

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC123572.11 + phase: 0
(475 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
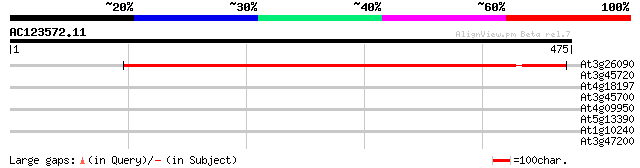
Score E
Sequences producing significant alignments: (bits) Value
At3g26090 unknown protein 509 e-144
At3g45720 putative transporter protein 34 0.15
At4g18197 unknown protein 33 0.45
At3g45700 putative transporter protein 32 0.76
At4g09950 AIG1-like protein 31 1.3
At5g13390 unknown protein 30 2.2
At1g10240 unknown protein 30 3.8
At3g47200 unknown protein 28 8.4
>At3g26090 unknown protein
Length = 393
Score = 509 bits (1310), Expect = e-144
Identities = 241/376 (64%), Positives = 307/376 (81%), Gaps = 5/376 (1%)
Query: 97 LFPVWGEGPLGFGLLLSSRITQAFQLYFIFVKRRLPLIRSFLLIPLILLPWIIGAAVIHI 156
L+ VW EGPLGFGLL+S RITQAFQLYFIFVK+RLP ++S++ +PL+LLPWI GAA+IH
Sbjct: 20 LWAVWIEGPLGFGLLMSCRITQAFQLYFIFVKKRLPPVKSYIFLPLVLLPWIFGAAIIHA 79
Query: 157 KKPLSNRCHMSVQWTIPVVCLHALYVATLVGVTAAVHHIEFRFDELRDLWRGILVSSVSV 216
KPL+++CHM +QWT PV LHALYV L+ T AV H+EFRFDELRDLW+GILVS+ S+
Sbjct: 80 TKPLNDKCHMGLQWTFPVAGLHALYVLALIAFTRAVRHVEFRFDELRDLWKGILVSATSI 139
Query: 217 AVWVTAYILNEIHDNISWLQVVSRFLLLVLASILVLAFFSISSSQPLLSQISLRRRESRE 276
+WVTA++LNEIH+ ISWLQV SRF+LLV ILV+ FFSISS+QPLLSQISL++R++ E
Sbjct: 140 VIWVTAFVLNEIHEEISWLQVASRFVLLVTGGILVVVFFSISSNQPLLSQISLKKRQNFE 199
Query: 277 FRTMGQALGIPDSGVLTQSEPISRVDPNEPLDKLLLNKKFRQSFMGFADSCLAGESVHFF 336
F+ MGQALGIPDSG+L + E VDPNEPLDKLLLNK+FR SFM FADSC AGE++HFF
Sbjct: 200 FQRMGQALGIPDSGLLFRKEEFRPVDPNEPLDKLLLNKRFRHSFMEFADSCYAGETLHFF 259
Query: 337 DEVHELSKISEHDCVRRIYMARHIIEKYMVAGAAMEINISHRSKQEILSTSDLARADLFH 396
+EV+E KI E D +RRIYMARHI+EK++VAGA ME+N+SH+++QEIL+T DL DLF
Sbjct: 260 EEVYEHGKIPEDDSIRRIYMARHIMEKFIVAGAEMELNLSHKTRQEILTTQDLTHTDLFK 319
Query: 397 NALNEIVHLMKTNLAKDYWSSMFFLKFQEECDMRCNGYELEQMTGWNY-SPRLSSVHGTD 455
NALNE++ L+K NL +DYWSS++F+KF+EE +E G+++ SPRLSSV G+D
Sbjct: 320 NALNEVMQLIKMNLVRDYWSSIYFIKFKEEESC----HEAMHKEGYSFSSPRLSSVQGSD 375
Query: 456 DPFHHDHLLKNSECGN 471
DPF+ +H+ K+S C +
Sbjct: 376 DPFYQEHMSKSSRCSS 391
>At3g45720 putative transporter protein
Length = 555
Score = 34.3 bits (77), Expect = 0.15
Identities = 31/125 (24%), Positives = 56/125 (44%), Gaps = 24/125 (19%)
Query: 151 AAVIHIK--KPLSNRCHMSVQWTIPVVCLHALYVATLVGVTAAVHH---IEFRFDELRDL 205
+A++ K K + N MSV W +P + +VG+ A H+ + + E +
Sbjct: 418 SAIVEAKRLKTVENEHPMSVLWLLPPL--------VIVGIGDAFHYMANVAVFYGEFPES 469
Query: 206 WRGILVSSVSVAVWVTAY----ILNEIHDNISWL-------QVVSRFLLLVLASILVLAF 254
R S SVA ++ Y ++N I +WL +V + + +LV+ +L L +
Sbjct: 470 MRNTATSVTSVAFGISFYLSTALINLIQRTTAWLPDDINHGRVDNVYWVLVIGGVLNLGY 529
Query: 255 FSISS 259
F + S
Sbjct: 530 FFVCS 534
>At4g18197 unknown protein
Length = 390
Score = 32.7 bits (73), Expect = 0.45
Identities = 58/285 (20%), Positives = 119/285 (41%), Gaps = 36/285 (12%)
Query: 9 KGGCPTDYVAVTVSILSFILLLIWSIFPFIVHKVPRTKGSGF-WIPVIQVVASFNLLLSI 67
+ G + YV + ++ F +L+++ F I + P++ + F P +AS L +
Sbjct: 68 ENGGNSTYVVTLLQLIGFPVLVLFRFFSRI--RQPKSTDTNFSQSPSFTTLASVYLCTGL 125
Query: 68 MTISSNLKRAIGGSHVTCGQVSFHNHTLCLFPVWGEGPLGFGLLLSSRIT-QAFQLYFIF 126
+ + A+G L PV F L+L+S++ AF YF+
Sbjct: 126 LVSAYAYLSAVG---------------LLYLPV-----STFSLILASQLAFTAFFSYFLN 165
Query: 127 VKRRLPLIRSFLLIPLILLPWIIGAAVIHIKKPLSNRCHMS-VQWTIPVVCLHALYVATL 185
++ PLI S LL+ + +A++ + N ++S VQ+ I +C + +
Sbjct: 166 SQKFTPLIVSSLLL------LTVSSALLVVNTDSENSTNVSRVQYVIGFIC--TIGASAG 217
Query: 186 VGVTAAVHHIEFRFDELRDLWRGILVSSVSVAVWVTAYILNEIHDNISWLQVVS---RFL 242
+G+ ++ + FR + + ++ ++ + +L + + W + S +
Sbjct: 218 IGLLLSLIQMLFRKVFTKHTSSAVTDLAIYQSLVASCVVLIGLFASGEWETLPSEMRNYK 277
Query: 243 LLVLASILVLAFFSISSSQPLLSQISLRRRESREFRTMGQALGIP 287
L ++ +L LA +IS L + L S F A+G+P
Sbjct: 278 LGKVSYVLTLASAAISWQVYTLGLVGLIFESSSVFSNSITAVGLP 322
>At3g45700 putative transporter protein
Length = 548
Score = 32.0 bits (71), Expect = 0.76
Identities = 25/118 (21%), Positives = 55/118 (46%), Gaps = 10/118 (8%)
Query: 151 AAVIHIK--KPLSNRCHMSVQWTIPVVCLHALYVATLVGVTAAVHHIEFRFDELRDLWRG 208
+AV+ K K + N MSV W +P + ++ + A A+ + EF + LR+
Sbjct: 410 SAVVEAKRLKTVENSHLMSVLWLVPALVINGIGEAFHFPANIAIFYGEFP-ESLRNTATS 468
Query: 209 ILVSSVSVAVWVTAYILNEIHDNISWL-------QVVSRFLLLVLASILVLAFFSISS 259
+ + ++ +++ +++ I WL +V + +L+LV+ + +F + S
Sbjct: 469 LTSVVMGISFYLSTALIDVIQRTTKWLPNDINHGRVDNVYLVLVIIGVSNFGYFLVCS 526
>At4g09950 AIG1-like protein
Length = 336
Score = 31.2 bits (69), Expect = 1.3
Identities = 15/68 (22%), Positives = 32/68 (47%)
Query: 396 HNALNEIVHLMKTNLAKDYWSSMFFLKFQEECDMRCNGYELEQMTGWNYSPRLSSVHGTD 455
H+ LN + + K N K Y + + + E ++ ++E+M GW+ +S +
Sbjct: 194 HDLLNLVEQISKKNNGKSYMADLSHELRENEATIKEKQKQIEEMKGWSSKQEISQMKKEL 253
Query: 456 DPFHHDHL 463
+ H++ L
Sbjct: 254 EKSHNEML 261
>At5g13390 unknown protein
Length = 1126
Score = 30.4 bits (67), Expect = 2.2
Identities = 32/122 (26%), Positives = 54/122 (44%), Gaps = 21/122 (17%)
Query: 132 PLIRSFLLIPLILLPWIIGAAVIHIKKPLSNRC------------HMSVQWTIPVVCLHA 179
P S+LLI +LL ++I IK + R ++S ++ + LHA
Sbjct: 728 PTWPSWLLIVSLLLILAAATSLIPIKYVVELRAFYSIAMGLALGVYISAEFFLQAAVLHA 787
Query: 180 LYVATLVGVTAAVHHIEFRFDELRDLWRGILVSSVSVAVWVTAYIL-------NEIHDNI 232
L V TL V A+V I F V ++ VA++ Y+L N++++N+
Sbjct: 788 LIVVTL--VCASVFVIFTHFPSASSTKLLPWVFALLVALFPVTYLLEGQVRIKNDLNENV 845
Query: 233 SW 234
+W
Sbjct: 846 TW 847
>At1g10240 unknown protein
Length = 680
Score = 29.6 bits (65), Expect = 3.8
Identities = 28/120 (23%), Positives = 50/120 (41%), Gaps = 18/120 (15%)
Query: 86 GQVSFHNHTLCLFPVWGEGPLGFGLLLSSRI----TQAFQLYFIFVKRRLPLIRSFLLIP 141
G++ H LC++ V G+ P F L R + ++LY L + F
Sbjct: 354 GEMPATKHALCIWMVVGKFPSWFNAGLGERYNDWKAEFYRLY------HLESVEEF---- 403
Query: 142 LILLPW--IIGAAVIHIKKPLSNRCHMSVQWTIPVVCLHALYVATLVGVTAAVHHIEFRF 199
L W ++ + +H + ++N W++P + H L TL G + A++ RF
Sbjct: 404 --ELGWRDMVNSFGLHTNRHINNLYASRSLWSLPYLRSHFLAGMTLTGRSKAINAFIQRF 461
>At3g47200 unknown protein
Length = 476
Score = 28.5 bits (62), Expect = 8.4
Identities = 15/59 (25%), Positives = 31/59 (52%), Gaps = 9/59 (15%)
Query: 141 PLILLPWIIGAAVIHIKKPLSNRCHMSVQWTIPVVCLHALYVATLVGVTAAVHHIEFRF 199
P+ +PW++ + I+ L + ++ +P L LYV + +GV++ ++ I F F
Sbjct: 161 PIFSIPWLLSS----IQSDL-----LLLENQVPFFVLQTLYVGSKIGVSSDLNRIAFHF 210
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.327 0.140 0.432
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,355,328
Number of Sequences: 26719
Number of extensions: 417039
Number of successful extensions: 1216
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1212
Number of HSP's gapped (non-prelim): 10
length of query: 475
length of database: 11,318,596
effective HSP length: 103
effective length of query: 372
effective length of database: 8,566,539
effective search space: 3186752508
effective search space used: 3186752508
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC123572.11


