

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC122161.8 - phase: 0 /pseudo
(473 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
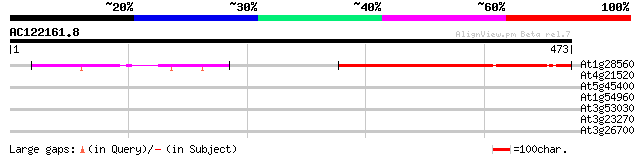
Score E
Sequences producing significant alignments: (bits) Value
At1g28560 hypothetical protein 256 2e-68
At4g21520 unknown protein 32 0.76
At5g45400 replication protein A1-like 32 0.99
At1g54960 NPK1-related protein kinase 2 (ANP2) 30 2.2
At3g53030 serine protein kinase - like 30 2.9
At3g23270 hypothetical protein 29 4.9
At3g26700 protein kinase -like protein 29 6.4
>At1g28560 hypothetical protein
Length = 482
Score = 256 bits (654), Expect = 2e-68
Identities = 122/196 (62%), Positives = 149/196 (75%), Gaps = 4/196 (2%)
Query: 278 QKKGQHDPSGYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEAQKKWEYIINGELQQK 337
QK G++DPSGYFLIEDVF+ DLR+PSA D + PILDWL NSK+EA KKWE ++ GELQ+K
Sbjct: 290 QKAGKYDPSGYFLIEDVFHNDLRNPSAKDYSYPILDWLWNSKDEALKKWECVLTGELQKK 349
Query: 338 QKAIVGEASVSHLPRFASFEMHKIHFCDLGFRLGAGYLYCHQGDCTHTLVIRDMRLIHAD 397
QK ++GEA LPR+ + +M HFCD+ FR+GA Y+YCHQGDC HT+VIRDMR+ H +
Sbjct: 350 QKLVLGEAKSVDLPRYRTADMQSTHFCDIRFRVGASYVYCHQGDCKHTIVIRDMRMSHPE 409
Query: 398 DVQNWAVYPIVTFQLKIRFQKCGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAED 457
DVQN A YPI+ F K R QKCGVCKI RA+KV VDDKW +N YFCD CF LLH +E+
Sbjct: 410 DVQNRAAYPIM-FWPKRRIQKCGVCKIKRASKVAVDDKWASENSSYFCDVCFELLH-SEE 467
Query: 458 GGSPMYTDFIEYDYNH 473
G P+ DF +DY H
Sbjct: 468 G--PLNCDFPVFDYVH 481
Score = 77.8 bits (190), Expect = 1e-14
Identities = 59/184 (32%), Positives = 88/184 (47%), Gaps = 42/184 (22%)
Query: 19 IPRGGPIYVSNMSGPIVRVPLFQDSILTQLHSLQSELPPDS------NHDISVDDLKVFT 72
+PRGGPIY+ NM G I P F+ S L L L++ L DS D+S+D LK++T
Sbjct: 22 VPRGGPIYLPNMVGNISTTPEFKSSFLNVLQDLETHLSLDSTSSSSQQFDVSIDSLKIYT 81
Query: 73 EDDLMDMALKQVFQGRDNNQDPPNAELVTSLPLFFFIL*YH*LIMMDVVVLNNID*ILNL 132
+++L +MA+K+ F +Q+ EL SL N+ N
Sbjct: 82 DEELTEMAMKEAFPEDYLSQE----ELEPSL---------------------NVSHHENP 116
Query: 133 VAG-------VRRSNSGESLV*QINPFLMLQSNCKE----KVEEVVRIKQKQEEDKAQVR 181
+AG V+ + + + M + +E KVE++ ++KQKQEEDKA V
Sbjct: 117 LAGRAKRKRTVKNTEVKKRTLKNTEVMKMTEKKTEEAYLVKVEQLAKLKQKQEEDKAAVT 176
Query: 182 LHSF 185
LH F
Sbjct: 177 LHCF 180
>At4g21520 unknown protein
Length = 425
Score = 32.0 bits (71), Expect = 0.76
Identities = 13/28 (46%), Positives = 16/28 (56%), Gaps = 3/28 (10%)
Query: 435 KWTPDNPCYFC---DECFSLLHLAEDGG 459
KW+PD C+ D SL HL +DGG
Sbjct: 58 KWSPDGSCFLASSEDNTLSLFHLPQDGG 85
>At5g45400 replication protein A1-like
Length = 853
Score = 31.6 bits (70), Expect = 0.99
Identities = 16/46 (34%), Positives = 24/46 (51%), Gaps = 2/46 (4%)
Query: 240 QKGQHTG--SSGAHTCAGFRSCSLCRDFP*FSKRCQAIQCQKKGQH 283
Q GQ+ SS A + GF SC++CR S C + + +GQ+
Sbjct: 790 QVGQYGNQYSSDARSLGGFTSCNVCRSNSHVSANCPTLMSEPQGQY 835
>At1g54960 NPK1-related protein kinase 2 (ANP2)
Length = 651
Score = 30.4 bits (67), Expect = 2.2
Identities = 12/40 (30%), Positives = 24/40 (60%)
Query: 321 EAQKKWEYIINGELQQKQKAIVGEASVSHLPRFASFEMHK 360
E ++KW+ ++ EL++K++ I +A + PR S H+
Sbjct: 602 EIERKWKEELDQELERKRREITRQAGMGSSPRDRSLSRHR 641
>At3g53030 serine protein kinase - like
Length = 529
Score = 30.0 bits (66), Expect = 2.9
Identities = 14/42 (33%), Positives = 21/42 (49%)
Query: 215 LQDQSICQQINQNSKDDVS*VHKFRQKGQHTGSSGAHTCAGF 256
+ + +I QQI + DD V K +H+G +G H C F
Sbjct: 84 MDEITILQQIAEGDTDDTKCVVKLLDHFKHSGPNGQHVCMVF 125
>At3g23270 hypothetical protein
Length = 1045
Score = 29.3 bits (64), Expect = 4.9
Identities = 16/42 (38%), Positives = 19/42 (45%), Gaps = 2/42 (4%)
Query: 419 CGVCKIFRATKVTVDDKWTPDNPCYFCDECFSLLHLAEDGGS 460
C C +A K + TP P CD C+S L AE G S
Sbjct: 629 CHACSSKKALKAALAP--TPGKPHRVCDACYSKLKAAESGYS 668
>At3g26700 protein kinase -like protein
Length = 380
Score = 28.9 bits (63), Expect = 6.4
Identities = 13/36 (36%), Positives = 21/36 (58%)
Query: 287 GYFLIEDVFYTDLRDPSAIDLTRPILDWLQNSKEEA 322
G FL+E V + +PS T+ ++DW+QN + A
Sbjct: 266 GVFLLELVSGREASEPSPSSSTQTLVDWMQNLTDYA 301
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.334 0.146 0.467
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,753,188
Number of Sequences: 26719
Number of extensions: 452285
Number of successful extensions: 1806
Number of sequences better than 10.0: 7
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 1795
Number of HSP's gapped (non-prelim): 10
length of query: 473
length of database: 11,318,596
effective HSP length: 103
effective length of query: 370
effective length of database: 8,566,539
effective search space: 3169619430
effective search space used: 3169619430
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC122161.8


