

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122164.8 + phase: 0 /pseudo
(640 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
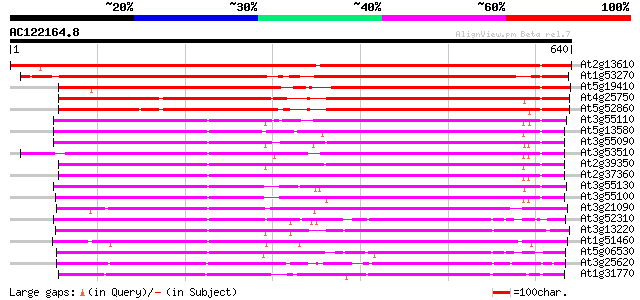
Score E
Sequences producing significant alignments: (bits) Value
At2g13610 putative ABC transporter 887 0.0
At1g53270 putative ABC transporter emb|AAD22683.1 523 e-148
At5g19410 membrane transporter - like protein 469 e-132
At4g25750 putative membrane transporter 459 e-129
At5g52860 ABC transporter-like protein 456 e-128
At3g55110 ABC transporter - like protein 381 e-106
At5g13580 ABC transporter-like protein 375 e-104
At3g55090 ABC transporter - like protein 374 e-103
At3g53510 ABC transporter -like protein 373 e-103
At2g39350 ABC transporter like protein 371 e-103
At2g37360 ABC transporter like protein 370 e-102
At3g55130 ABC transporter - like protein 363 e-100
At3g55100 ABC transporter - like protein 358 4e-99
At3g21090 ABC transporter, putative 236 2e-62
At3g52310 ABC transporter-like protein 235 5e-62
At3g13220 ABC transporter 234 1e-61
At1g51460 ATP-dependent transmembrane transporter like protein 233 2e-61
At5g06530 ABC transporter like protein 233 3e-61
At3g25620 membrane transporter, putative 231 9e-61
At1g31770 unknown protein 226 3e-59
>At2g13610 putative ABC transporter
Length = 649
Score = 887 bits (2291), Expect = 0.0
Identities = 460/646 (71%), Positives = 531/646 (81%), Gaps = 10/646 (1%)
Query: 1 MKMQGCEVEVIGINYKIHTNKAE-HPFKIFSKSP-----QLVNTNVQETEEVEKGCSGVR 54
M+ QGCE+E + I+Y I K +PF IF + P Q V T + + ++ + V+
Sbjct: 1 MEKQGCEIEALDIDYNIFVRKINVNPFGIFRRKPRPEADQPVKTEEESLKLEDETGNKVK 60
Query: 55 HVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQKGSVLLNQKPVDKSQFRKLS 114
HVLK V+ +A+PWEILAIVGPSGAGKSSLLEILA + PQ GSV +N++PVD++ F+K+S
Sbjct: 61 HVLKGVTCRAKPWEILAIVGPSGAGKSSLLEILAARLIPQTGSVYVNKRPVDRANFKKIS 120
Query: 115 GYVTQKDTLFPLLTVEETMMFSAKLRLKLPQQQQCSRVKSLIKELGLDHVAGTRIGDDRV 174
GYVTQKDTLFPLLTVEET++FSAKLRLKLP + SRVKSL+ ELGL+ VA R+GDD V
Sbjct: 121 GYVTQKDTLFPLLTVEETLLFSAKLRLKLPADELRSRVKSLVHELGLEAVATARVGDDSV 180
Query: 175 RGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIILSI 234
RGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSAL IIDMLK MAETRGRTIIL+I
Sbjct: 181 RGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALLIIDMLKHMAETRGRTIILTI 240
Query: 235 HQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSIDVIQ 294
HQPGFRIVK FNS+LLLANGS L G+ D L V LR GL PLH N+VEFAI+SI+ I
Sbjct: 241 HQPGFRIVKQFNSVLLLANGSTLKQGSVDQLGVYLRSNGLHPPLHENIVEFAIESIESIT 300
Query: 295 QQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDEDIINKTGT 354
+QQ+ Q L T ++ R ++ QG+ KSGKFTLQQLFQQ++V D +N
Sbjct: 301 KQQRLQESRRAAHVLTPQTTLQEKRSEDSQGESKSGKFTLQQLFQQTRVADVGTMN---I 357
Query: 355 GMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFCNLKDDLR 414
+F+ DFANSRL ETMILTHRFSKNIFRTKELFACRT+QML SG+VLG IF NLKDDL+
Sbjct: 358 ATEFTRDFANSRLEETMILTHRFSKNIFRTKELFACRTVQMLGSGIVLGLIFHNLKDDLK 417
Query: 415 GTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVYLPFL 474
G +ERVGLFAFILTFLL+S+IEALPIFLQEREILMKETS GSYRVSSYA+ANGLVYLPFL
Sbjct: 418 GARERVGLFAFILTFLLTSTIEALPIFLQEREILMKETSSGSYRVSSYAVANGLVYLPFL 477
Query: 475 LILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIVGNSVIN 534
LILAILF+ P+YWLVGLN +F AFLHF LLIWL+LYTANSVVVCFSALVPNFIVGNSVI+
Sbjct: 478 LILAILFSTPVYWLVGLNPSFMAFLHFSLLIWLILYTANSVVVCFSALVPNFIVGNSVIS 537
Query: 535 GVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFSNSKKCLEYMFGACVM 594
GV+GSFFLFSGYFISNHEIP YWIFMHYISLFKYPFEGFLINEFS S KCLEY FG C++
Sbjct: 538 GVMGSFFLFSGYFISNHEIPGYWIFMHYISLFKYPFEGFLINEFSKSNKCLEYGFGKCLV 597
Query: 595 KGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYVILRYKCSER 640
ED+LKEE Y GE SRW+NV + +CF+++YRFISYVILR +CS+R
Sbjct: 598 TEEDLLKEERY-GEESRWRNVVIMLCFVLLYRFISYVILRCRCSQR 642
>At1g53270 putative ABC transporter emb|AAD22683.1
Length = 590
Score = 523 bits (1346), Expect = e-148
Identities = 292/628 (46%), Positives = 402/628 (63%), Gaps = 60/628 (9%)
Query: 13 INYKIHTNKAEHPFKIFSKSPQLVNTNVQETEEVEKGCSGVRHVLKNVSFQARPWEILAI 72
I+Y++ T ++I +P+ N +E+ EK +LK+VS AR EI AI
Sbjct: 15 ISYRLETKNLS--YRIGGNTPKFSNLCGLLSEKEEKV------ILKDVSCDARSAEITAI 66
Query: 73 VGPSGAGKSSLLEILAGK--HRPQKGSVLLNQKPVDKSQFRKLSGYVTQKDTLFPLLTVE 130
GPSGAGK++LLEILAGK H G VL+N +P+D ++R++SG+V Q+D LFP LTV+
Sbjct: 67 AGPSGAGKTTLLEILAGKVSHGKVSGQVLVNGRPMDGPEYRRVSGFVPQEDALFPFLTVQ 126
Query: 131 ETMMFSAKLRLKLPQQQQCSRVKSLIKELGLDHVAGTRIGDDRVRGISGGERRRVSIGVE 190
ET+ +SA LRLK ++ ++VK LI+ELGL+HVA +RIG GISGGERRRVSIGVE
Sbjct: 127 ETLTYSALLRLKTKRKDAAAKVKRLIQELGLEHVADSRIGQGSRSGISGGERRRVSIGVE 186
Query: 191 VIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIILSIHQPGFRIVKLFNSLLL 250
++HDP V+++DEPTSGLDS SALQ++ +LK M +G+TI+L+IHQPGFRI++ + ++L
Sbjct: 187 LVHDPNVILIDEPTSGLDSASALQVVTLLKDMTIKQGKTIVLTIHQPGFRILEQIDRIVL 246
Query: 251 LANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSIDVIQQQQQWQVETETPRRLQ 310
L+NG V+ +G+ L ++ G ++P VNV+E+AID ++ P R Q
Sbjct: 247 LSNGMVVQNGSVYSLHQKIKFSGHQIPRRVNVLEYAIDIAGSLE-----------PIRTQ 295
Query: 311 GTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDEDIINKTGTGMDFSYDFANSRLRET 370
R+ G K+ K G + S +NS L E
Sbjct: 296 SC------REISCYGHSKTWK---------------SCYISAGGELHQSDSHSNSVLEEV 334
Query: 371 MILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFCNLKDDLRGTQE-RVGLFAFILTF 429
IL R KNIFRTK+LF R +Q I+GL+LGSI+ N+ + + + R G FAFILTF
Sbjct: 335 QILGQRSCKNIFRTKQLFTTRALQASIAGLILGSIYLNVGNQKKEAKVLRTGFFAFILTF 394
Query: 430 LLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLV 489
LLSS+ E LPIFLQ+R ILM+ETS +YRV SY +A+ L+++PFLLI+++LF P+YWLV
Sbjct: 395 LLSSTTEGLPIFLQDRRILMRETSRRAYRVLSYVLADTLIFIPFLLIISMLFATPVYWLV 454
Query: 490 GLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFIS 549
GL FL+F L+IW+VL +NS V CFSALVPNFI+G SVI+G++GSFFLFSGYFI+
Sbjct: 455 GLRRELDGFLYFSLVIWIVLLMSNSFVACFSALVPNFIMGTSVISGLMGSFFLFSGYFIA 514
Query: 550 NHEIPSYWIFMHYISLFKYPFEGFLINEFSNSKKCLEYMFGACVMKGEDVLKEEGYGGEG 609
IP YW FMHY+SLFKYPFE +INE+ +G+ LK++ E
Sbjct: 515 KDRIPVYWEFMHYLSLFKYPFECLMINEY----------------RGDVFLKQQDL-KES 557
Query: 610 SRWKNVGVTVCFIMVYRFISYVILRYKC 637
+W N+G+ FI+ YR + + IL Y+C
Sbjct: 558 QKWSNLGIMASFIVGYRVLGFFILWYRC 585
>At5g19410 membrane transporter - like protein
Length = 624
Score = 469 bits (1207), Expect = e-132
Identities = 251/593 (42%), Positives = 389/593 (65%), Gaps = 46/593 (7%)
Query: 56 VLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK--HR---PQKGSVLLNQKPVDKSQF 110
+L +VS A +ILA+VGPSG GKS+LL+I++G+ H+ P ++ N+K D +Q
Sbjct: 66 ILNSVSLAAESSKILAVVGPSGTGKSTLLKIISGRVNHKALDPSSAVLMNNRKITDYNQL 125
Query: 111 RKLSGYVTQKDTLFPLLTVEETMMFSAKLRLK-LPQQQQCSRVKSLIKELGLDHVAGTRI 169
R+L G+V Q D L PLLTV+ET+M+SAK L+ +++ RV+SL+ +LGL V + +
Sbjct: 126 RRLCGFVPQDDDLLPLLTVKETLMYSAKFSLRDSTAKEREERVESLLSDLGLVLVQDSFV 185
Query: 170 G--DDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRG 227
G D+ RG+SGGER+RVSI VE+I DP +L+LDEPTSGLDS ++LQ++++L MA+++
Sbjct: 186 GEGDEEDRGVSGGERKRVSIAVEMIRDPPILLLDEPTSGLDSRNSLQVVELLATMAKSKQ 245
Query: 228 RTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAI 287
RT++ SIHQP +RI+ + L+L+ GSV+H G+ + L ++ +G ++P +N +EFA+
Sbjct: 246 RTVLFSIHQPSYRILDYISDYLILSRGSVIHLGSLEHLEDSIAKLGFQIPEQLNPIEFAM 305
Query: 288 DSIDVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDED 347
+ ++ ++ + V + + ++ G + ++ F +V+D
Sbjct: 306 EIVESLRTFKPNSVAVVESSSMW------------PENNENDGIISKKEAF---RVLD-- 348
Query: 348 IINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFC 407
+ E L RF K I+RTK+LF RT+Q +++GL LGS++
Sbjct: 349 -------------------VTEISYLCSRFCKIIYRTKQLFLARTMQAVVAGLGLGSVYT 389
Query: 408 NLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANG 467
LK D G ER+GLFAF L+FLLSS++EALPI+L+ER +LMKE+S GSYR+SSY IAN
Sbjct: 390 RLKRDEEGVAERLGLFAFSLSFLLSSTVEALPIYLRERRVLMKESSRGSYRISSYMIANT 449
Query: 468 LVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFI 527
+ ++PFL ++++LF++P+YW+VGLN + AF F+L +WL++ A+S+V+ SA+ P+FI
Sbjct: 450 IAFVPFLFVVSLLFSIPVYWIVGLNPSIQAFSFFVLCVWLIILMASSLVLFLSAVSPDFI 509
Query: 528 VGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEF-SNSKKCLE 586
GNS+I V+G+FFLFSGYFI +IP W+FM+Y+SL++YP E ++NE+ S ++C
Sbjct: 510 SGNSLICTVLGAFFLFSGYFIPKEKIPKPWMFMYYVSLYRYPLESMVVNEYWSMREECFS 569
Query: 587 YMFGACVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYVILRYKCSE 639
C+M GEDVLKE G + +RW NVG+ + F + YR + + IL K S+
Sbjct: 570 SGNMGCLMTGEDVLKERGL-DKDTRWINVGIMLAFFVFYRILCWGILLRKASK 621
>At4g25750 putative membrane transporter
Length = 577
Score = 459 bits (1182), Expect = e-129
Identities = 251/588 (42%), Positives = 373/588 (62%), Gaps = 49/588 (8%)
Query: 56 VLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQKGSVLLNQKPVDKSQFRKLSG 115
+L+N++ + P +ILAI+GPSGAGKS+LL+ILA + P GS+LLN ++ S +RK+S
Sbjct: 30 ILRNITLTSHPSQILAIIGPSGAGKSTLLDILAARTSPTSGSILLNSVLINPSSYRKISS 89
Query: 116 YVTQKDTLFPLLTVEETMMFSAKLRLKLPQQQQCSRVKSLIKELGLDHVAGTRIGDDRVR 175
YV Q DT FPLLTV ET FSA L L + S V SL+KEL L H+A TR+G +
Sbjct: 90 YVPQHDTFFPLLTVSETFTFSASLLLPKNLSKVSSVVASLLKELNLTHLAHTRLG----Q 145
Query: 176 GISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIILSIH 235
G+SGGERRRVSIG+ ++HDP+VL+LDEPTSGLDS SA ++ +LK +A +R R +ILSIH
Sbjct: 146 GLSGGERRRVSIGLSLLHDPEVLLLDEPTSGLDSKSAFDVVQILKSIATSRERIVILSIH 205
Query: 236 QPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSIDVIQQ 295
QP F+I+ L + +LLL+ G++++HG DLL L G +P +N +E+A++ + I+
Sbjct: 206 QPSFKILSLIDRVLLLSKGTIVYHGRLDLLEAFLLSKGFTVPSQLNSLEYAMEILQNIRD 265
Query: 296 QQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDEDIINKTGTG 355
+ P + +Q + +QS V
Sbjct: 266 PYE-NANIALPDHCPESKKQNQ---------------------KQSIV------------ 291
Query: 356 MDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFCNLKDDLRG 415
+ +SR+ E +L+ RF K I+RT++L ++ L+ GLVLG+I+ N+ G
Sbjct: 292 -----RYKSSRITEISLLSSRFWKIIYRTRQLLLTNILESLVVGLVLGTIYLNIGTGKEG 346
Query: 416 TQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVYLPFLL 475
++R GLFAF LTFLLSS+ + LPIF+ ER IL++ETS G YR+SS+ +AN LV+LP+LL
Sbjct: 347 IRKRFGLFAFTLTFLLSSTTQTLPIFIDERPILLRETSSGLYRLSSHILANTLVFLPYLL 406
Query: 476 ILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIVGNSVING 535
++AI+++V LY+LVGL ++ A +F+L+IW+++ ANS V+ S+L PN+I G S +
Sbjct: 407 LIAIIYSVSLYFLVGLCFSWQALAYFVLVIWIIVLMANSFVLFLSSLAPNYIAGTSSVTI 466
Query: 536 VIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFS-NSKKCL----EYMFG 590
++ +FFLFSGYFIS +P YW+FM++ S++KY + LINE+S KCL E
Sbjct: 467 LLAAFFLFSGYFISKESLPKYWLFMYFFSMYKYALDALLINEYSCLHNKCLVWFEEASVN 526
Query: 591 ACVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYVILRYKCS 638
+C++ G DVL + G E RW NV + + F ++YR + +++L + S
Sbjct: 527 SCLVTGGDVLDKNGL-HERQRWFNVYMLLGFFVLYRVLCFLVLLKRVS 573
>At5g52860 ABC transporter-like protein
Length = 589
Score = 456 bits (1173), Expect = e-128
Identities = 252/588 (42%), Positives = 373/588 (62%), Gaps = 51/588 (8%)
Query: 56 VLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQKGSVLLNQKPVDKSQFRKLSG 115
+L+N++ A P EILA+VGPSGAGKS+LL+ILA K P GS+LLN P++ S +RK+S
Sbjct: 44 ILRNITLTAHPTEILAVVGPSGAGKSTLLDILASKTSPTSGSILLNSIPINPSSYRKISS 103
Query: 116 YVTQKDTLFPLLTVEETMMFSAKLRLKLPQQQQCSRVKSLIKELGLDHVAGTRIGDDRVR 175
YV Q D+ FPLLTV ET F+A L L P V SL+ EL L H++ TR+ +
Sbjct: 104 YVPQHDSFFPLLTVSETFSFAACLLLPNPSIVS-ETVTSLLSELNLTHLSHTRLA----Q 158
Query: 176 GISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIILSIH 235
G+SGGERRRVSIG+ ++HDP L+LDEPTSGLDS SA +I +LK +A +R RT+ILSIH
Sbjct: 159 GLSGGERRRVSIGLSLLHDPCFLLLDEPTSGLDSKSAFDVIHILKSIAVSRQRTVILSIH 218
Query: 236 QPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSIDVIQQ 295
QP F+I+ + + LLLL+ G+V++HG D L L G +P +N +E+A++ + +++
Sbjct: 219 QPSFKILSIIDRLLLLSKGTVVYHGRLDSLEGFLLFKGFTVPPQLNSLEYAMEILQELRE 278
Query: 296 QQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDEDIINKTGTG 355
T P + + +K R+ +QS V
Sbjct: 279 SDGNTDATALP-----SIENRKQRE------------------KQSIV------------ 303
Query: 356 MDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFCNLKDDLRG 415
+ SR+ E +L RF K I+RT++L ++ L+ GLVLG+I+ N+ G
Sbjct: 304 -----RYRKSRITEISLLARRFWKIIYRTRQLLLTNALEALVVGLVLGTIYINIGIGKAG 358
Query: 416 TQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVYLPFLL 475
++R G+FAF LTFLLSS+ E LPIF+ ER IL++ETS G YR+SS+ +AN LV+LP+L
Sbjct: 359 IEKRFGMFAFTLTFLLSSTTETLPIFINERPILLRETSSGIYRLSSHILANTLVFLPYLF 418
Query: 476 ILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIVGNSVING 535
+++I+++V +Y+L+GL + AF +F+L+IW++L ANS V+ S+L PN+I G S++
Sbjct: 419 VISIIYSVSVYFLIGLCPTWQAFGYFVLVIWIILLMANSFVLFLSSLAPNYITGTSLVTI 478
Query: 536 VIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFS-NSKKCLEYMFGA--- 591
++ +FFLFSGYFIS +P YW+FM++ S++KY + LINE+S + KCL ++ A
Sbjct: 479 LLAAFFLFSGYFISKESLPKYWLFMYFFSMYKYALDALLINEYSCLASKCLVWLEEAQTK 538
Query: 592 -CVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYVILRYKCS 638
C++ G DVLK++G E RW NV V + F ++YR + ++ L + S
Sbjct: 539 ICMVTGGDVLKKKGL-HEKQRWFNVYVLLGFFVLYRVLCFLALLRRVS 585
>At3g55110 ABC transporter - like protein
Length = 708
Score = 381 bits (979), Expect = e-106
Identities = 226/634 (35%), Positives = 360/634 (56%), Gaps = 67/634 (10%)
Query: 51 SGVRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQ--KGSVLLNQKPVDKS 108
+ V+ +L +++ +AR EILA++G SGAGKS+L++ LAG+ KG+V LN + V +S
Sbjct: 86 ASVKTLLDDITGEARDGEILAVLGGSGAGKSTLIDALAGRVAEDSLKGTVTLNGEKVLQS 145
Query: 109 QFRK-LSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVA 165
+ K +S YV Q D LFP+LTV+ET+MF+++ RL LP+ ++ RV++LI +LGL + A
Sbjct: 146 RLLKVISAYVMQDDLLFPMLTVKETLMFASEFRLPRSLPKSKKMERVETLIDQLGLRNAA 205
Query: 166 GTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAET 225
T IGD+ RG+SGGERRRVSIG+++IHDP +L LDEPTSGLDST+A ++ +LK +A++
Sbjct: 206 DTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTNAFMVVQVLKRIAQS 265
Query: 226 RGRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEF 285
G +I+SIHQP RI+ L + L++L++G + +G+ L G +P N+ EF
Sbjct: 266 -GSVVIMSIHQPSARIIGLLDRLIILSHGKSVFNGSPVSLPSFFSSFGRPIPEKENITEF 324
Query: 286 AIDSI-----------DVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTL 334
A+D I D+++ ++WQ + +T R TTQ + + GK
Sbjct: 325 ALDVIRELEGSSEGTRDLVEFNEKWQ-QNQTAR---ATTQSRVSLKEAIAASVSRGKLV- 379
Query: 335 QQLFQQSKVIDEDIINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQ 394
M+ +AN L ET IL R+ KN RT EL R
Sbjct: 380 -----------SGSSGANPISMETVSSYANPPLAETFILAKRYIKNWIRTPELIGMRIGT 428
Query: 395 MLISGLVLGSIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSC 454
++++GL+L +++ L + RG QER+G FAF ++ + + +P+F+QER I ++ET+
Sbjct: 429 VMVTGLLLATVYWRLDNTPRGAQERMGFFAFGMSTMFYCCADNIPVFIQERYIFLRETTH 488
Query: 455 GSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANS 514
+YR SSY I++ LV LP LL L+I F +W VGL+ +F ++ L+I+ ++ +S
Sbjct: 489 NAYRTSSYVISHALVSLPQLLALSIAFAATTFWTVGLSGGLESFFYYCLIIYAAFWSGSS 548
Query: 515 VVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFL 574
+V S L+PN ++ V + L G++I+ IP YWI+ HYISL KYP+E L
Sbjct: 549 IVTFISGLIPNVMMSYMVTIAYLSYCLLLGGFYINRDRIPLYWIWFHYISLLKYPYEAVL 608
Query: 575 INEFSNSKKC------------------------LEYMFGA---------CVMKGEDVLK 601
INEF + +C L+ + G+ C+ G D+L
Sbjct: 609 INEFDDPSRCFVKGVQVFDGTLLAEVSHVMKVKLLDTLSGSLGTKITESTCLRTGPDLLM 668
Query: 602 EEGYGGEGSRWKNVGVTVCFIMVYRFISYVILRY 635
++G + S+W + +T+ + + +R + Y+ L +
Sbjct: 669 QQGI-TQLSKWDCLWITLAWGLFFRILFYLSLLF 701
>At5g13580 ABC transporter-like protein
Length = 727
Score = 375 bits (964), Expect = e-104
Identities = 225/623 (36%), Positives = 350/623 (56%), Gaps = 47/623 (7%)
Query: 51 SGVRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK--HRPQKGSVLLNQKPVDKS 108
S + +L ++ +AR EILA++G SG+GKS+L++ LA + KG+V LN + ++
Sbjct: 103 SKTKTLLNGITGEARDGEILAVLGASGSGKSTLIDALANRIAKGSLKGNVTLNGEVLNSK 162
Query: 109 QFRKLSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAG 166
+ +S YV Q D LFP+LTVEET+MF+A+ RL L + ++ RV++LI +LGL + A
Sbjct: 163 MQKAISAYVMQDDLLFPMLTVEETLMFAAEFRLPRSLSKSKKSLRVQALIDQLGLRNAAN 222
Query: 167 TRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETR 226
T IGD+ RGISGGERRRVSIG+++IHDP +L LDEPTSGLDSTSAL +I +LK +A++
Sbjct: 223 TVIGDEGHRGISGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSALSVIKVLKRIAQS- 281
Query: 227 GRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFA 286
G +I+++HQP +R+++L + LL L+ G + G+ +L G +P H N EFA
Sbjct: 282 GSMVIMTLHQPSYRLLRLLDRLLFLSRGQTVFSGSPAMLPRFFAEFGHPIPEHENRTEFA 341
Query: 287 IDSIDVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDE 346
+D +I++ + T + +Q+K Q G S K + + K++
Sbjct: 342 LD---LIRELEGSAGGTRSLVEFNKGFRQRKAEPRSQTG--LSLKEAISASISKGKLVSG 396
Query: 347 DIINKTGTG---MDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLG 403
+G + FAN E +L R N R ELF R +L++G +L
Sbjct: 397 ATTTTHSSGSSPVSTIPTFANPFWVELAVLAKRSMTNSRRQPELFGIRLGAVLVTGFILA 456
Query: 404 SIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYA 463
++F L + +G QER+G FAF ++ + +ALP+FLQER I M+ET+ +YR SSY
Sbjct: 457 TMFWQLDNSPKGVQERLGCFAFAMSTTFYTCADALPVFLQERFIFMRETAYNAYRRSSYV 516
Query: 464 IANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALV 523
+++ LV LP L+IL++ F +W VGL+ FL + L+I + +S V S +V
Sbjct: 517 LSHSLVALPSLIILSLAFAAITFWGVGLDGGLMGFLFYFLVILASFWAGSSFVTFLSGVV 576
Query: 524 PNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFSNSKK 583
P+ ++G +++ ++ F LFSG+FI+ IP YWI+ HYISL KYP+E L+NEF + K
Sbjct: 577 PHVMLGYTIVVAILAYFLLFSGFFINRDRIPGYWIWFHYISLVKYPYEAVLLNEFGDPTK 636
Query: 584 C---------------------------------LEYMFGACVMKGEDVLKEEGYGGEGS 610
C + C+ G D+L+++G + +
Sbjct: 637 CFVRGVQIFDNTPLVAVPQGMKVRLLATMSKSLGMRITSSTCLTTGYDILQQQGV-TDLT 695
Query: 611 RWKNVGVTVCFIMVYRFISYVIL 633
+W + VTV + +R + Y L
Sbjct: 696 KWNCLWVTVAWGFFFRILFYFSL 718
>At3g55090 ABC transporter - like protein
Length = 720
Score = 374 bits (959), Expect = e-103
Identities = 226/636 (35%), Positives = 351/636 (54%), Gaps = 63/636 (9%)
Query: 51 SGVRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK--HRPQKGSVLLNQKPVDKS 108
S + +L N+S + R EILA++G SG+GKS+L++ LA + KG+V LN + +
Sbjct: 86 SKTKTLLDNISGETRDGEILAVLGASGSGKSTLIDALANRIAKGSLKGTVTLNGEALQSR 145
Query: 109 QFRKLSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAG 166
+ +S YV Q D LFP+LTVEET+MF+A+ RL LP+ ++ RV++LI +LG+ + A
Sbjct: 146 MLKVISAYVMQDDLLFPMLTVEETLMFAAEFRLPRSLPKSKKKLRVQALIDQLGIRNAAK 205
Query: 167 TRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETR 226
T IGD+ RGISGGERRRVSIG+++IHDP VL LDEPTSGLDSTSA ++ +LK +AE+
Sbjct: 206 TIIGDEGHRGISGGERRRVSIGIDIIHDPIVLFLDEPTSGLDSTSAFMVVKVLKRIAES- 264
Query: 227 GRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFA 286
G II+SIHQP R++ L + L+ L+ G + G+ L G +P + N EFA
Sbjct: 265 GSIIIMSIHQPSHRVLSLLDRLIFLSRGHTVFSGSPASLPSFFAGFGNPIPENENQTEFA 324
Query: 287 IDSI-----------DVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQ 335
+D I +++ ++WQ ++ + + + + K +
Sbjct: 325 LDLIRELEGSAGGTRGLVEFNKKWQ-------EMKKQSNPQTLTPPASPNPNLTLKEAIS 377
Query: 336 QLFQQSKVID-----EDIINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFAC 390
+ K++ +IN G + FAN E LT R N R EL
Sbjct: 378 ASISRGKLVSGGGGGSSVINHGGGTLAVPA-FANPFWIEIKTLTRRSILNSRRQPELLGM 436
Query: 391 RTIQMLISGLVLGSIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMK 450
R ++++G +L ++F L + +G QER+G FAF ++ + + +ALP+FLQER I M+
Sbjct: 437 RLATVIVTGFILATVFWRLDNSPKGVQERLGFFAFAMSTMFYTCADALPVFLQERYIFMR 496
Query: 451 ETSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLY 510
ET+ +YR SSY +++ +V P L+ L++ F V +W VGL FL + L+I +
Sbjct: 497 ETAYNAYRRSSYVLSHAIVTFPSLIFLSLAFAVTTFWAVGLEGGLMGFLFYCLIILASFW 556
Query: 511 TANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPF 570
+ +S V S +VP+ ++G +++ ++ F LFSG+FI+ IP YWI+ HY+SL KYP+
Sbjct: 557 SGSSFVTFLSGVVPHVMLGYTIVVAILAYFLLFSGFFINRDRIPQYWIWFHYLSLVKYPY 616
Query: 571 EGFLINEFSNSKKCL-------------EYMFG--------------------ACVMKGE 597
E L NEFS+ +C E +G C+ G
Sbjct: 617 EAVLQNEFSDPTECFVRGVQLFDNSPLGELTYGMKLRLLDSVSRSIGMRISSSTCLTTGA 676
Query: 598 DVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYVIL 633
DVLK++G + S+W + +TV F ++R + Y+ L
Sbjct: 677 DVLKQQGV-TQLSKWNCLLITVGFGFLFRILFYLCL 711
>At3g53510 ABC transporter -like protein
Length = 739
Score = 373 bits (957), Expect = e-103
Identities = 235/662 (35%), Positives = 359/662 (53%), Gaps = 65/662 (9%)
Query: 13 INYKIHTNKAEHPFKIFSKSPQLVNTNVQETEEVEKGCSGVRHVLKNVSFQARPWEILAI 72
+ Y + K PF SP N T+ + G SG +AR E++A+
Sbjct: 93 LTYSVKIKKKFKPFPCCGNSPFDGNDMEMNTKVLLNGISG----------EAREGEMMAV 142
Query: 73 VGPSGAGKSSLLEILAGKHRPQ--KGSVLLNQKPVDKSQFRKLSGYVTQKDTLFPLLTVE 130
+G SG+GKS+L++ LA + + +G + LN + ++ S + +S YV Q D LFP+LTVE
Sbjct: 143 LGASGSGKSTLIDALANRISKESLRGDITLNGEVLESSLHKVISAYVMQDDLLFPMLTVE 202
Query: 131 ETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTRIGDDRVRGISGGERRRVSIG 188
ET+MFSA+ RL L ++++ +RV++LI +LGL + A T IGD+ RG+SGGERRRVSIG
Sbjct: 203 ETLMFSAEFRLPSSLSKKKKKARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIG 262
Query: 189 VEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIILSIHQPGFRIVKLFNSL 248
++IHDP +L LDEPTSGLDSTSA ++ +L+ +A++ G +I+SIHQP +RI+ L + L
Sbjct: 263 TDIIHDPIILFLDEPTSGLDSTSAYMVVKVLQRIAQS-GSIVIMSIHQPSYRILGLLDKL 321
Query: 249 LLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSIDVIQQQQQWQ---VETET 305
+ L+ G+ ++ G+ L G +P + N EFA+D I ++ + VE
Sbjct: 322 IFLSRGNTVYSGSPTHLPQFFSEFGHPIPENENKPEFALDLIRELEDSPEGTKSLVEFHK 381
Query: 306 PRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDEDIINKTGTGMDFSYD-FAN 364
R + T+ Q + + D S + +L + T + S+ FAN
Sbjct: 382 QWRAKQTSSQSRRNTNVSLKDAISASISRGKLVSGA------------TNLRSSFQTFAN 429
Query: 365 SRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFCNLKDDLRGTQERVGLFA 424
E +++ R N R ELF R +L++G++L +IF L + RG QER+G FA
Sbjct: 430 PFWTEMLVIGKRSILNSRRQPELFGIRLGAVLVTGMILATIFWKLDNSPRGIQERLGFFA 489
Query: 425 FILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVYLPFLLILAILFTVP 484
F ++ + EA+P+FLQER I M+ET+ +YR SSY +A+ ++ +P L+IL+ F
Sbjct: 490 FAMSTTFYTCAEAIPVFLQERYIFMRETAYNAYRRSSYVLAHTIISIPALIILSAAFAAS 549
Query: 485 LYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIVGNSVINGVIGSFFLFS 544
+ VGL FL F I + +S V S +V + ++G +V+ ++ F LFS
Sbjct: 550 TFSAVGLAGGSEGFLFFFFTILTAFWAGSSFVTFLSGVVSHVMIGFTVVVAILAYFLLFS 609
Query: 545 GYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFSNSKKC-------------------- 584
G+FIS IP YWI+ HY+SL KYP+EG L NEF + KC
Sbjct: 610 GFFISRDRIPLYWIWFHYLSLVKYPYEGVLQNEFEDPTKCFVRGIQMFDNSPLGQVPTAV 669
Query: 585 ----LEYMFG---------ACVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYV 631
L+ M G CV G D+LK++G E S+W + +TV + +R + Y
Sbjct: 670 KISLLKSMSGVLGINVTAETCVTTGIDILKQQGI-TEISKWNCLWITVAWGFFFRVLFYF 728
Query: 632 IL 633
L
Sbjct: 729 TL 730
>At2g39350 ABC transporter like protein
Length = 740
Score = 371 bits (953), Expect = e-103
Identities = 219/626 (34%), Positives = 345/626 (54%), Gaps = 51/626 (8%)
Query: 56 VLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK--HRPQKGSVLLNQKPVDKSQFRKL 113
+L N+S + R EI+A++G SG+GKS+L++ LA + KG+V LN + + + +
Sbjct: 109 LLNNISGETRDGEIMAVLGASGSGKSTLIDALANRIAKGSLKGTVKLNGETLQSRMLKVI 168
Query: 114 SGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTRIGD 171
S YV Q D LFP+LTVEET+MF+A+ RL LP+ ++ RV++LI +LG+ + A T IGD
Sbjct: 169 SAYVMQDDLLFPMLTVEETLMFAAEFRLPRSLPKSKKKLRVQALIDQLGIRNAAKTIIGD 228
Query: 172 DRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTII 231
+ RGISGGERRRVSIG+++IHDP +L LDEPTSGLDSTSA ++ +LK +A++ G +I
Sbjct: 229 EGHRGISGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLKRIAQS-GSIVI 287
Query: 232 LSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSI- 290
+SIHQP R++ L + L+ L+ G ++ G+ L G +P + N EFA+D I
Sbjct: 288 MSIHQPSHRVLGLLDRLIFLSRGHTVYSGSPASLPRFFTEFGSPIPENENRTEFALDLIR 347
Query: 291 ----------DVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQ 340
+I+ ++WQ + R T + + + + +L
Sbjct: 348 ELEGSAGGTRGLIEFNKKWQEMKKQSNRQPPLTPPSSPYPNLTLKEAIAASISRGKLVSG 407
Query: 341 SKVIDEDIINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGL 400
+ + T + FAN E L+ R N R ELF R ++I+G
Sbjct: 408 GESVAHGGATTNTTTLAVP-AFANPMWIEIKTLSKRSMLNSRRQPELFGIRIASVVITGF 466
Query: 401 VLGSIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVS 460
+L ++F L + +G QER+G FAF ++ + + +ALP+FLQER I M+ET+ +YR S
Sbjct: 467 ILATVFWRLDNSPKGVQERLGFFAFAMSTMFYTCADALPVFLQERYIFMRETAYNAYRRS 526
Query: 461 SYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFS 520
SY +++ +V P L+ L++ F YW VGL+ T L + L+I ++ +S V S
Sbjct: 527 SYVLSHAIVSFPSLIFLSVAFAATTYWAVGLDGGLTGLLFYCLIILASFWSGSSFVTFLS 586
Query: 521 ALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFSN 580
+VP+ ++G +++ ++ F LFSG+FI+ + IP YWI+ HY+SL KYP+E L NEFS+
Sbjct: 587 GVVPSVMLGYTIVVAILAYFLLFSGFFINRNRIPDYWIWFHYMSLVKYPYEAVLQNEFSD 646
Query: 581 SKKC---------------------------------LEYMFGACVMKGEDVLKEEGYGG 607
+ KC + C+ G D+L+++G
Sbjct: 647 ATKCFVRGVQIFDNTPLGELPEVMKLKLLGTVSKSLGVTISSTTCLTTGSDILRQQGV-V 705
Query: 608 EGSRWKNVGVTVCFIMVYRFISYVIL 633
+ S+W + +TV F +R + Y L
Sbjct: 706 QLSKWNCLFITVAFGFFFRILFYFTL 731
>At2g37360 ABC transporter like protein
Length = 755
Score = 370 bits (950), Expect = e-102
Identities = 217/616 (35%), Positives = 346/616 (55%), Gaps = 40/616 (6%)
Query: 56 VLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQ--KGSVLLNQKPVDKSQFRKL 113
+L +S +AR E++A++G SG+GKS+L++ LA + +GS+ LN + ++ S + +
Sbjct: 133 LLNGISGEAREGEMMAVLGASGSGKSTLIDALANRIAKDSLRGSITLNGEVLESSMQKVI 192
Query: 114 SGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTRIGD 171
S YV Q D LFP+LTVEET+MFSA+ RL L ++++ +RV++LI +LGL A T IGD
Sbjct: 193 SAYVMQDDLLFPMLTVEETLMFSAEFRLPRSLSKKKKKARVQALIDQLGLRSAAKTVIGD 252
Query: 172 DRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTII 231
+ RG+SGGERRRVSIG ++IHDP +L LDEPTSGLDSTSA +I +L+ +A++ G +I
Sbjct: 253 EGHRGVSGGERRRVSIGNDIIHDPIILFLDEPTSGLDSTSAYMVIKVLQRIAQS-GSIVI 311
Query: 232 LSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSID 291
+SIHQP +RI+ L + L+ L+ G+ ++ G+ L +P + N EFA+D I
Sbjct: 312 MSIHQPSYRIMGLLDQLIFLSKGNTVYSGSPTHLPQFFSEFKHPIPENENKTEFALDLIR 371
Query: 292 VIQQQQQWQVE-TETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDEDIIN 350
++ + E ++ + ++++ + S K + + K++ N
Sbjct: 372 ELEYSTEGTKPLVEFHKQWRAKQAPSYNNNNKRNTNVSSLKEAITASISRGKLVSGATNN 431
Query: 351 KTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFCNLK 410
+ FAN E +++ R N R EL R ++++G++L ++F NL
Sbjct: 432 NSSNLTPSFQTFANPFWIEMIVIGKRAILNSRRQPELLGMRLGAVMVTGIILATMFTNLD 491
Query: 411 DDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGLVY 470
+ +G QER+G FAF ++ + EA+P+FLQER I M+ET+ +YR SSY ++ ++
Sbjct: 492 NSPKGAQERLGFFAFAMSTTFYTCAEAIPVFLQERYIFMRETAYNAYRRSSYVLSQSIIS 551
Query: 471 LPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIVGN 530
+P L++L+ F +W VGL+ F F I + +S V S ++PN ++G
Sbjct: 552 IPALIVLSASFAATTFWAVGLDGGANGFFFFYFTILASFWAGSSFVTFLSGVIPNVMLGF 611
Query: 531 SVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFSNSKKC------ 584
+V+ ++ F LFSG+FIS IP YW++ HYISL KYP+EG L NEF N +C
Sbjct: 612 TVVVAILAYFLLFSGFFISRDRIPVYWLWFHYISLVKYPYEGVLQNEFQNPTRCFARGVQ 671
Query: 585 ------------------LEYMFG---------ACVMKGEDVLKEEGYGGEGSRWKNVGV 617
L+ M G CV G D+LK++G + S+W + +
Sbjct: 672 LFDNSPLGEFPNDVKVNLLKSMSGVLGTNVTAETCVTTGIDILKQQGI-TDISKWNCLWI 730
Query: 618 TVCFIMVYRFISYVIL 633
TV + +R + Y L
Sbjct: 731 TVAWGFFFRVLFYFTL 746
>At3g55130 ABC transporter - like protein
Length = 725
Score = 363 bits (933), Expect = e-100
Identities = 225/644 (34%), Positives = 356/644 (54%), Gaps = 78/644 (12%)
Query: 51 SGVRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK--HRPQKGSVLLNQKPVDKS 108
+GV+ +L +VS +A +ILA++G SGAGKS+L++ LAG+ +GSV LN + V +S
Sbjct: 94 NGVKTLLDDVSGEASDGDILAVLGASGAGKSTLIDALAGRVAEGSLRGSVTLNGEKVLQS 153
Query: 109 QFRK-LSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVA 165
+ K +S YV Q D LFP+LTV+ET+MF+++ RL L + ++ RV++LI +LGL + A
Sbjct: 154 RLLKVISAYVMQDDLLFPMLTVKETLMFASEFRLPRSLSKSKKMERVEALIDQLGLRNAA 213
Query: 166 GTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAET 225
T IGD+ RG+SGGERRRVSIG+++IHDP VL LDEPTSGLDST+A ++ +LK +A++
Sbjct: 214 NTVIGDEGHRGVSGGERRRVSIGIDIIHDPIVLFLDEPTSGLDSTNAFMVVQVLKRIAQS 273
Query: 226 RGRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEF 285
G +I+SIHQP RIV+L + L++L+ G + +G+ L G +P N+ EF
Sbjct: 274 -GSIVIMSIHQPSARIVELLDRLIILSRGKSVFNGSPASLPGFFSDFGRPIPEKENISEF 332
Query: 286 AIDSIDVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVID 345
A+D + R L+G+ + K D + ++ K +L Q Q+ +D
Sbjct: 333 ALDLV----------------RELEGSNEGTKALVDFNEKWQQN-KISLIQSAPQTNKLD 375
Query: 346 ED-------IINKT--------------GTGMDFSYDFANSRLRETMILTHRFSKNIFRT 384
+D IN + T M+ +AN L ET IL R+ KN R
Sbjct: 376 QDRSLSLKEAINASVSRGKLVSGSSRSNPTSMETVSSYANPSLFETFILAKRYMKNWIRM 435
Query: 385 KELFACRTIQMLISGLVLGSIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQE 444
EL R ++++G +L +++ L RG QER+ LFAF++ + ++ +P+F+QE
Sbjct: 436 PELVGTRIATVMVTGCLLATVYWKLDHTPRGAQERLTLFAFVVPTMFYCCLDNVPVFIQE 495
Query: 445 REILMKETSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLL 504
R I ++ET+ +YR SSY I++ LV LP LL +++F+ +W VGL+ F+ + LL
Sbjct: 496 RYIFLRETTHNAYRTSSYVISHSLVSLPQLLAPSLVFSAITFWTVGLSGGLEGFVFYCLL 555
Query: 505 IWLVLYTANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYIS 564
I+ ++ +SVV S +VPN ++ V + L SG++++ IP YW + HYIS
Sbjct: 556 IYASFWSGSSVVTFISGVVPNIMLCYMVSITYLAYCLLLSGFYVNRDRIPFYWTWFHYIS 615
Query: 565 LFKYPFEGFLINEFSNSKKCL---------------------------------EYMFGA 591
+ KYP+E LINEF + +C +
Sbjct: 616 ILKYPYEAVLINEFDDPSRCFVRGVQVFDSTLLGGVSDSGKVKLLETLSKSLRTKITEST 675
Query: 592 CVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYVILRY 635
C+ G D+L ++G + S+W + +T + +R + Y L +
Sbjct: 676 CLRTGSDLLAQQGI-TQLSKWDCLWITFASGLFFRILFYFALLF 718
>At3g55100 ABC transporter - like protein
Length = 662
Score = 358 bits (920), Expect = 4e-99
Identities = 219/624 (35%), Positives = 341/624 (54%), Gaps = 61/624 (9%)
Query: 53 VRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK--HRPQKGSVLLNQKPVDKSQF 110
++ +L ++ +A+ EILAI+G SGAGKS+L++ LAG+ KG+V LN + +
Sbjct: 48 IKTLLNGITGEAKEGEILAILGASGAGKSTLIDALAGQIAEGSLKGTVTLNGEALQSRLL 107
Query: 111 RKLSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTR 168
R +S YV Q+D LFP+LTVEET+MF+A+ RL L + ++ +RV++LI +LGL V T
Sbjct: 108 RVISAYVMQEDLLFPMLTVEETLMFAAEFRLPRSLSKSKKRNRVETLIDQLGLTTVKNTV 167
Query: 169 IGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGR 228
IGD+ RG+SGGERRRVSIG ++IHDP VL LDEPTSGLDSTSA ++ +LK +A + G
Sbjct: 168 IGDEGHRGVSGGERRRVSIGTDIIHDPIVLFLDEPTSGLDSTSAFMVVQVLKKIARS-GS 226
Query: 229 TIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAID 288
+I+SIHQP RI++ + +++L++G ++ + L + G +P N+ EF +D
Sbjct: 227 IVIMSIHQPSGRIMEFLDRVIVLSSGQIVFSDSPATLPLFFSEFGSPIPEKENIAEFTLD 286
Query: 289 SIDVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDEDI 348
I + L+G+ + +G + + Q+ S + E I
Sbjct: 287 LI----------------KDLEGSPEGTRGLVEFNRNWQHRKLRVSQEPHHNSSSLGEAI 330
Query: 349 INKTGTG--MDFSY----DFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVL 402
G + SY + N ET+IL R+ N RT EL R ++++G +L
Sbjct: 331 NASISRGKLVSTSYRSIPSYVNPWWVETVILAKRYMINWTRTPELIGTRVFIVMMTGFLL 390
Query: 403 GSIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSY 462
+++ + D RG QER+ F+F + + S + LP F+QER I ++ET+ +YR SSY
Sbjct: 391 ATVYWKVDDSPRGVQERLSFFSFAMATMFYSCADGLPAFIQERYIFLRETAHNAYRRSSY 450
Query: 463 AIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSAL 522
I++ LV LP L L+I F +W VGLN F+++L++I+ ++ S V S +
Sbjct: 451 VISHSLVTLPHLFALSIGFAATTFWFVGLNGGLAGFIYYLMIIFASFWSGCSFVTFVSGV 510
Query: 523 VPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFSNSK 582
+PN ++ V G + LFSG++++ I YWI++HYISL KYP+E L NEF +
Sbjct: 511 IPNVMMSYMVTFGYLSYCLLFSGFYVNRDRIHLYWIWIHYISLLKYPYEAVLHNEFDDPS 570
Query: 583 KC------------------------LEYMFG---------ACVMKGEDVLKEEGYGGEG 609
+C LE M G C+ G D+LK+ G +
Sbjct: 571 RCFVRGNQVFDNTIMEGVSETTKAKLLETMSGYLGMELTESTCLTTGSDLLKQHGI-EQL 629
Query: 610 SRWKNVGVTVCFIMVYRFISYVIL 633
+W + VT+ + +R + Y L
Sbjct: 630 DKWGCLWVTLAWGFFFRILFYFSL 653
>At3g21090 ABC transporter, putative
Length = 680
Score = 236 bits (603), Expect = 2e-62
Identities = 181/598 (30%), Positives = 294/598 (48%), Gaps = 36/598 (6%)
Query: 54 RHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK---HRPQKGSVLLNQKPVDKSQF 110
R +L+ ++ A P I+AI+GPSG+GKS+LL+ LAG+ + G++LLN K
Sbjct: 43 RRLLQRLNGYAEPGRIMAIMGPSGSGKSTLLDSLAGRLARNVVMTGNLLLNGKKARLDY- 101
Query: 111 RKLSGYVTQKDTLFPLLTVEETMMFSAKLRLK--LPQQQQCSRVKSLIKELGLDHVAGTR 168
L YVTQ+D L LTV ET+ +SA LRL + +++ V+ I ELGL +
Sbjct: 102 -GLVAYVTQEDVLLGTLTVRETITYSAHLRLPSDMSKEEVSDIVEGTIMELGLQDCSDRV 160
Query: 169 IGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGR 228
IG+ RG+SGGER+RVSI +E++ P++L LDEPTSGLDS SA +I L+ +A GR
Sbjct: 161 IGNWHARGVSGGERKRVSIALEILTRPQILFLDEPTSGLDSASAFFVIQALRNIARD-GR 219
Query: 229 TIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAID 288
T+I S+HQP + LF+ L LL++G ++ G A G P N + +
Sbjct: 220 TVISSVHQPSSEVFALFDDLFLLSSGESVYFGEAKSAVEFFAESGFPCPKKRNPSDHFLR 279
Query: 289 SIDVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDE-- 346
I+ + T T + Q + D K L + +++SK
Sbjct: 280 CIN-----SDFDTVTATLKGSQRIQETPATSDPLMNLATSVIKARLVENYKRSKYAKSAK 334
Query: 347 ----DIINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVL 402
++ N G M+ + ++ LT R N+ R + R I ++ + +
Sbjct: 335 SRIRELSNIEGLEMEIRKGSEATWWKQLRTLTARSFINMCRDVGYYWTRIISYIVVSISV 394
Query: 403 GSIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSY 462
G+IF ++ RV FI F+ SI P FL+E ++ KE G Y VS Y
Sbjct: 395 GTIFYDVGYSYTSILARVSCGGFITGFMTFMSIGGFPSFLEEMKVFYKERLSGYYGVSVY 454
Query: 463 AIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSAL 522
++N + PFL+ ++++ Y LV F+ + F L I+ + S+++ +++
Sbjct: 455 ILSNYISSFPFLVAISVITGTITYNLVKFRPGFSHYAFFCLNIFFSVSVIESLMMVVASV 514
Query: 523 VPNFIVGNSVINGVIGSFFLFSGYFISNHEIPS-YWIF-MHYISLFKYPFEGFLINEFSN 580
VPNF++G G+IG + SG+F ++P +W + + YIS + +
Sbjct: 515 VPNFLMGLITGAGLIGIIMMTSGFFRLLPDLPKIFWRYPVSYISYGSWAIQ--------- 565
Query: 581 SKKCLEYMF-GACVMKGEDVL-KEEGYGGEGSRWKNVGVTVCFIMVYRFISYVILRYK 636
E +F G M GE+V+ K G S+W ++ V ++ YR + +V+L+ +
Sbjct: 566 ----FEPLFPGEPKMTGEEVIEKVFGVKVTYSKWWDLAAVVAILVCYRLLFFVVLKLR 619
>At3g52310 ABC transporter-like protein
Length = 737
Score = 235 bits (600), Expect = 5e-62
Identities = 188/603 (31%), Positives = 304/603 (50%), Gaps = 54/603 (8%)
Query: 51 SGVRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQK--GSVLLNQKPVDKS 108
S + +L +S A P E+LA++GPSG+GK++LL L G+ Q GSV N KP K
Sbjct: 162 SSEKSILNGISGSAYPGELLALMGPSGSGKTTLLNALGGRFNQQNIGGSVSYNDKPYSKH 221
Query: 109 QFRKLSGYVTQKDTLFPLLTVEETMMFSAKLRLK--LPQQQQCSRVKSLIKELGLDHVAG 166
++ G+VTQ D LFP LTV+ET+ ++A LRL L +Q++ R S+I+ELGL+
Sbjct: 222 LKTRI-GFVTQDDVLFPHLTVKETLTYTALLRLPKTLTEQEKEQRAASVIQELGLERCQD 280
Query: 167 TRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETR 226
T IG VRG+SGGER+RV IG E++ +P +L+LDEPTS LDST+AL+I+ ML +A+
Sbjct: 281 TMIGGSFVRGVSGGERKRVCIGNEIMTNPSLLLLDEPTSSLDSTTALKIVQMLHCIAKA- 339
Query: 227 GRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFA 286
G+TI+ +IHQP R+ F+ L++L+ GS+L+ G A +G L +N EF
Sbjct: 340 GKTIVTTIHQPSSRLFHRFDKLVVLSRGSLLYFGKASEAMSYFSSIGCSPLLAMNPAEFL 399
Query: 287 IDSIDVIQQQQQWQVETETPRRLQGTTQQKKGR----DDEQQGDDKSGKFTLQQLFQQSK 342
+D ++ V + +++ + R D E Q +++ K T + ++ K
Sbjct: 400 LDLVN--GNMNDISVPSALKEKMKIIRLELYVRNVKCDVETQYLEEAYK-TQIAVMEKMK 456
Query: 343 V-----IDEDI-----INKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRT 392
+ +DE++ K G+ + + LR H + + R
Sbjct: 457 LMAPVPLDEEVKLMITCPKREWGLSWWEQYCLLSLRGIKERRHDYFSWL---------RV 507
Query: 393 IQMLISGLVLGSIFCNLKDDLRGTQERVGLFAFILTFL-LSSSIEALPIFLQEREILMKE 451
Q+L + ++LG ++ D R GL FI F A+ F QER +L KE
Sbjct: 508 TQVLSTAIILGLLWWQ-SDITSQRPTRSGLLFFIAVFWGFFPVFTAIFTFPQERAMLSKE 566
Query: 452 TSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYT 511
YR+S+Y +A LP LIL +LF V +Y++ GL +F +L ++L +
Sbjct: 567 RESNMYRLSAYFVARTTSDLPLDLILPVLFLVVVYFMAGLRLRAESFFLSVLTVFLCIVA 626
Query: 512 ANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFE 571
A + + A + + ++ + + +F L GYF+ ++P + ++ ++S F Y
Sbjct: 627 AQGLGLAIGASLMDLKKATTLASVTVMTFMLAGGYFVK--KVPFFIAWIRFMS-FNYHTY 683
Query: 572 GFLINEFSNSKKCLEYMFGACVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYV 631
L+ ++Y + GE++ E G K V V I+ YR ++Y
Sbjct: 684 KLLVK--------VQYEEIMESVNGEEI--ESGL-------KEVSALVAMIIGYRLVAYF 726
Query: 632 ILR 634
LR
Sbjct: 727 SLR 729
>At3g13220 ABC transporter
Length = 647
Score = 234 bits (596), Expect = 1e-61
Identities = 184/607 (30%), Positives = 299/607 (48%), Gaps = 49/607 (8%)
Query: 53 VRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQ-KGSVLLNQKPVDKSQFR 111
++H+LK ++ P EILA++GPSG+GK++LL+I+ G+ KG + N P S R
Sbjct: 65 LKHILKGITGSTGPGEILALMGPSGSGKTTLLKIMGGRLTDNVKGKLTYNDIPYSPSVKR 124
Query: 112 KLSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTRI 169
++ G+VTQ D L P LTVEET+ F+A LRL + ++Q+ ++++ +IKELGL+ TR+
Sbjct: 125 RI-GFVTQDDVLLPQLTVEETLAFAAFLRLPSSMSKEQKYAKIEMIIKELGLERCRRTRV 183
Query: 170 GDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRT 229
G V+GISGGER+R SI E++ DP +L+LDEPTSGLDSTSA +++ +L+ +A+ GRT
Sbjct: 184 GGGFVKGISGGERKRASIAYEILVDPSLLLLDEPTSGLDSTSATKLLHILQGVAKA-GRT 242
Query: 230 IILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDS 289
+I +IHQP R+ +F+ LLL++ G +G A + + + +N EF +D
Sbjct: 243 VITTIHQPSSRMFHMFDKLLLISEGHPAFYGKARESMEYFSSLRILPEIAMNPAEFLLDL 302
Query: 290 I-----DVIQQQQQWQVETETPRRLQGTTQQKKGR--------DDEQQGDDKSGKFTLQQ 336
D+ + +T P + + K R + E+ ++ LQ
Sbjct: 303 ATGQVSDISLPDELLAAKTAQPDSEEVLLKYLKQRYKTDLEPKEKEENHRNRKAPEHLQI 362
Query: 337 LFQQSKVIDEDIINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELF-ACRTIQM 395
Q K D+ S + +IL+ R + R ++ F R +Q
Sbjct: 363 AIQVKK------------------DWTLSWWDQFLILSRRTFRE--RRRDYFDKLRLVQS 402
Query: 396 LISGLVLGSIFCNLKDDLRG-TQERVGLFAFILTFLLSSSI-EALPIFLQEREILMKETS 453
L +VLG ++ K D +++VGL +I F SSS+ A+ +F E+ L+KE
Sbjct: 403 LGVAVVLGLLWWKSKTDTEAHLRDQVGLMFYICIFWTSSSLFGAVYVFPFEKIYLVKERK 462
Query: 454 CGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTAN 513
YR+S Y + + L + ++ F + +Y++ N N FL +L I L+ T+
Sbjct: 463 AEMYRLSVYYVCSTLCDMVAHVLYPTFFMIIVYFMAEFNRNIPCFLFTVLTILLIAITSQ 522
Query: 514 SVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGF 573
A V + + + V+ F L GY++ + IP + ++ Y+S Y F
Sbjct: 523 GAGEFLGASVLSIKRAGMIASLVLMLFLLTGGYYVQH--IPKFMQWLKYLSFMHYGFRLL 580
Query: 574 LINEFSNSKKCLEYMFGAC--VMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYV 631
L ++S + G C + GG W V + YR +Y
Sbjct: 581 LKVQYSADQLFECGSKGGCRTLQSSSSFDTINLNGGLQELW----VLLAMAFGYRLCAYF 636
Query: 632 ILRYKCS 638
LR K S
Sbjct: 637 CLRKKIS 643
>At1g51460 ATP-dependent transmembrane transporter like protein
Length = 678
Score = 233 bits (595), Expect = 2e-61
Identities = 172/611 (28%), Positives = 310/611 (50%), Gaps = 31/611 (5%)
Query: 49 GCSGVRHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQKGSVLLNQKPVDKS 108
G + +L V+ P ILAI+GPSG+GKS+LL+ LAG+ G+V+++ K +
Sbjct: 23 GEGATKRLLNGVNGCGEPNRILAIMGPSGSGKSTLLDALAGR---LAGNVVMSGKVLVNG 79
Query: 109 QFRKL----SGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLD 162
+ R+L + YVTQ+D L LTV E++ +SA LRL KL +++ V++ I ++GL+
Sbjct: 80 KKRRLDFGAAAYVTQEDVLLGTLTVRESISYSAHLRLPSKLTREEISDIVEATITDMGLE 139
Query: 163 HVAGTRIGDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVM 222
+ IG+ +RGISGGE++R+SI +EV+ P +L LDEPTSGLDS SA ++ +L+ +
Sbjct: 140 ECSDRTIGNWHLRGISGGEKKRLSIALEVLTKPSLLFLDEPTSGLDSASAFFVVQILRNI 199
Query: 223 AETRGRTIILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNV 282
A + G+T++ SIHQP + LF+ LLLL+ G ++ G A+ + G P N
Sbjct: 200 ASS-GKTVVSSIHQPSGEVFALFDDLLLLSGGETVYFGEAESATKFFGEAGFPCPSRRNP 258
Query: 283 VEFAIDSID--------VIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKS---GK 331
+ + ++ + + ++ + + +L TT DD + ++ K
Sbjct: 259 SDHFLRCVNSDFDNVTAALVESRRINDSSFSLHQLHETTNTLDPLDDIPTAEIRTTLVRK 318
Query: 332 FTLQQLFQQSKVIDEDIINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACR 391
F S+ ++I + G + + ++ ILT R N+ R + R
Sbjct: 319 FKCSLYAAASRARIQEIASIVGIVTERKKGSQTNWWKQLRILTQRSFINMSRDLGYYWMR 378
Query: 392 TIQMLISGLVLGSIFCNLKDDLRGTQERVGLFAFILTFLLSSSIEALPIFLQEREILMKE 451
++ + +GSIF N+ + F+ F+ SI F++E ++ +E
Sbjct: 379 IAVYIVLSICVGSIFFNVGRNHTNVMSTAACGGFMAGFMTFMSIGGFQSFIEEMKVFSRE 438
Query: 452 TSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYT 511
G Y V+ Y ++N L LPF++++ + + ++V + + F + L + + T
Sbjct: 439 RLNGHYGVAVYTVSNLLSSLPFIILMCLSTSSITIYMVRFQSGGSHFFYNCLDLICAITT 498
Query: 512 ANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPS-YWIF-MHYISLFKYP 569
S ++ +++VPNF++G + G IG L +G+F ++P +W + + YI+ +
Sbjct: 499 VESCMMMIASVVPNFLMGVMLGAGYIGIMVLSAGFFRFFPDLPMVFWRYPVSYINYGAWA 558
Query: 570 FEGFLINEFSNSKKCLEYMFGACV---MKGEDVLKEE-GYGGEGSRWKNVGVTVCFIMVY 625
+G NE +EY + MKGE +L+ G E S+W ++ V + ++ Y
Sbjct: 559 LQGAYKNEMIG----VEYDSPLPLVPKMKGELILQTVLGINPESSKWLDLAVVMMILIGY 614
Query: 626 RFISYVILRYK 636
R + IL+++
Sbjct: 615 RIAFFAILKFR 625
>At5g06530 ABC transporter like protein
Length = 627
Score = 233 bits (593), Expect = 3e-61
Identities = 191/605 (31%), Positives = 302/605 (49%), Gaps = 60/605 (9%)
Query: 54 RHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQK--GSVLLNQKPVDKSQFR 111
+ +L +S P E+LA++GPSG+GK++LL +LAG+ GSV N KP K
Sbjct: 53 KEILTGISGSVNPGEVLALMGPSGSGKTTLLSLLAGRISQSSTGGSVTYNDKPYSKYLKS 112
Query: 112 KLSGYVTQKDTLFPLLTVEETMMFSAKLRLK--LPQQQQCSRVKSLIKELGLDHVAGTRI 169
K+ G+VTQ D LFP LTV+ET+ ++A+LRL L ++Q+ R +I+ELGL+ T I
Sbjct: 113 KI-GFVTQDDVLFPHLTVKETLTYAARLRLPKTLTREQKKQRALDVIQELGLERCQDTMI 171
Query: 170 GDDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRT 229
G VRG+SGGER+RVSIG E+I +P +L+LDEPTSGLDST+AL+ I ML +AE G+T
Sbjct: 172 GGAFVRGVSGGERKRVSIGNEIIINPSLLLLDEPTSGLDSTTALRTILMLHDIAEA-GKT 230
Query: 230 IILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAID- 288
+I +IHQP R+ F+ L+LL GS+L+ G + +G + +N EF +D
Sbjct: 231 VITTIHQPSSRLFHRFDKLILLGRGSLLYFGKSSEALDYFSSIGCSPLIAMNPAEFLLDL 290
Query: 289 ------------SIDVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQ- 335
+D Q ET+T + + E + ++ K L
Sbjct: 291 ANGNINDISVPSELDDRVQVGNSGRETQTGKPSPAAVHEYLVEAYETRVAEQEKKKLLDP 350
Query: 336 -QLFQQSKVIDEDIINKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFA-CRTI 393
L +++K + + GT Y IL R K R E F+ R
Sbjct: 351 VPLDEEAKAKSTRLKRQWGTCWWEQY----------CILFCRGLKE--RRHEYFSWLRVT 398
Query: 394 QMLISGLVLGSIFCNLKDDLR---GTQERVGLFAFILTFL-LSSSIEALPIFLQEREILM 449
Q+L + ++LG ++ + D+R G Q++ GL FI F A+ F QER +L
Sbjct: 399 QVLSTAVILGLLW--WQSDIRTPMGLQDQAGLLFFIAVFWGFFPVFTAIFAFPQERAMLN 456
Query: 450 KETSCGSYRVSSYAIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVL 509
KE + YR+S+Y +A LP IL LF + +Y++ GL + F +L ++L +
Sbjct: 457 KERAADMYRLSAYFLARTTSDLPLDFILPSLFLLVVYFMTGLRISPYPFFLSMLTVFLCI 516
Query: 510 YTANSVVVCFSALVPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYP 569
A + + A++ + ++ + + +F L G+F+ ++P + ++ Y+S F Y
Sbjct: 517 IAAQGLGLAIGAILMDLKKATTLASVTVMTFMLAGGFFVK--KVPVFISWIRYLS-FNYH 573
Query: 570 FEGFLINEFSNSKKCLEYMFGACVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFIS 629
L+ ++Y A + G + + V V I YR ++
Sbjct: 574 TYKLLLK--------VQYQDFAVSINGMRI---------DNGLTEVAALVVMIFGYRLLA 616
Query: 630 YVILR 634
Y+ LR
Sbjct: 617 YLSLR 621
>At3g25620 membrane transporter, putative
Length = 646
Score = 231 bits (589), Expect = 9e-61
Identities = 183/587 (31%), Positives = 313/587 (53%), Gaps = 44/587 (7%)
Query: 54 RHVLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGKHRPQ-KGSVLLNQKPVDKSQFRK 112
R VLK VS +P E+LA++GPSG+GK++L+ LAG+ + + G+V N +P S RK
Sbjct: 96 RLVLKCVSGIVKPGELLAMLGPSGSGKTTLVTALAGRLQGKLSGTVSYNGEPFTSSVKRK 155
Query: 113 LSGYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTRIG 170
+G+VTQ D L+P LTV ET+ ++A LRL +L ++++ +V+ ++ +LGL + IG
Sbjct: 156 -TGFVTQDDVLYPHLTVMETLTYTALLRLPKELTRKEKLEQVEMVVSDLGLTRCCNSVIG 214
Query: 171 DDRVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTI 230
+RGISGGER+RVSIG E++ +P +L+LDEPTSGLDST+A +I+ L+ +A GRT+
Sbjct: 215 GGLIRGISGGERKRVSIGQEMLVNPSLLLLDEPTSGLDSTTAARIVATLRSLAR-GGRTV 273
Query: 231 ILSIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLEL-PLHVNVVEFAIDS 289
+ +IHQP R+ ++F+ +L+L+ G ++ G + + +G + VN +F +D
Sbjct: 274 VTTIHQPSSRLYRMFDKVLVLSEGCPIYSGDSGRVMEYFGSIGYQPGSSFVNPADFVLDL 333
Query: 290 IDVIQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQSKVIDEDII 349
+ I + + ET RL +Q + +Q S K L ++ +
Sbjct: 334 ANGITSDTKQYDQIETNGRLDRLEEQ----NSVKQSLISSYKKNLYPPLKEE-------V 382
Query: 350 NKTGTGMDFSYDFANSRLRETMILTHRFSKNIFRTKELFACRTIQMLISGLVLGSIFCNL 409
++T F D N+RLR+ I T+R+ + + + R G ++ +
Sbjct: 383 SRT-----FPQDQTNARLRKKAI-TNRWPTSWWMQFSVLLKR-----------GLLWWHS 425
Query: 410 KDDLRGTQERVG-LFAFILTFLLSSSIEALPIFLQEREILMKETSCGSYRVSSYAIANGL 468
+ + Q++VG LF F + + A+ F QER +L+KE S G YR+SSY IA +
Sbjct: 426 R--VAHLQDQVGLLFFFSIFWGFFPLFNAIFTFPQERPMLIKERSSGIYRLSSYYIARTV 483
Query: 469 VYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSALVPNFIV 528
LP LIL +F YW+ GL + T F+ L+++ + A V + A++ +
Sbjct: 484 GDLPMELILPTIFVTITYWMGGLKPSLTTFIMTLMIVLYNVLVAQGVGLALGAILMDAKK 543
Query: 529 GNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFSNSKKCLEYM 588
++ + ++ F L GY+I + IP + ++ Y+S Y ++ L+ + E
Sbjct: 544 AATLSSVLMLVFLLAGGYYIQH--IPGFIAWLKYVSFSHYCYK-LLVGVQYTWDEVYECG 600
Query: 589 FGA-CVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYVILR 634
G C + + +K G W + + V +++YR ++Y+ LR
Sbjct: 601 SGLHCSVMDYEGIKNLRIG--NMMWDVLALAV-MLLLYRVLAYLALR 644
>At1g31770 unknown protein
Length = 648
Score = 226 bits (576), Expect = 3e-59
Identities = 175/593 (29%), Positives = 294/593 (49%), Gaps = 46/593 (7%)
Query: 56 VLKNVSFQARPWEILAIVGPSGAGKSSLLEILAGK-HRPQKGSVLLNQKPVDKSQFRKLS 114
+L ++ P E LA++GPSG+GK++LL L G+ + G V+ N +P ++ +
Sbjct: 81 ILNGITGMVCPGEFLAMLGPSGSGKTTLLSALGGRLSKTFSGKVMYNGQPFSGC-IKRRT 139
Query: 115 GYVTQKDTLFPLLTVEETMMFSAKLRL--KLPQQQQCSRVKSLIKELGLDHVAGTRIGDD 172
G+V Q D L+P LTV ET+ F+A LRL L + ++ V +I ELGL+ + IG
Sbjct: 140 GFVAQDDVLYPHLTVWETLFFTALLRLPSSLTRDEKAEHVDRVIAELGLNRCTNSMIGGP 199
Query: 173 RVRGISGGERRRVSIGVEVIHDPKVLILDEPTSGLDSTSALQIIDMLKVMAETRGRTIIL 232
RGISGGE++RVSIG E++ +P +L+LDEPTSGLDST+A +I+ +K +A + GRT++
Sbjct: 200 LFRGISGGEKKRVSIGQEMLINPSLLLLDEPTSGLDSTTAHRIVTTIKRLA-SGGRTVVT 258
Query: 233 SIHQPGFRIVKLFNSLLLLANGSVLHHGTADLLSVNLRLMGLELPLHVNVVEFAIDSIDV 292
+IHQP RI +F+ ++LL+ GS +++G A +G L VN + +D +
Sbjct: 259 TIHQPSSRIYHMFDKVVLLSEGSPIYYGAASSAVEYFSSLGFSTSLTVNPADLLLDLANG 318
Query: 293 IQQQQQWQVETETPRRLQGTTQQKKGRDDEQQGDDKSGKFTLQQLFQQ--SKVIDEDIIN 350
I Q K+ + EQ K+ K TL +++ S + ++ N
Sbjct: 319 IPPDTQ-----------------KETSEQEQ----KTVKETLVSAYEKNISTKLKAELCN 357
Query: 351 KTGTGMDFSYDFANSRLRETMILTHRFSKNIF-------RTKELFACRTIQMLISGLVLG 403
+++ A + E T + + R E F I +IS LG
Sbjct: 358 AESHSYEYTKAAAKNLKSEQWCTTWWYQFTVLLQRGVRERRFESFNKLRIFQVISVAFLG 417
Query: 404 SIFCNLKDDLRGTQERVGLFAFILTFL-LSSSIEALPIFLQEREILMKETSCGSYRVSSY 462
+ Q+R L F F A+ F QE+ +L+KE S G YR+SSY
Sbjct: 418 GLLW-WHTPKSHIQDRTALLFFFSVFWGFYPLYNAVFTFPQEKRMLIKERSSGMYRLSSY 476
Query: 463 AIANGLVYLPFLLILAILFTVPLYWLVGLNTNFTAFLHFLLLIWLVLYTANSVVVCFSAL 522
+A + LP L L F +YW+ GL + T F+ LL++ + A + + F AL
Sbjct: 477 FMARNVGDLPLELALPTAFVFIIYWMGGLKPDPTTFILSLLVVLYSVLVAQGLGLAFGAL 536
Query: 523 VPNFIVGNSVINGVIGSFFLFSGYFISNHEIPSYWIFMHYISLFKYPFEGFLINEFSNSK 582
+ N ++ + F + GY++ +IP + +++ Y+S Y ++ L ++++
Sbjct: 537 LMNIKQATTLASVTTLVFLIAGGYYV--QQIPPFIVWLKYLSYSYYCYKLLLGIQYTDDD 594
Query: 583 --KCLEYMFGACVMKGEDVLKEEGYGGEGSRWKNVGVTVCFIMVYRFISYVIL 633
+C + ++ C + +K G + W +V V ++ YR ++Y+ L
Sbjct: 595 YYECSKGVW--CRVGDFPAIKSMGL---NNLWIDVFVMGVMLVGYRLMAYMAL 642
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.140 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,879,002
Number of Sequences: 26719
Number of extensions: 602993
Number of successful extensions: 2257
Number of sequences better than 10.0: 140
Number of HSP's better than 10.0 without gapping: 125
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 1734
Number of HSP's gapped (non-prelim): 258
length of query: 640
length of database: 11,318,596
effective HSP length: 106
effective length of query: 534
effective length of database: 8,486,382
effective search space: 4531727988
effective search space used: 4531727988
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC122164.8


