLGHYA 276
+ + K ++ G GKP K K +G G R + CG+L H
Sbjct: 301 VSPAVQTKNTQQQVTPSKG-GKPAQGQKRKWDHPSRAGQGGRAGY-----FSCGSLDHTV 354
Query: 277 NECMNDLNCH 286
+C + H
Sbjct: 355 ADCTERGHLH 364
>At4g10580 putative reverse-transcriptase -like protein
Length = 1240
Score = 399 bits (1024), Expect = e-111
Identities = 201/385 (52%), Positives = 269/385 (69%), Gaps = 11/385 (2%)
Query: 416 DLVCLPLKNMDVIFGMDWLLAFGVSINCLTKSVTFSKPVEEL---------GREFLTAEQ 466
DL+ P++ DVI GMDWL + V ++ V F +P L G ++A Q
Sbjct: 368 DLIISPVELYDVILGMDWLDYYRVHLDWHRGRVFFERPEGRLVYQGVRPISGSLVISAVQ 427
Query: 467 VKKSLDGEACVFMMFASLKVS-GEKGASVLPVVQEFPEVFPEDITELPSEREVEFAIDLV 525
+K ++ +++ S+ S G+ S + VVQEF +VF + + LP + F I+L
Sbjct: 428 AEKMIEKGCEAYLVTISMPESVGQVAVSDIRVVQEFQDVF-QSLQGLPPSQSDPFTIELE 486
Query: 526 PGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVD 585
PGT+P+S APYRM+ +E+ ELKKQL++LL K FIRPS SP GAPVL VKKKDGS RLC+D
Sbjct: 487 PGTAPLSKAPYRMAPAEMAELKKQLKDLLGKGFIRPSTSPWGAPVLFVKKKDGSFRLCID 546
Query: 586 YRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYG 645
YR+LN++T+KNRYPLPRID+ +DQL GA FSKIDL SGYHQI + D+ KTAFRTRYG
Sbjct: 547 YRELNRVTVKNRYPLPRIDELLDQLRGATCFSKIDLTSGYHQIPIAEADVRKTAFRTRYG 606
Query: 646 HYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQ 705
H+E+ VMPFG++NAP VFM MN +F +LD FV++FIDDILVYSKS EE HLR V++
Sbjct: 607 HFEFVVMPFGLTNAPAVFMRLMNSVFQEFLDEFVIIFIDDILVYSKSPEEQEVHLRRVME 666
Query: 706 VLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWG 765
L+E +L AKLSKC FW +E+ FLGH++S G+SVDP K++A+ W P + EI++ G
Sbjct: 667 KLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWPRPTNATEIRSFLG 726
Query: 766 LDGYYRRFIEGFSKLALPLTQLTQK 790
GYYRRF++GF+ +A P+T+LT K
Sbjct: 727 WAGYYRRFVKGFASMAQPMTKLTGK 751
Score = 48.9 bits (115), Expect = 1e-05
Identities = 27/91 (29%), Positives = 42/91 (45%), Gaps = 2/91 (2%)
Query: 46 RRLDRFLRNNPPTFKGRFDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFAEEAEYWW 105
R +++ R + F G PE A +W + R F + + +V LA H +A WW
Sbjct: 139 RMMEQLQRIDTGYFSGGTSPEEADSWRSRVGRNFGSSRCPAEYRVDLAVHFLEGDAHLWW 198
Query: 106 INAKGRLELGGDVVTWAKFKVEFLRKYFPED 136
+ R ++WA F EF KYFP++
Sbjct: 199 RSVTARRRQTD--MSWADFVAEFKAKYFPQE 227
>At2g07660 putative retroelement pol polyprotein
Length = 949
Score = 397 bits (1020), Expect = e-110
Identities = 208/393 (52%), Positives = 273/393 (68%), Gaps = 17/393 (4%)
Query: 411 VDFKLDLVCLPLKNMDVIFGMDWLLAFGVSINCLTKSVTFSKPVEEL---------GREF 461
V+ DL+ +PLK DVI GMDWL + I+C + + F + L G
Sbjct: 26 VNMPADLIIVPLKKHDVILGMDWLGKYKGHIDCHRERIQFERDEGMLKFQGIRTTSGSLV 85
Query: 462 LTAEQVKKSLDGEACVFMMFASLKVSGEKGASV----LPVVQEFPEVFPEDITELPSERE 517
++A Q ++ L G+ C + + + E GAS + +V EF +VF ++ +P +R
Sbjct: 86 ISAIQAERML-GKGCE--AYLATITTKEVGASAELKDILIVNEFSDVFAA-VSGVPPDRS 141
Query: 518 VEFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKD 577
F I+L PGT+PIS APYRM+ +E+ ELKKQLEELL K FIRPS SP GAPVL VKKKD
Sbjct: 142 DPFTIELEPGTTPISKAPYRMAPAEMAELKKQLEELLAKGFIRPSSSPWGAPVLFVKKKD 201
Query: 578 GSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISK 637
GS RLC+DYR LNK+T+KN+YPLPRID+ MDQL GA+ FSKIDL SGYHQI ++ D+ K
Sbjct: 202 GSFRLCIDYRGLNKVTVKNKYPLPRIDELMDQLGGAQWFSKIDLASGYHQIPIEPTDVRK 261
Query: 638 TAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHA 697
TAFRTRYGH+E+ VMPFG++NAP FM+ MN +F +LD FV++FIDDILV+SKS E H
Sbjct: 262 TAFRTRYGHFEFVVMPFGLTNAPAAFMKMMNGVFRDFLDEFVIIFIDDILVHSKSWEAHQ 321
Query: 698 EHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSV 757
EHLR VL+ L+E++L AKLSK FW + V FLGHVIS G+SVDP K+ ++ +W P++
Sbjct: 322 EHLRAVLERLREHELFAKLSKFSFWQRSVGFLGHVISDQGVSVDPEKIRSIKEWPRPRNA 381
Query: 758 FEIKAIWGLDGYYRRFIEGFSKLALPLTQLTQK 790
EI++ GL GYYRRF+ F+ +A PLT+LT K
Sbjct: 382 TEIRSFLGLAGYYRRFVMSFASMAQPLTRLTGK 414
>At2g14650 putative retroelement pol polyprotein
Length = 1328
Score = 369 bits (947), Expect = e-102
Identities = 199/447 (44%), Positives = 267/447 (59%), Gaps = 24/447 (5%)
Query: 344 GMCFINSTPLIAIIDTGATHSFISASCVERLGLVVTPLLRGMVIDTPASGSGTTSFMCAK 403
G + + D+GA+H FI+ R + P + +
Sbjct: 304 GTLLVGGVEAHVLFDSGASHCFITPESASRGNIRGDPGEQLGAVKVAGGQFVAVLGRTKG 363
Query: 404 CPVNFGDVDFKLDLVCLPLKNMDVIFGMDWLLAFGVSINCLTKSVTFSKPVEELGREFLT 463
+ DL+ P++ DVI GMDWL + V ++C V+F +P L
Sbjct: 364 VDIQIAGESMPADLIISPVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGSL------ 417
Query: 464 AEQVKKSLDGEACVFMMFASLKVSGEKGASVLPVVQEFPEVFPEDITELPSEREVEFAID 523
Q + G + + A + EKG EF +VF + + LP R F I+
Sbjct: 418 VYQGVRPTSGSLVISAVQAEKMI--EKGCEAY---LEFEDVF-QSLQGLPPSRSDPFTIE 471
Query: 524 LVPGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLC 583
L PGT+P+S APYRM+ +E+ ELKKQLE+LL GAPVL VKKKDGS RLC
Sbjct: 472 LEPGTAPLSKAPYRMAPAEMAELKKQLEDLL------------GAPVLFVKKKDGSFRLC 519
Query: 584 VDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTR 643
+DYR LN +T+KN+YPLPRID+ +DQL GA FSKIDL SGYH I + D+ KTAFRTR
Sbjct: 520 IDYRGLNWVTVKNKYPLPRIDELLDQLRGATCFSKIDLTSGYHLIPIAEADVRKTAFRTR 579
Query: 644 YGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIV 703
YGH+E+ VMPFG++NAP FM MN +F LD FV++FIDDILVYSKS EEH HLR V
Sbjct: 580 YGHFEFVVMPFGLTNAPAAFMRLMNSVFQEVLDEFVIIFIDDILVYSKSLEEHEVHLRRV 639
Query: 704 LQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAI 763
++ L+E +L AKLSKC FW +E+ FLGH++S G+SVDP K++A+ W +P + EI++
Sbjct: 640 MEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWHTPTNATEIRSF 699
Query: 764 WGLDGYYRRFIEGFSKLALPLTQLTQK 790
GL GYYRRF++GF+ +A P+T+LT K
Sbjct: 700 LGLAGYYRRFVKGFASMAQPMTKLTGK 726
Score = 62.8 bits (151), Expect = 7e-10
Identities = 41/142 (28%), Positives = 62/142 (42%), Gaps = 7/142 (4%)
Query: 15 IAQALATMAQVLAQGNENTAANRRNEGEAEE-----RRLDRFLRNNPPTFKGRFDPEGAQ 69
+ + LA + V A A ++ AEE R +++ R F G PE A
Sbjct: 107 LLERLAGVVPVEAPVAPRVAEVQQRAAVAEEVPSYLRMMEQLQRIGTGYFSGGTSPEVAD 166
Query: 70 TWVQGMERIFRAMVTGDDQKVRLATHMFAEEAEYWWINAKGRLELGGDVVTWAKFKVEFL 129
+W +ER F + + ++ LA H +A WW + R ++WA F EF
Sbjct: 167 SWRSRVERNFGSSRCPAEYRIDLAVHFLEGDAHLWWRSVTARRRQAD--ISWADFVAEFN 224
Query: 130 RKYFPEDLRTKKEVEFLNLKQG 151
KYFP++ + E FL L QG
Sbjct: 225 AKYFPQEALDRMEAHFLELTQG 246
>At1g36590 hypothetical protein
Length = 1499
Score = 334 bits (857), Expect = 1e-91
Identities = 257/807 (31%), Positives = 391/807 (47%), Gaps = 100/807 (12%)
Query: 44 EERRLDRFLRNNPPTFKG-----RFDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFA 98
++ R D F N T G RFD + W+ +E F T +D KV++A F
Sbjct: 93 QQVRSDHFNAYNNLTRLGKIDFPRFDGTRLKEWLFKVEEFFGVDSTPEDMKVKMAAIHFD 152
Query: 99 EEAEYW---WINAKGRLELGGDVVTWAKFKVEFLRKYFPEDLRTKK-EVEFLNLKQGIMS 154
A W +I + LE+ D W + V+ L++ F +D E++ L GI+
Sbjct: 153 SHASTWHQSFIQSGVGLEVLYD---WKGY-VKLLKERFEDDCDDPMAELKHLQETDGII- 207
Query: 155 VAEYAAKFEELSRFCPYIN-AEDAMVSKCV---KFESGLRPVIYQYMCVQEIRDFDTLVH 210
+Y KFE + +N +E+ +VS + + ++ + ++Q V+
Sbjct: 208 --DYHQKFELIKT---RVNLSEEYLVSVYLAGLRTDTQMHVRMFQPQTVRHCLFLGKTYE 262
Query: 211 KCRMFDDAGKAKSN--------YYKVQNEKKGKGHGVG-----KPYSKDKGKKRESGGGS 257
K + A S Y K Q E + K G KP S+ K +
Sbjct: 263 KAHLKKPANTTWSTNRSAPTGGYNKYQKEGESKTDHYGNKGNFKPVSQQPKKMSQQEMSD 322
Query: 258 RPSLADVKCYRCG---TLGHYANECMNDLNCHKCGKAGHKAVDCRVVAREITCYNCGEKG 314
R S CY C T HY HK + VD N ++
Sbjct: 323 RRSKG--LCYFCDEKYTPEHYL--------VHKKTQLFRMDVDEEFEDAREELVNDDDE- 371
Query: 315 HISTKPKKAAGKVFALNAKEVEQPDNLIRGMCFINSTPLIAIIDTGATHSFISASCVERL 374
H+ A + V+ + + + FI +ID+G+TH+F+ + +L
Sbjct: 372 HMPQISVNAVSGIAGYKTMRVKGTYD--KKIIFI-------LIDSGSTHNFLDPNTAAKL 422
Query: 375 GLVVTPLLRGMVIDTPASG-----SGTTSFMCAKCPVNFGDVDFKLDLVCLPLKNMDVIF 429
G V G+ + A G G + K F+ D++ +PL+ +D++
Sbjct: 423 GCKVDTA--GLTRVSVADGRKLRVEGKVTDFSWKLQTT----TFQSDILLIPLQGIDMVL 476
Query: 430 GMDWLLAFG-------------------VSINCLTKSVTFSKPVEELGR--------EFL 462
G+ WL G V ++ LT ++L + L
Sbjct: 477 GVQWLETLGRISWEFKKLEMRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQEDQVQLAML 536
Query: 463 TAEQVKKSLDGEACVFMMFASLKVSGEKGASVLPVVQEFPEVFPEDITELPSEREVEFAI 522
++V +S +GE C S GE+ V V+ E+P++F E P + I
Sbjct: 537 CVQEVSESTEGELCTINALTS--ELGEESV-VEEVLNEYPDIFIEPTALPPFREKHNHKI 593
Query: 523 DLVPGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRL 582
L+ G++P++ PYR S + E+ K +E+LL ++ S SP +PV+LVKKKDG+ RL
Sbjct: 594 KLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSPYASPVVLVKKKDGTWRL 653
Query: 583 CVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRT 642
CVDYR+LN +T+K+ +P+P I+D MD+L GA +FSKIDLR+GYHQ+R+ +DI KTAF+T
Sbjct: 654 CVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKTAFKT 713
Query: 643 RYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRI 702
GH+EY VMPFG++NAP F MN IF P+L +FV+VF DDILVYS S EEH +HL+
Sbjct: 714 HSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQHLKQ 773
Query: 703 VLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKA 762
V +V++ N+L AKLSKC F + +V +LGH IS GI DP+K+ AV +W P ++ +++
Sbjct: 774 VFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLKQLRG 833
Query: 763 IWGLDGYYRRFIEGFSKLALPLTQLTQ 789
GL GYYRRF+ F +A PL LT+
Sbjct: 834 FLGLAGYYRRFVRSFGVIAGPLHALTK 860
>At3g11970 hypothetical protein
Length = 1499
Score = 333 bits (854), Expect = 2e-91
Identities = 257/810 (31%), Positives = 393/810 (47%), Gaps = 106/810 (13%)
Query: 44 EERRLDRFLRNNPPTFKG-----RFDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFA 98
++ R D F N T G RFD + W+ +E F T +D KV++A F
Sbjct: 93 QQVRSDHFNAYNNLTRLGKIDFPRFDGTRLKEWLFKVEEFFGVDSTPEDMKVKMAAIHFD 152
Query: 99 EEAEYW---WINAKGRLELGGDVVTWAKFKVEFLRKYFPEDLRTKK-EVEFLNLKQGIMS 154
A W +I + LE+ D W + V+ L++ F +D E++ L GI+
Sbjct: 153 SHASTWHQSFIQSGVGLEVLYD---WKGY-VKLLKERFEDDCDDPMAELKHLQETDGII- 207
Query: 155 VAEYAAKFEELSRFCPYIN-AEDAMVSKCVKFESGLRPVIYQYMCVQEIRDFDTLVHKCR 213
+Y KFE + +N +E+ +VS + +GLR ++ + + + + +
Sbjct: 208 --DYHQKFELIKT---RVNLSEEYLVSV---YLAGLRTDTQMHVRMFQPQTVRHCLFLGK 259
Query: 214 MFDDAGKAK--------------SNYYKVQNEKKGKGHGVG-----KPYSKDKGKKRESG 254
++ A K Y K Q E + K G KP S+ K +
Sbjct: 260 TYEKAHPKKPANTTWSTNRSAPTGGYNKYQKEGESKTDHYGNKGNFKPVSQQPKKMSQQE 319
Query: 255 GGSRPSLADVKCYRCG---TLGHYANECMNDLNCHKCGKAGHKAVDCRVVAREITCYNCG 311
R S CY C T HY HK + VD N
Sbjct: 320 MSDRRSKG--LCYFCDEKYTPEHYL--------VHKKTQLFRMDVDEEFEDAREELVNDD 369
Query: 312 EKGHISTKPKKAAGKVFALNAKEVEQPDNLIRGMCFINSTPLIAIIDTGATHSFISASCV 371
++ H+ A + V+ + + + FI +ID+G+TH+F+ +
Sbjct: 370 DE-HMPQISVNAVSGIAGYKTMRVKGTYD--KKIIFI-------LIDSGSTHNFLDPNTA 419
Query: 372 ERLGLVVTPLLRGMVIDTPASG-----SGTTSFMCAKCPVNFGDVDFKLDLVCLPLKNMD 426
+LG V G+ + A G G + K F+ D++ +PL+ +D
Sbjct: 420 AKLGCKVDTA--GLTRVSVADGRKLRVEGKVTDFSWKLQTT----TFQSDILLIPLQGID 473
Query: 427 VIFGMDWLLAFG-------------------VSINCLTKSVTFSKPVEELGR-------- 459
++ G+ WL G V ++ LT ++L +
Sbjct: 474 MVLGVQWLETLGRISWEFKKLEMRFKFNNQKVLLHGLTSGSVREVKAQKLQKLQEDQVQL 533
Query: 460 EFLTAEQVKKSLDGEACVFMMFASLKVSGEKGASVLPVVQEFPEVFPEDITELPSEREVE 519
L ++V +S +GE C S GE+ V V+ E+P++F E P +
Sbjct: 534 AMLCVQEVSESTEGELCTINALTS--ELGEESV-VEEVLNEYPDIFIEPTALPPFREKHN 590
Query: 520 FAIDLVPGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGS 579
I L+ G++P++ PYR S + E+ K +E+LL ++ S SP +PV+LVKKKDG+
Sbjct: 591 HKIKLLEGSNPVNQRPYRYSIHQKNEIDKLVEDLLTNGTVQASSSPYASPVVLVKKKDGT 650
Query: 580 MRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTA 639
RLCVDYR+LN +T+K+ +P+P I+D MD+L GA +FSKIDLR+GYHQ+R+ +DI KTA
Sbjct: 651 WRLCVDYRELNGMTVKDSFPIPLIEDLMDELGGAVIFSKIDLRAGYHQVRMDPDDIQKTA 710
Query: 640 FRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEH 699
F+T GH+EY VMPFG++NAP F MN IF P+L +FV+VF DDILVYS S EEH +H
Sbjct: 711 FKTHSGHFEYLVMPFGLTNAPATFQGLMNFIFKPFLRKFVLVFFDDILVYSSSLEEHRQH 770
Query: 700 LRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFE 759
L+ V +V++ N+L AKLSKC F + +V +LGH IS GI DP+K+ AV +W P ++ +
Sbjct: 771 LKQVFEVMRANKLFAKLSKCAFAVPKVEYLGHFISAQGIETDPAKIKAVKEWPQPTTLKQ 830
Query: 760 IKAIWGLDGYYRRFIEGFSKLALPLTQLTQ 789
++ GL GYYRRF+ F +A PL LT+
Sbjct: 831 LRGFLGLAGYYRRFVRSFGVIAGPLHALTK 860
>At1g35370 hypothetical protein
Length = 1447
Score = 291 bits (744), Expect = 1e-78
Identities = 173/474 (36%), Positives = 267/474 (55%), Gaps = 49/474 (10%)
Query: 348 INSTPLIAIIDTGATHSFISASC-------VERLGLVVTPLLRGMVIDTPASGSGTTSFM 400
++ L +ID+G+TH+FI ++ VE GL + G ++ G T
Sbjct: 375 VDKRDLFILIDSGSTHNFIDSTVAAKLGCHVESAGLTKVAVADGRKLNVDGQIKGFTW-- 432
Query: 401 CAKCPVNFGDVDFKLDLVCLPLKNMDVIFGMDWLLAFG-VSINCLTKSVTFSKPVEEL-- 457
F+ D++ +PL+ +D++ G+ WL G +S + F + +
Sbjct: 433 ------KLQSTTFQSDILLIPLQGVDMVLGVQWLETLGRISWEFKKLEMQFFYKNQRVWL 486
Query: 458 ------GREFLTAEQVKKSLDGEACVFMMFASLKVSGEKG---------------ASVLP 496
+ A +++K+ + + M+ VS E+ + V
Sbjct: 487 HGIITGSVRDIKAHKLQKTQADQIQLAMVCVREVVSDEEQEIGSISALTSDVVEESVVQN 546
Query: 497 VVQEFPEVFPEDITELPSEREV-EFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLE 555
+V+EFP+VF E T+LP RE + I L+ G +P++ PYR + E+ K ++++++
Sbjct: 547 IVEEFPDVFAEP-TDLPPFREKHDHKIKLLEGANPVNQRPYRYVVHQKDEIDKIVQDMIK 605
Query: 556 KQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEV 615
I+ S SP +PV+LVKKKDG+ RLCVDY +LN +T+K+R+ +P I+D MD+L G+ V
Sbjct: 606 SGTIQVSSSPFASPVVLVKKKDGTWRLCVDYTELNGMTVKDRFLIPLIEDLMDELGGSVV 665
Query: 616 FSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYL 675
FSKIDLR+GYHQ+R+ +DI KTAF+T GH+EY VM FG++NAP F MN +F +L
Sbjct: 666 FSKIDLRAGYHQVRMDPDDIQKTAFKTHNGHFEYLVMLFGLTNAPATFQSLMNSVFRDFL 725
Query: 676 DRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISK 735
+FV+VF DDIL+YS S EEH EHLR+V +V++ ++L AK SK LGH IS
Sbjct: 726 RKFVLVFFDDILIYSSSIEEHKEHLRLVFEVMRLHKLFAKGSK--------EHLGHFISA 777
Query: 736 GGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLALPLTQLTQ 789
I DP+K+ AV +W +P +V +++ G GYYRRF+ F +A PL LT+
Sbjct: 778 REIETDPAKIQAVKEWPTPTTVKQVRGFLGFAGYYRRFVRNFGVIAGPLHALTK 831
>At2g06890 putative retroelement integrase
Length = 1215
Score = 283 bits (725), Expect = 2e-76
Identities = 221/727 (30%), Positives = 331/727 (45%), Gaps = 120/727 (16%)
Query: 130 RKYFPEDLRTKKEVEFLNLKQGIMSVAEYAAKFEELSRFCPYINAEDAMVSKCVKFESGL 189
+++ P + ++ NL QG SV EY + E L +A +S+ F L
Sbjct: 7 KRFVPSHYHRELHLKLRNLTQGNRSVEEYYKEMETLMLRADISEDREATLSR---FLGDL 63
Query: 190 RPVIYQYMCVQEIRDFDTLVHKCRMFDDAGKAKSNYYK-----------VQNEKKGKGH- 237
I + Q + ++HK +F+ K KS+ Q E++ +
Sbjct: 64 NRDIQDRLETQYYVQIEEMLHKAILFEQQVKRKSSSRSSYGSGTIAKPTYQREERTSSYH 123
Query: 238 --------GVGKPYSK---DKGKKRESGGGSRPSLADVKCYRCGTLGHYANECMNDLNCH 286
KPY+ KGK S R DV+CY+C GHYANEC N
Sbjct: 124 NKPIVSPRAESKPYAAVQDHKGKAEISTSRVR----DVRCYKCQGKGHYANECPN----- 174
Query: 287 KCGKAGHKAVDCRVVAREITCYNCGEKGHISTKPKK---------AAGKVFALNAK--EV 335
K V + EI S K + A + ++ K E
Sbjct: 175 -------KRVMILLDNGEIEPEEEIPDSPSSLKENEELPAQGELLVARRTLSVQTKTDEQ 227
Query: 336 EQPDNLIRGMCFINSTPLIAIIDTGATHSFISASCVERLGL--------------VVTPL 381
EQ NL C ++ IID G+ + S + V++LGL VV P+
Sbjct: 228 EQRKNLFHTRCHVHGKVCSLIIDGGSCTNVASETMVKKLGLKWLNDSGKMRVKNQVVVPI 287
Query: 382 LRGMVIDTPASGSGTTSFMCAKCPVNFGDV--------DFKLDLVCLPLKNMDVIFGMDW 433
+ G D +C P+ G + D K+ ++ G
Sbjct: 288 VIGKYED---------EILCDVLPMEAGHILLGRPWQSDRKVMHDGFTNRHSFEFKGGKT 338
Query: 434 LLAFGVSINCLTKSVTFSKPVEELGRE---FLTAEQVKKSLDG-EACVFMMFASLKVSGE 489
+L + + E++ ++ F + +VK + + + +F S
Sbjct: 339 ILVSMTPHEVYQDQIHLKQKKEQVVKQPNFFAKSGEVKSAYSSKQPMLLFVFKEALTSLT 398
Query: 490 KGASVLP-----VVQEFPEVFPEDITE-LPSEREVEFAIDLVPGTSPISIAPYRMSASEL 543
A VLP ++Q++ +VFPED + LP R +E ID VPG S + YR + E
Sbjct: 399 NFAPVLPSEMTSLLQDYKDVFPEDNPKGLPPIRGIEHQIDFVPGASLPNRPAYRTNPVET 458
Query: 544 GELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRI 603
EL++Q KDGS R+C D R +N +T+K +P+PR+
Sbjct: 459 KELQRQ--------------------------KDGSWRMCFDCRAINNVTVKYCHPIPRL 492
Query: 604 DD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVF 663
DD +D+L G+ +FSKIDL+SGYHQIR+ D KTAF+T++G YE+ VMPFG+++AP F
Sbjct: 493 DDMLDELHGSSIFSKIDLKSGYHQIRMNEGDEWKTAFKTKHGLYEWLVMPFGLTHAPSTF 552
Query: 664 MEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWL 723
M MN + ++ FV+V+ DDILVYS+S EH EHL VL VL++ +L A L KC F
Sbjct: 553 MRLMNHVLRAFIGIFVIVYFDDILVYSESLREHIEHLDSVLNVLRKEELYANLKKCTFCT 612
Query: 724 KEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRFIEGFSKLALP 783
+ FLG V+S G+ VD KV A+ W SPK+V E+++ GL G+YRRF + FS + P
Sbjct: 613 DNLVFLGFVVSADGVKVDEEKVKAIRDWPSPKTVGEVRSFHGLAGFYRRFFKDFSTIVAP 672
Query: 784 LTQLTQK 790
LT++ +K
Sbjct: 673 LTEVMKK 679
>At2g05610 putative retroelement pol polyprotein
Length = 780
Score = 281 bits (720), Expect = 8e-76
Identities = 136/277 (49%), Positives = 190/277 (68%)
Query: 497 VVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLEK 556
VV EFP++F E P + I+L+ ++P++ PYR + E+ K ++++L
Sbjct: 10 VVTEFPDIFVEPTDLPPFRAPHDHKIELLEDSNPVNQRPYRYVVHQKNEIDKIVDDMLAS 69
Query: 557 QFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVF 616
I+ S SP +PV+LVKKKDG+ RLCVDYR+LN +T+K+R+P+P I+D MD+L G+ V+
Sbjct: 70 GTIQASSSPYASPVVLVKKKDGTWRLCVDYRELNGMTVKDRFPIPLIEDLMDELGGSNVY 129
Query: 617 SKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLD 676
SKIDLR+GYHQ+R+ DI KTAF+T GHYEY VMPFG++NAP F MN F P+L
Sbjct: 130 SKIDLRAGYHQVRMDPLDIHKTAFKTHNGHYEYLVMPFGLTNAPASFQSLMNSFFKPFLR 189
Query: 677 RFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKG 736
+FV+VF DDIL+YS S EEH +HL V +V++ + L AK+SKC F + V +LGH IS
Sbjct: 190 KFVLVFFDDILIYSTSMEEHKKHLEAVFEVMRVHHLFAKMSKCAFAVPRVEYLGHFISGE 249
Query: 737 GISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRF 773
GI+ DP+K+ AV W P ++ ++ GL GYYRRF
Sbjct: 250 GIATDPAKIKAVQDWPVPVNLKQLCGFLGLTGYYRRF 286
>At1g36120 putative reverse transcriptase gb|AAD22339.1
Length = 1235
Score = 263 bits (672), Expect = 3e-70
Identities = 202/589 (34%), Positives = 283/589 (47%), Gaps = 78/589 (13%)
Query: 124 FKVEFLRKYFPEDLRTKKEVEFLNLKQGIMSVAEYAAKFEELSRFCPYINAEDAMVSKCV 183
F EF KYFP++ + E FL L +G +SV EY +F+ L + +D +
Sbjct: 157 FVAEFNAKYFPQEALDRMEARFLELTKGELSVREYGREFKRLLVYAGRGMEDDQAQMR-- 214
Query: 184 KFESGLRPVIYQYMCVQEIRDFDTLVHKCRMFDDAGKAKSNYYKVQNEKKGKGHGVGKPY 243
+F GLRP L +CR+ A KA +V+ + + + V
Sbjct: 215 RFLRGLRP---------------DLRVRCRVSQYATKAALTAAEVEEDLQRQVVAVSPSV 259
Query: 244 SKDKGKKR--ESGGGSRPSLADVKC---YRCGTLGHYANECMNDLNCHKCGKAGHKAVDC 298
K +++ S GG + K +R G G A +D + G A A
Sbjct: 260 QPKKIQQQVAPSKGGKPAQVQKRKWDHPFRAGQGGR-AGSAADDSSTSTAGGAAQSAGAA 318
Query: 299 RVVAREITCYNCGEKGHISTKPKKAAGKVFALNAKEVEQPDNLIRGMCFINSTPLIAIID 358
R A E +ST +A +LI G C + + D
Sbjct: 319 RSAA---------ECAAVSTYCSGTTWLYYA----------DLISGRCRGH-----VLFD 354
Query: 359 TGATHSFISASCVERLGLVVTPLLRGMVIDTPASGSGTTSFMCAK-CPVNFGDVDFKLDL 417
+GA+H FI+ R + P + + A G T AK + + DL
Sbjct: 355 SGASHCFITPESASRDNIRGEPGEQFGAVKI-AGGQFLTVLGRAKGVNIQIEEESMPADL 413
Query: 418 VCLPLKNMDVIFGMDWLLAFGVSINCLTKSVTFSKP--------VEELGREFLTAE-QVK 468
+ P+ D I GMDWL + V ++C V+F +P V R + +E QV+
Sbjct: 414 IINPVVLYDAILGMDWLDHYRVHLDCHRGRVSFERPEGRLVYQGVRPTSRSLVISEVQVE 473
Query: 469 KSLDGEACVFMMFASLKVS-GEKGASVLPVVQEFPEVFPEDITELPSEREVEFAIDLVPG 527
K ++ +++ S+ S G+ S + VVQEF +VF + + LP R F I+L G
Sbjct: 474 KMIEKGCEAYLVTISMSESVGQVAVSDIWVVQEFEDVF-QSLQGLPPSRSDPFTIELELG 532
Query: 528 TSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYR 587
T+P+S PYRM +E+ ELKKQLE+LL K FIRPS S GAP
Sbjct: 533 TAPLSKTPYRMVPAEIAELKKQLEDLLGKGFIRPSTSRWGAP------------------ 574
Query: 588 QLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHY 647
LN++T+KN+YPLPRID+ +DQL GA FSKIDL GYHQ + D+ KTAFRTRYGH+
Sbjct: 575 GLNRVTLKNKYPLPRIDELLDQLRGATCFSKIDLTPGYHQFPIAEADVRKTAFRTRYGHF 634
Query: 648 EYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEH 696
E+ VMPFG++NAP M MN +F +LD FV++FIDDILVY KS EEH
Sbjct: 635 EFVVMPFGLTNAPTALMRLMNSVFQEFLDEFVIIFIDDILVYFKSPEEH 683
>At4g03650 putative reverse transcriptase
Length = 839
Score = 207 bits (526), Expect = 2e-53
Identities = 92/157 (58%), Positives = 121/157 (76%)
Query: 634 DISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSE 693
D+ KTAFRTRYGH+E+ VMPFG++NAP FM MN +F +LD FV++FIDDILVYSKS
Sbjct: 520 DVRKTAFRTRYGHFEFVVMPFGLTNAPAAFMRLMNSVFQEFLDEFVIIFIDDILVYSKSP 579
Query: 694 EEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWES 753
EEH HLR V++ L+E +L AKLSKC FW +E+ FLGH++S G+SVDP K++A+ W
Sbjct: 580 EEHEVHLRRVMEKLREQKLFAKLSKCSFWQREMGFLGHIVSAEGVSVDPEKIEAIRDWPR 639
Query: 754 PKSVFEIKAIWGLDGYYRRFIEGFSKLALPLTQLTQK 790
P + EI++ GL GYYRRFI+GF+ +A P+T+LT K
Sbjct: 640 PTNATEIRSFLGLAGYYRRFIKGFASMAQPMTKLTGK 676
Score = 87.0 bits (214), Expect = 4e-17
Identities = 56/145 (38%), Positives = 83/145 (56%), Gaps = 9/145 (6%)
Query: 416 DLVCLPLKNMDVIFGMDWLLAFGVSINCLTKSVTFSKPVEELGREFLTAEQVKKSLDGEA 475
DL+ ++ DVI GMDWL + V ++C V+F +P GRE + + L G+
Sbjct: 387 DLIISHVELYDVILGMDWLDHYRVHLDCHRGRVSFERPEGSAGRE--NDREGLRGLPGDD 444
Query: 476 CVFMMFASLKVSGEKGASVLPVVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAP 535
++A V G+ S + VVQEF VF + + LP R F I+L P T+P+S AP
Sbjct: 445 ----IYAG--VCGQVAVSDIRVVQEFQYVF-QSLQGLPPSRSDPFTIELEPKTAPLSKAP 497
Query: 536 YRMSASELGELKKQLEELLEKQFIR 560
YRM+ +++ ELKKQLE+LL + +R
Sbjct: 498 YRMAPAKMAELKKQLEDLLAEADVR 522
Score = 73.6 bits (179), Expect = 4e-13
Identities = 46/146 (31%), Positives = 65/146 (44%), Gaps = 4/146 (2%)
Query: 46 RRLDRFLRNNPPTFKGRFDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFAEEAEYWW 105
R + + R F G PE A +W +ER F + + +V L H +A WW
Sbjct: 143 RMMKQLQRIGTEYFSGGTSPEEADSWRSQVERNFGSSRCPAEYRVDLTVHFLEGDAHLWW 202
Query: 106 INAKGRLELGGDVVTWAKFKVEFLRKYFPEDLRTKKEVEFLNLKQGIMSVAEYAAKFEEL 165
+ R ++WA F EF KYFP + + E FL L QG+ SV EY KF L
Sbjct: 203 RSVTARRRQAD--MSWADFMAEFNAKYFPREALDRMEARFLELTQGVRSVREYDRKFNRL 260
Query: 166 SRFCPYINAEDAMVSKCVKFESGLRP 191
+ +D ++ +F GLRP
Sbjct: 261 LVYAGRGMEDDQ--AQMRRFLRGLRP 284
>At4g04230 putative transposon protein
Length = 315
Score = 189 bits (481), Expect = 4e-48
Identities = 105/266 (39%), Positives = 155/266 (57%), Gaps = 43/266 (16%)
Query: 461 FLTAEQVKKSLDGEACVFMM-FASLKVSG----EKGASVLPVVQEFPEVFPEDITELPSE 515
+L+++ V K++ + V +M F +G E A V V+ + ++FPE+I
Sbjct: 65 YLSSKDVCKTMSAKGTVLLMVFKECLSTGIGDPELSAEVQVVLHWYKDLFPEEIP----- 119
Query: 516 REVEFAIDLVPGTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKK 575
PG PI K +IR S+SP PVLLV K
Sbjct: 120 ----------PGLPPI-----------------------HKGYIRESLSPCAVPVLLVPK 146
Query: 576 KDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDI 635
KDG+ R+C+D R +N +TIK R+P+PR+ D +D+L GA +FSK+DLRSGYHQ+R++ D
Sbjct: 147 KDGTWRMCLDCRAINNITIKYRHPIPRLYDMLDELSGAIIFSKVDLRSGYHQVRMREGDE 206
Query: 636 SKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEE 695
KTAF+T+ G YE VMPFG++NAP FM MN++ ++ +FVVV+ DDIL+Y+KS +
Sbjct: 207 WKTAFKTKQGLYECLVMPFGLTNAPSTFMRLMNQVLRSFIGKFVVVYFDDILIYNKSYSD 266
Query: 696 HAEHLRIVLQVLKENQLCAKLSKCEF 721
H + L ++L+ L++ L A L KC F
Sbjct: 267 HIQPLELILKTLRKEGLYANLKKCTF 292
>At4g08100 putative polyprotein
Length = 1054
Score = 184 bits (467), Expect = 2e-46
Identities = 87/175 (49%), Positives = 122/175 (69%)
Query: 585 DYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRY 644
D R +N +T+K R+P+PR+DD +D+L G+ +FSKIDL+SGYHQ R+K D KTA +T+
Sbjct: 527 DCRAINNITVKYRHPIPRLDDTLDKLHGSSIFSKIDLKSGYHQTRMKEGDEWKTAIKTKQ 586
Query: 645 GHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVL 704
YE+ VMPFG++NAP FM MN + ++ FV+V+ DDILVYSK+ E H HL++VL
Sbjct: 587 RLYEWLVMPFGLTNAPNTFMRLMNHVLRKHIGVFVIVYFDDILVYSKNLEYHVMHLKLVL 646
Query: 705 QVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFE 759
+L++ +L A L KC F + FLG V+S GI VD KV A+ +W +PK+V E
Sbjct: 647 DLLRKEKLYANLKKCTFCTDNLVFLGFVVSADGIKVDEEKVKAIREWPNPKNVSE 701
Score = 73.6 bits (179), Expect = 4e-13
Identities = 59/242 (24%), Positives = 97/242 (39%), Gaps = 20/242 (8%)
Query: 57 PTFKGRFDPEGAQTWVQGMERIFRAMVTGDDQKVRLATHMFAEEAEYWW--INAKGRLEL 114
P F G +P+ W Q +E +F KV++A F A WW + R
Sbjct: 97 PAFHGTDNPDTYLEWEQKIELVFLCQECLQSNKVKIAATKFYNYALSWWDQLVTSRRRTR 156
Query: 115 GGDVVTWAKFKVEFLRKYFPEDLRTKKEVEFLNLKQGIMSVAEYAAKFEELSRFCPYINA 174
+ TW + K +++ P + NL QG +V EY + E L
Sbjct: 157 DYPIKTWNQLKFVMRKRFVPSYYHRELHQRLRNLVQGSKTVEEYFLEMETLMLRADLQED 216
Query: 175 EDAMVSKCVKFESGLRPVIYQYMCVQEIRDFDTLVHKCRMFDD--------AGKAKSNYY 226
+A++S +F GL I + Q + + ++HK MF+ + K+NY
Sbjct: 217 GEAVMS---RFMGGLNREIQDRLETQHYVELEEMLHKAVMFEQQIKRKNARSSHTKTNYS 273
Query: 227 ----KVQNEKKGKGHGVGKPYSKDKGKKRESGGGSRPSLA---DVKCYRCGTLGHYANEC 279
Q E+K KP+ K K + G + + D++C++ L HYA++C
Sbjct: 274 SGKPSYQKEEKFGYQNDSKPFVKPKPVDLDPKGKGKEVITRARDIRCFKSQGLRHYASKC 333
Query: 280 MN 281
N
Sbjct: 334 SN 335
>At1g37060 Athila retroelment ORF 1, putative
Length = 1734
Score = 166 bits (420), Expect = 5e-41
Identities = 113/317 (35%), Positives = 161/317 (50%), Gaps = 35/317 (11%)
Query: 509 ITELPSERE-VEFAIDLVPGTSPI--------------SIAPYRMSASELGEL-KKQLEE 552
ITEL R+ + +++D + G SP SI P R L E+ KK++ +
Sbjct: 862 ITELMKYRKAIGYSLDDIKGISPTLCTHRIHLENESYSSIEPQRRLNPNLKEVVKKEILK 921
Query: 553 LLEKQFIRP-SVSPRGAPVLLVKKKDGSM------------------RLCVDYRQLNKLT 593
LL+ I P S S +PV V KK G R+C++YR+LN +
Sbjct: 922 LLDAGVIYPISDSTWVSPVHCVPKKGGMTVVKNSKDELIPTRTITGHRMCIEYRKLNVAS 981
Query: 594 IKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMP 653
K +PLP ID +++L + +D SG+ QI + D KT F YG + Y MP
Sbjct: 982 RKEHFPLPFIDHMLERLANHPYYCFLDSYSGFFQIPIHPNDQGKTTFTCPYGTFAYKRMP 1041
Query: 654 FGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQVLKENQLC 713
FG+ NAP F M IF ++ V VF+DD VY S +L VL+ +E L
Sbjct: 1042 FGLCNAPATFQRCMTSIFSDLIEEMVEVFMDDFSVYGSSFSSCLLNLCRVLKRCEETNLV 1101
Query: 714 AKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWGLDGYYRRF 773
KC F ++E LG IS+ GI VD +K+D ++Q + PK+V +I++ G G+YR F
Sbjct: 1102 LNWEKCHFMVREGIVLGRKISEEGIEVDKAKIDVMMQLQPPKTVKDIRSFLGHAGFYRIF 1161
Query: 774 IEGFSKLALPLTQLTQK 790
I+ FSKLA PLT+L K
Sbjct: 1162 IKDFSKLARPLTRLLCK 1178
>At2g14400 putative retroelement pol polyprotein
Length = 1466
Score = 159 bits (403), Expect = 4e-39
Identities = 87/262 (33%), Positives = 146/262 (55%), Gaps = 1/262 (0%)
Query: 527 GTSPISIAPYRMSASELGELKKQLEELLEKQFIRPSVSPRG-APVLLVKKKDGSMRLCVD 585
G S + ++ + ++++LL+ IR P A ++VKKK+G R+C+D
Sbjct: 461 GISRADVNRRKLGVERAKAVNDEVDKLLKIGSIREVQYPDWLANTVVVKKKNGKDRVCID 520
Query: 586 YRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYG 645
+ LNK K+ +PLP ID ++ G E+ S +D SGY+QI + ED KT F T G
Sbjct: 521 FTDLNKACPKDSFPLPHIDRLVESTAGNELLSFMDAFSGYNQIMMNPEDQEKTLFITDRG 580
Query: 646 HYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHLRIVLQ 705
Y Y VMPFG+ NA + +N++F ++ + + V+IDD+L+ S +E+H +HL
Sbjct: 581 IYCYKVMPFGLRNAGATYPRLVNKMFSEHVGKTMEVYIDDMLIKSLKKEDHVKHLEECFA 640
Query: 706 VLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEIKAIWG 765
+L + Q+ +KC F + FLG++++K GI +P++++A L SPK+ E++ + G
Sbjct: 641 ILNQYQMKLNPAKCTFGVPSGEFLGYIVTKRGIEANPNQINAFLNMPSPKNFKEVQRLTG 700
Query: 766 LDGYYRRFIEGFSKLALPLTQL 787
RFI + +LP Q+
Sbjct: 701 RIAALNRFISRSTDKSLPFYQI 722
>At2g14640 putative retroelement pol polyprotein
Length = 945
Score = 159 bits (402), Expect = 6e-39
Identities = 125/382 (32%), Positives = 174/382 (44%), Gaps = 56/382 (14%)
Query: 316 ISTKPKKAAGKV-FALNAKEVEQPDNLIRGMCFINSTPLIAIIDTGATHSFISASCVERL 374
+ + +A GK L A E + IR I+ LIA+I++G+TH+FI V +
Sbjct: 291 VEEESTEAEGKPEITLYALEGVDTTSTIRVRATIHRNRLIALINSGSTHNFIGEKAVRGM 350
Query: 375 GLVVT---PLLRGMVIDTPASGSGTTSFMCAKCPVNFGDVDFKLDLVCLPLKNMDVIFGM 431
L T P +V P + PV G V F + L LPL +D+ G+
Sbjct: 351 NLKATTTKPFTVRVVNGMPLVCRSRYEAI----PVVMGGVVFPVTLYALPLMGLDLAMGV 406
Query: 432 DWLLAFGVSINCLTKSVTFS-------------KPVEELGREFLTAEQVKKSLDGEACVF 478
WL G ++ C K T KP G E T KK+ G
Sbjct: 407 QWLSTLGPTL-CNWKEQTLQFHWAGDEVRLMGIKPTGLRGVEHKTI--TKKARMGHTIFA 463
Query: 479 MMFA-SLKVSGEKGASVLPVVQEFPEVFPEDITELPSEREVEFAIDLVPGTSPISIAPYR 537
+ A + + +++EF +F E T+LPSER + I L GT P+++ PYR
Sbjct: 464 ITMAHNSSDPNTPDGEIKKLIEEFAGLF-ETPTQLPSERPIVHRIALKKGTDPVNVRPYR 522
Query: 538 MSASELGELKKQLEELLEKQFIRPSVSPRGAPVLLVKKKDGSMRLCVDYRQLNKLTIKNR 597
+ Q +E+ IRPS SP +PVLL R
Sbjct: 523 YAFY-------QKDEMSRAGIIRPSSSPFLSPVLL-----------------------ER 552
Query: 598 YPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAFRTRYGHYEYSVMPFGVS 657
+P+P +DD +D+L GA F+K+DL +GY Q+R+ + DI K AF+T GHYEY VMPFG+
Sbjct: 553 FPIPTVDDMLDELNGAVYFTKLDLTAGYQQVRMHSPDIPKIAFQTHNGHYEYLVMPFGLC 612
Query: 658 NAPGVFMEYMNRIFHPYLDRFV 679
NAP F MN IF P L R V
Sbjct: 613 NAPSTFQALMNEIFWPLLSRSV 634
>At2g15410 putative retroelement pol polyprotein
Length = 1787
Score = 157 bits (396), Expect = 3e-38
Identities = 83/217 (38%), Positives = 123/217 (56%)
Query: 571 LLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRV 630
++VKKK+G R+C+D+ LNK K+ +PLP ID ++ G E+ S +D SGY+QI +
Sbjct: 824 VVVKKKNGKWRVCIDFTDLNKACPKDSFPLPHIDRLVEATAGNELLSFMDAFSGYNQILM 883
Query: 631 KAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYS 690
D KT F T G Y Y VMPFG+ NA + +N++F LD + V+IDD+LV S
Sbjct: 884 HQNDREKTVFITDQGTYCYKVMPFGLKNAGATYPRLVNQMFTDQLDHSMEVYIDDMLVKS 943
Query: 691 KSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQ 750
EEH HLR QVL + SKC F + FLG+++++ GI +P ++ A++
Sbjct: 944 LRAEEHITHLRQCFQVLNRYNMKLNPSKCTFGVTSGEFLGYLVTRRGIEANPKQISAIID 1003
Query: 751 WESPKSVFEIKAIWGLDGYYRRFIEGFSKLALPLTQL 787
SP++ E++ + G RFI + LP QL
Sbjct: 1004 LPSPRNTREVQRLIGRIAALNRFISRSTDKCLPFYQL 1040
>At1g20390 hypothetical protein
Length = 1791
Score = 155 bits (393), Expect = 6e-38
Identities = 90/270 (33%), Positives = 144/270 (53%), Gaps = 1/270 (0%)
Query: 522 IDLVPGTSPISIAPYRMSASELGELKKQLEELLEK-QFIRPSVSPRGAPVLLVKKKDGSM 580
+++ P P+ ++ + +++E+LL+ Q I A ++VKKK+G
Sbjct: 769 LNVDPTFKPVKQKRRKLGPERARAVNEEVEKLLKAGQIIEVKYPEWLANPVVVKKKNGKW 828
Query: 581 RLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRVKAEDISKTAF 640
R+CVDY LNK K+ YPLP ID ++ G + S +D SGY+QI + +D KT+F
Sbjct: 829 RVCVDYTDLNKACPKDSYPLPHIDRLVEATSGNGLLSFMDAFSGYNQILMHKDDQEKTSF 888
Query: 641 RTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYSKSEEEHAEHL 700
T G Y Y VM FG+ NA + ++N++ + R V V+IDD+LV S E+H EHL
Sbjct: 889 VTDRGTYCYKVMSFGLKNAGATYQRFVNKMLADQIGRTVEVYIDDMLVKSLKPEDHVEHL 948
Query: 701 RIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQWESPKSVFEI 760
VL + +KC F + FLG+V++K GI +P ++ A+L+ SP++ E+
Sbjct: 949 SKCFDVLNTYGMKLNPTKCTFGVTSGEFLGYVVTKRGIEANPKQIRAILELPSPRNAREV 1008
Query: 761 KAIWGLDGYYRRFIEGFSKLALPLTQLTQK 790
+ + G RFI + LP L ++
Sbjct: 1009 QRLTGRIAALNRFISRSTDKCLPFYNLLKR 1038
>At2g14380 putative retroelement pol polyprotein
Length = 764
Score = 148 bits (373), Expect = 1e-35
Identities = 76/217 (35%), Positives = 125/217 (57%)
Query: 571 LLVKKKDGSMRLCVDYRQLNKLTIKNRYPLPRIDD*MDQLVGAEVFSKIDLRSGYHQIRV 630
++VKKK+ R+C+D+ LNK K+ +PLP ID ++ G E+ S +D SGY+QI +
Sbjct: 539 VVVKKKNDKWRVCIDFTDLNKACPKDSFPLPHIDRMVEATTGNELLSFMDAFSGYNQIPM 598
Query: 631 KAEDISKTAFRTRYGHYEYSVMPFGVSNAPGVFMEYMNRIFHPYLDRFVVVFIDDILVYS 690
+D KT+F G Y Y VMPFG+ N + +N++F P L + + V+IDD+LV S
Sbjct: 599 HKDDQEKTSFIIDRGTYCYKVMPFGLKNVGARYQRLVNQMFAPQLGKTMEVYIDDMLVKS 658
Query: 691 KSEEEHAEHLRIVLQVLKENQLCAKLSKCEFWLKEVSFLGHVISKGGISVDPSKVDAVLQ 750
+H +HL+ + L + + +KC F + FLG++++K GI +P ++ A+L
Sbjct: 659 TRSADHIDHLKACFETLNKYNMKLNPAKCLFGVTSGEFLGYIVTKRGIEANPKQIRAILD 718
Query: 751 WESPKSVFEIKAIWGLDGYYRRFIEGFSKLALPLTQL 787
+SP++ E++ + G RFI + +LP QL
Sbjct: 719 LQSPRNKKEVQRLTGRIAGLNRFIARSTDKSLPFYQL 755
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.321 0.138 0.414
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,947,234
Number of Sequences: 26719
Number of extensions: 797643
Number of successful extensions: 2552
Number of sequences better than 10.0: 127
Number of HSP's better than 10.0 without gapping: 81
Number of HSP's successfully gapped in prelim test: 47
Number of HSP's that attempted gapping in prelim test: 2148
Number of HSP's gapped (non-prelim): 296
length of query: 790
length of database: 11,318,596
effective HSP length: 107
effective length of query: 683
effective length of database: 8,459,663
effective search space: 5777949829
effective search space used: 5777949829
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC126008.1


