

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC121244.7 - phase: 0
(418 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
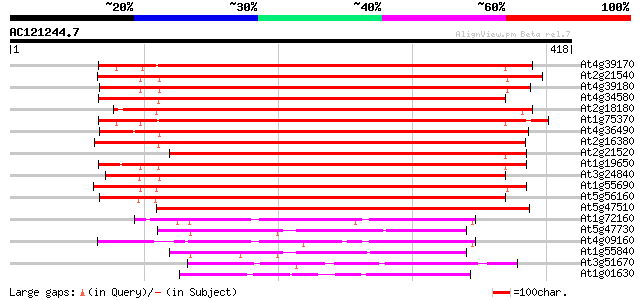
Sequences producing significant alignments: (bits) Value
At4g39170 unknown protein 407 e-114
At2g21540 putative phosphatidylinositol/phophatidylcholine trans... 391 e-109
At4g39180 SEC14 - like protein 385 e-107
At4g34580 putative protein 373 e-103
At2g18180 putative phosphatidylinositol/phophatidylcholine trans... 370 e-102
At1g75370 sec14 cytosolic factor, putative 369 e-102
At4g36490 hypothetical protein 366 e-101
At2g16380 putative phosphatidylinositol/phosphatidylcholine tran... 366 e-101
At2g21520 putative phosphatidylinositol/phosphatidylcholine tran... 352 2e-97
At1g19650 sec14 cytosolic factor- like protein 344 5e-95
At3g24840 phosphatidylinositol transfer protein like 322 2e-88
At1g55690 Phosphatidylinositol Transfer Protein like 299 2e-81
At5g56160 putative protein 278 3e-75
At5g47510 putative protein 246 2e-65
At1g72160 cytosolic factor, putative 71 1e-12
At5g47730 putative protein 69 5e-12
At4g09160 putative protein 64 1e-10
At1g55840 polyphosphoinositide binding protein, putative 63 3e-10
At3g51670 unknown protein 62 8e-10
At1g01630 polyphosphoinositide binding protein, putative 57 1e-08
>At4g39170 unknown protein
Length = 614
Score = 407 bits (1045), Expect = e-114
Identities = 208/381 (54%), Positives = 261/381 (67%), Gaps = 59/381 (15%)
Query: 67 SDENRKERRSDF--------------KKKLLNAFSKFKHSFKKKR-----------VDEL 101
S + +KER+SDF KKK +NA +KFKHS KKKR ++++
Sbjct: 18 SSDEKKERKSDFENSEDERRTRIGSLKKKAINASTKFKHSLKKKRRKSDVRVSSVSIEDV 77
Query: 102 QAVNVVPAVVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKE 161
+ V + A VD FR+ L+M+ELLP KHDDYHMMLRFLKARKFDI KAKHMWADM++WRKE
Sbjct: 78 RDVEELQA-VDEFRQALVMEELLPHKHDDYHMMLRFLKARKFDIEKAKHMWADMIQWRKE 136
Query: 162 FGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVK 221
FG DTI++DF+F E++EV+KY PHGYH VDKEGRPV+IER K+D NKLMQVTT+DRY++
Sbjct: 137 FGTDTIIQDFQFEEIDEVLKYYPHGYHSVDKEGRPVYIERLGKVDPNKLMQVTTLDRYIR 196
Query: 222 YHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDN 281
YH + E +KFPACTIA+K++IDSS TILD+QG+G N ++ RE++ R KI DN
Sbjct: 197 YHVKEFERSFMLKFPACTIAAKKYIDSSTTILDVQGVGLKNFTKSARELITRLQKIDGDN 256
Query: 282 YPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFL 341
YP+T Q FIIN G R L S + F+DPK SK+HV+G +YQ KLL++ID+SELP FL
Sbjct: 257 YPETLHQMFIINAGPGFRLLWSTVKSFLDPKTTSKIHVLGCKYQSKLLEIIDSSELPEFL 316
Query: 342 GGTCTCANQGGCLRSDKGPWNNPEILK--------------------------------- 368
GG CTCA+QGGC+ SDKGPW NPEI+K
Sbjct: 317 GGACTCADQGGCMLSDKGPWKNPEIVKMVLHGGAHRAKQVVKVLNSDGKVIAYAKPSYPW 376
Query: 369 VKGSDTLTAESSSEAEDNVIA 389
+KGSDT TAES SEAED V++
Sbjct: 377 IKGSDTSTAESGSEAEDIVVS 397
>At2g21540 putative phosphatidylinositol/phophatidylcholine transfer
protein
Length = 371
Score = 391 bits (1005), Expect = e-109
Identities = 188/343 (54%), Positives = 251/343 (72%), Gaps = 11/343 (3%)
Query: 66 GSDENRKERRSDFKKKLLNAFSKFKHSFKKK--RVDELQAVNVVPAV-------VDAFRK 116
GS++ +K + KKK +NA +KFKHSF K+ R + +V++V + VDAFR+
Sbjct: 19 GSEDEKKTKLCSLKKKAINASNKFKHSFTKRTRRNSRVMSVSIVDDIDLEELQAVDAFRQ 78
Query: 117 LLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNEL 176
LI+DELLP KHDD+HMMLRFL+ARKFD+ KAK MW DM+ WRKEFG DTIMEDF+F E+
Sbjct: 79 ALILDELLPSKHDDHHMMLRFLRARKFDLEKAKQMWTDMIHWRKEFGVDTIMEDFDFKEI 138
Query: 177 NEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFP 236
+EV+KY P GYHGVDK+GRPV+IER ++D KLMQVTTIDRYVKYH + E+ IK P
Sbjct: 139 DEVLKYYPQGYHGVDKDGRPVYIERLGQVDATKLMQVTTIDRYVKYHVREFEKTFNIKLP 198
Query: 237 ACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGL 296
AC+IA+K+HID S TILD+QG+G + +A R++++R KI DNYP+T + FIIN G
Sbjct: 199 ACSIAAKKHIDQSTTILDVQGVGLKSFSKAARDLLQRIQKIDSDNYPETLNRMFIINAGS 258
Query: 297 ELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRS 356
R L S + F+DPK +K+HV+G++YQ KLL++ID++ELP FLGG CTCA++GGC+RS
Sbjct: 259 GFRLLWSTVKSFLDPKTTAKIHVLGNKYQSKLLEIIDSNELPEFLGGNCTCADKGGCMRS 318
Query: 357 DKGPWNNPEILKV--KGSDTLTAESSSEAEDNVIAVCEEVNVV 397
DKGPWN+P+I K+ G ++ S E+ I+V E +V
Sbjct: 319 DKGPWNDPDIFKMVQNGEGKCPRKTLSNIEEKTISVDENTTMV 361
>At4g39180 SEC14 - like protein
Length = 554
Score = 385 bits (989), Expect = e-107
Identities = 186/332 (56%), Positives = 241/332 (72%), Gaps = 11/332 (3%)
Query: 68 DENRKERRSDFKKKLLNAFSKFKHSFKKK--RVDELQAVNVVPAV-------VDAFRKLL 118
D+ R + KKK +NA +KFKHS KK R + V++V + VDAFR+ L
Sbjct: 22 DDKRLTKLCSLKKKAINATNKFKHSMTKKGRRHSRVACVSIVDEIDTEELQAVDAFRQAL 81
Query: 119 IMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNE 178
I+DELLP KHDD+HMMLRFL+ARKFD+ KAK MW+DML WRKE+GADTIMEDF+F E+ E
Sbjct: 82 ILDELLPSKHDDHHMMLRFLRARKFDLEKAKQMWSDMLNWRKEYGADTIMEDFDFKEIEE 141
Query: 179 VIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPAC 238
V+KY P GYHGVDKEGRP++IER ++D KLM+VTTIDRYVKYH + E+ +KFPAC
Sbjct: 142 VVKYYPQGYHGVDKEGRPIYIERLGQVDATKLMKVTTIDRYVKYHVKEFEKTFNVKFPAC 201
Query: 239 TIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLEL 298
+IA+KRHID S TILD+QG+G N +A +++++ KI DNYP+T + FIIN G
Sbjct: 202 SIAAKRHIDQSTTILDVQGVGLSNFNKAAKDLLQSIQKIDNDNYPETLNRMFIINAGCGF 261
Query: 299 RSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDK 358
R L + + F+DPK +K+HV+G++YQ KLL++IDA+ELP FLGG CTCA++GGC+RSDK
Sbjct: 262 RLLWNTVKSFLDPKTTAKIHVLGNKYQTKLLEIIDANELPEFLGGKCTCADKGGCMRSDK 321
Query: 359 GPWNNPEILKV--KGSDTLTAESSSEAEDNVI 388
GPWN+PEI K+ G S S E+ I
Sbjct: 322 GPWNDPEIFKLVQNGEGRCLRRSLSGIEEKTI 353
>At4g34580 putative protein
Length = 560
Score = 373 bits (957), Expect = e-103
Identities = 178/314 (56%), Positives = 235/314 (74%), Gaps = 11/314 (3%)
Query: 67 SDENRK-ERRSDFKKKLLNAFSKFKHSFKKKRVDELQAVNVVPA----------VVDAFR 115
S+E RK + S KKK +NA ++FK+SFKKK V VP +DAFR
Sbjct: 11 SEEERKIVKISSLKKKAINASNRFKNSFKKKGRRSSSRVMSVPIEDDIDAEDLQALDAFR 70
Query: 116 KLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNE 175
+ LI+DELLP K DD HMMLRFL+ARKFDI KAK MW+DM++WRK+FGADTI+EDF+F E
Sbjct: 71 QALILDELLPSKLDDLHMMLRFLRARKFDIEKAKQMWSDMIQWRKDFGADTIIEDFDFEE 130
Query: 176 LNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKF 235
++EV+K+ P GYHGVDKEGRPV+IER ++D NKL+QVTT+DRYVKYH + E+ +KF
Sbjct: 131 IDEVMKHYPQGYHGVDKEGRPVYIERLGQIDANKLLQVTTMDRYVKYHVKEFEKTFKVKF 190
Query: 236 PACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVG 295
P+C++A+ +HID S TILD+QG+G N ++ RE+++R KI +NYP+T + FIIN G
Sbjct: 191 PSCSVAANKHIDQSTTILDVQGVGLKNFSKSARELLQRLCKIDNENYPETLNRMFIINAG 250
Query: 296 LELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLR 355
R L S + F+DPK +K+HV+G++Y KLL+VIDASELP F GG CTC ++GGC+R
Sbjct: 251 SGFRLLWSTVKSFLDPKTTAKIHVLGNKYHSKLLEVIDASELPEFFGGACTCEDKGGCMR 310
Query: 356 SDKGPWNNPEILKV 369
SDKGPWN+PE+LK+
Sbjct: 311 SDKGPWNDPEVLKI 324
>At2g18180 putative phosphatidylinositol/phophatidylcholine transfer
protein
Length = 558
Score = 370 bits (949), Expect = e-102
Identities = 182/324 (56%), Positives = 237/324 (72%), Gaps = 15/324 (4%)
Query: 78 FKKKLLNAFSKFKHSF-KKKRVDELQAVNVVP--------AVVDAFRKLLIMDELLPQKH 128
FKK+ + SK S KK+R ++ +V + VVDAFR++LI+DELLP KH
Sbjct: 20 FKKR---SCSKLSCSLTKKRRSSKVMSVEIFEDEHDAEELKVVDAFRQVLILDELLPDKH 76
Query: 129 DDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYH 188
DDYHMMLRFLKARKFD+ K MW+DML WRKEFGADT+MEDFEF E++EV+KY P G+H
Sbjct: 77 DDYHMMLRFLKARKFDLEKTNQMWSDMLRWRKEFGADTVMEDFEFKEIDEVLKYYPQGHH 136
Query: 189 GVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDS 248
GVDKEGRPV+IER ++D KLMQVTT+DRYV YH E +KFPAC+IA+K+HID
Sbjct: 137 GVDKEGRPVYIERLGQVDSTKLMQVTTMDRYVNYHVMEFERTFNVKFPACSIAAKKHIDQ 196
Query: 249 SITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYF 308
S TILD+QG+G N +A R+++ R K+ DNYP+T + FIIN G R L + + F
Sbjct: 197 STTILDVQGVGLKNFNKAARDLITRLQKVDGDNYPETLNRMFIINAGSGFRMLWNTVKSF 256
Query: 309 MDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGPWNNPEILK 368
+DPK +K+HV+G++YQ KLL++IDASELP FLGG+CTCA+ GGC+RSDKGPWNNP+I+K
Sbjct: 257 LDPKTTAKIHVLGNKYQSKLLEIIDASELPEFLGGSCTCADNGGCMRSDKGPWNNPDIMK 316
Query: 369 -VKGSDTLTAESS--SEAEDNVIA 389
V D + ++ S A +N+I+
Sbjct: 317 RVNNGDHICSKRSQADNAGENIIS 340
>At1g75370 sec14 cytosolic factor, putative
Length = 612
Score = 369 bits (946), Expect = e-102
Identities = 196/396 (49%), Positives = 254/396 (63%), Gaps = 65/396 (16%)
Query: 67 SDENRKERRSDF----------------KKKLLNAFSKFKHSFKKK-------------- 96
S++ R+ERRSDF KKK A SK +HS KKK
Sbjct: 18 SNDERRERRSDFEVSEDEKKTRIGNFNFKKKAAKASSKLRHSLKKKGSSRRRSSDRTFSL 77
Query: 97 RVDELQAVNVVPAVVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADML 156
++++ V + AV D FR LL+ + LLP DDYH+MLRFLKARKFDIGK K MW++M+
Sbjct: 78 TIEDIHDVEELRAV-DEFRNLLVSENLLPPTLDDYHIMLRFLKARKFDIGKTKLMWSNMI 136
Query: 157 EWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTI 216
+WRK+FG DTI EDFEF E +EV+KY PHGYHGVDKEGRPV+IER +D KLMQVTT+
Sbjct: 137 KWRKDFGTDTIFEDFEFEEFDEVLKYYPHGYHGVDKEGRPVYIERLGLVDPAKLMQVTTV 196
Query: 217 DRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLK 276
+R+++YH + E+ IK PAC IA+KRHIDSS TILD+QG+GF N + R+++ + K
Sbjct: 197 ERFIRYHVREFEKTVNIKLPACCIAAKRHIDSSTTILDVQGVGFKNFSKPARDLIIQLQK 256
Query: 277 ILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASE 336
I DNYP+T + FIIN G + + + + F+DPK +K+HVIG++YQ KLL++IDAS+
Sbjct: 257 IDNDNYPETLHRMFIINGGSGFKLVWATVKQFLDPKTVTKIHVIGNKYQNKLLEIIDASQ 316
Query: 337 LPTFLGGTCTCANQGGCLRSDKGPWNNPEILK---------------------------- 368
LP FLGGTCTCA++GGC+RSDKGPWN+PEILK
Sbjct: 317 LPDFLGGTCTCADRGGCMRSDKGPWNDPEILKMLQSGGPLCRHNSALNSFSRVSSCDKPS 376
Query: 369 ---VKGSDTLTAESSSEAEDNVIAVCEEVNVVLRLP 401
+K SDT TAES SE E+ +VN LR+P
Sbjct: 377 FSGIKASDTSTAESGSEVEE---MASPKVNRELRVP 409
>At4g36490 hypothetical protein
Length = 543
Score = 366 bits (939), Expect = e-101
Identities = 175/327 (53%), Positives = 233/327 (70%), Gaps = 9/327 (2%)
Query: 68 DENRKERRSDFKKKLLNAFSKFKHSFKKKRVDELQAVNVVPAV--------VDAFRKLLI 119
D K R FKK+ + ++ + K++R ++ +V ++ V VDAFR+ LI
Sbjct: 6 DAELKPRMGSFKKRSSSKNLRYSMT-KRRRSSKVMSVEIIEDVHDAEELKAVDAFRQSLI 64
Query: 120 MDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEV 179
+DELLP+KHDDYHMMLRFLKARKFD+ K K MW +ML WRKEFGADT+ME+F+F E++EV
Sbjct: 65 LDELLPEKHDDYHMMLRFLKARKFDLEKTKQMWTEMLRWRKEFGADTVMEEFDFKEIDEV 124
Query: 180 IKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACT 239
+KY P G+HGVDKEGRPV+IER +D KLMQVTT+DRYV YH E +KFPAC+
Sbjct: 125 LKYYPQGHHGVDKEGRPVYIERLGLVDSTKLMQVTTMDRYVNYHVMEFERTFNVKFPACS 184
Query: 240 IASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELR 299
IA+K+HID S TILD+QG+G N +A R+++ R K+ DNYP+T + FIIN G R
Sbjct: 185 IAAKKHIDQSTTILDVQGVGLKNFNKAARDLITRLQKVDGDNYPETLNRMFIINAGSGFR 244
Query: 300 SLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKG 359
L + + F+DPK +K+HV+G++YQ KLL++ID SELP FLGG+CTCA+ GGC+RSDKG
Sbjct: 245 MLWNTVKSFLDPKTTAKIHVLGNKYQSKLLEIIDESELPEFLGGSCTCADNGGCMRSDKG 304
Query: 360 PWNNPEILKVKGSDTLTAESSSEAEDN 386
PW NPEI+K + S+AE++
Sbjct: 305 PWKNPEIMKRVHNGDHKCSKGSQAENS 331
>At2g16380 putative phosphatidylinositol/phosphatidylcholine
transfer protein
Length = 582
Score = 366 bits (939), Expect = e-101
Identities = 178/332 (53%), Positives = 242/332 (72%), Gaps = 11/332 (3%)
Query: 64 LTGSDENRK-ERRSDFKKKLLNAFSKFKHSFKKK-RVDELQAVNVVPA---------VVD 112
+ S++ RK + S K+K ++A ++FK+SFKKK R + V+V V+
Sbjct: 6 MENSEDGRKLVKMSSLKQKAISASNRFKNSFKKKTRRTSSKIVSVANTDDINGDDYLSVE 65
Query: 113 AFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFE 172
AFR++L++D+LLP KHDD HMMLRFL+ARKFD KAK MW+DML+WR +FG DTI+EDFE
Sbjct: 66 AFRQVLVLDDLLPPKHDDLHMMLRFLRARKFDKEKAKQMWSDMLQWRMDFGVDTIIEDFE 125
Query: 173 FNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHA 232
F E+++V+K+ P GYHGVDKEGRPV+IER ++D NKL+Q TT+DRY KYH + E+M
Sbjct: 126 FEEIDQVLKHYPQGYHGVDKEGRPVYIERLGQIDANKLLQATTMDRYEKYHVKEFEKMFK 185
Query: 233 IKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFII 292
IKFP+C+ A+K+HID S TI D+QG+G N ++ RE+++R LKI DNYP+T + FII
Sbjct: 186 IKFPSCSAAAKKHIDQSTTIFDVQGVGLKNFNKSARELLQRLLKIDNDNYPETLNRMFII 245
Query: 293 NVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGG 352
N G R L + + F+DPK SK+HV+G++YQ KLL+ IDASELP F GG CTCA++GG
Sbjct: 246 NAGPGFRLLWAPIKKFLDPKTTSKIHVLGNKYQPKLLEAIDASELPYFFGGLCTCADKGG 305
Query: 353 CLRSDKGPWNNPEILKVKGSDTLTAESSSEAE 384
CLRSDKGPWN+PE+LK+ + + SE +
Sbjct: 306 CLRSDKGPWNDPELLKIARNPEARFSTISEED 337
>At2g21520 putative phosphatidylinositol/phosphatidylcholine
transfer protein
Length = 531
Score = 352 bits (903), Expect = 2e-97
Identities = 170/299 (56%), Positives = 214/299 (70%), Gaps = 33/299 (11%)
Query: 120 MDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEV 179
MDELLP +HDDYHMMLRFLKARKFD+ KAK MWADM++WRKEFG DTI++DF+F E+NEV
Sbjct: 1 MDELLPDRHDDYHMMLRFLKARKFDVEKAKQMWADMIQWRKEFGTDTIIQDFDFEEINEV 60
Query: 180 IKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACT 239
+K+ P YHGVDKEGRP++IER K+D N+LMQVT++DRYV+YH + E IKFP+CT
Sbjct: 61 LKHYPQCYHGVDKEGRPIYIERLGKVDPNRLMQVTSMDRYVRYHVKEFERSFMIKFPSCT 120
Query: 240 IASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELR 299
I++KRHIDSS TILD+QG+G N ++ R+++ R KI DNYP+T Q FIIN G R
Sbjct: 121 ISAKRHIDSSTTILDVQGVGLKNFNKSARDLITRLQKIDGDNYPETLHQMFIINAGPGFR 180
Query: 300 SLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKG 359
L + + F+DPK ++K+HV+G +Y KLL+VID +ELP FLGG CTCA+QGGC+ SDKG
Sbjct: 181 LLWNTVKSFLDPKTSAKIHVLGYKYLSKLLEVIDVNELPEFLGGACTCADQGGCMLSDKG 240
Query: 360 PWNNPEILK---------------------------------VKGSDTLTAESSSEAED 385
PW NPEI+K +KGSDT TAES S+AED
Sbjct: 241 PWKNPEIVKMVLHGGAHRARQVVKVLNSEGKVIAYAKPSYTWIKGSDTSTAESGSDAED 299
>At1g19650 sec14 cytosolic factor- like protein
Length = 608
Score = 344 bits (883), Expect = 5e-95
Identities = 180/360 (50%), Positives = 238/360 (66%), Gaps = 42/360 (11%)
Query: 67 SDENRKERRSDFKKKLLNAFSKFKHSFKKK------RVDELQAVNVVPA----VVDAFRK 116
S++ +K R KK ++ SKF+HS K++ R L ++ A V FR+
Sbjct: 29 SEDEKKTRIGGILKKK-SSKSKFRHSLKRRGSRSIDRTLSLTFEDIHDAEELRYVSEFRQ 87
Query: 117 LLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNEL 176
LI D LLP DDYH+MLRFL ARKFD+GKAK MW +M++WR++FG DTI+EDFEF EL
Sbjct: 88 SLISDHLLPPNLDDYHIMLRFLFARKFDLGKAKLMWTNMIQWRRDFGTDTILEDFEFPEL 147
Query: 177 NEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFP 236
+EV++Y P GYHGVDKEGRPV+IER K+D +KLMQVTT++RY++YH + E+ +KFP
Sbjct: 148 DEVLRYYPQGYHGVDKEGRPVYIERLGKVDASKLMQVTTLERYLRYHVKEFEKTITVKFP 207
Query: 237 ACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGL 296
AC IA+KRHIDSS TILD+QG+G N + R+++ + KI DNYP+T + FIIN G
Sbjct: 208 ACCIAAKRHIDSSTTILDVQGLGLKNFTKTARDLIIQLQKIDSDNYPETLHRMFIINAGS 267
Query: 297 ELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRS 356
+ L + F+DPK SK+HV+G++YQ KLL++IDAS+LP F GGTCTCA+QGGC+RS
Sbjct: 268 GFKLLWGTVKSFLDPKTVSKIHVLGNKYQNKLLEMIDASQLPDFFGGTCTCADQGGCMRS 327
Query: 357 DKGPWNNPEILK-------------------------------VKGSDTLTAESSSEAED 385
DKGPW + EILK +K SDT TA+S SE E+
Sbjct: 328 DKGPWKDSEILKMGRSGGTFCRHAGAFLSSDSQISSSDKPTYSLKVSDTSTAKSGSELEE 387
>At3g24840 phosphatidylinositol transfer protein like
Length = 579
Score = 322 bits (826), Expect = 2e-88
Identities = 154/308 (50%), Positives = 218/308 (70%), Gaps = 10/308 (3%)
Query: 72 KERRSDFKKKLLNAFSKFKHSFKK--KRVDELQAVNVVPAV--------VDAFRKLLIMD 121
+ R KKK + A +K HS +K KRV + A V+ V V+ FRK L+
Sbjct: 32 RTRSRSLKKKAIKASNKLTHSLRKRGKRVADQYAPIVIEDVRDEEEEKAVNVFRKALVSL 91
Query: 122 ELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIK 181
+LLP +HDDYH MLRFLKAR+FD+ K MW +ML+WRKE G DTI++DF ++E EV +
Sbjct: 92 DLLPPRHDDYHTMLRFLKARRFDLEKTVQMWEEMLKWRKENGVDTIIQDFVYDEYEEVQQ 151
Query: 182 YNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIA 241
Y PHGYHGVD+EGRPV+IER K+D KLM+VTT++R+++YH Q E+ + KFPAC+IA
Sbjct: 152 YYPHGYHGVDREGRPVYIERLGKIDPGKLMKVTTLERFLRYHVQGFEKTFSEKFPACSIA 211
Query: 242 SKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSL 301
+KRHI+SS TI+D+ G+ + + + ++++ R KI DNYP+T Q +IIN G + +
Sbjct: 212 AKRHINSSTTIIDVHGVSWMSFRKLAQDLVMRMQKIDGDNYPETLNQMYIINAGNGFKLV 271
Query: 302 RSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGPW 361
+ + F+DPK SK+HV+G++Y+ LL++ID SELP FLGG C CA++GGC+R +KGPW
Sbjct: 272 WNTVKGFLDPKTTSKIHVLGNKYRSHLLEIIDPSELPEFLGGNCKCAHEGGCMRFNKGPW 331
Query: 362 NNPEILKV 369
N+PEI+K+
Sbjct: 332 NDPEIMKL 339
>At1g55690 Phosphatidylinositol Transfer Protein like
Length = 625
Score = 299 bits (765), Expect = 2e-81
Identities = 157/338 (46%), Positives = 220/338 (64%), Gaps = 15/338 (4%)
Query: 63 FLTGSDENRKERRSDFKKKLLNAFSKFKHSFKKK---RVD-ELQAVNVVP-------AVV 111
F DE R+ + + KKK +NA +KF HS KK+ ++D + AV++ +VV
Sbjct: 20 FEISEDERRRSKIGNLKKKAINASTKFTHSLKKRGKRKIDYRVPAVSIEDVRDEKEESVV 79
Query: 112 DAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDF 171
FR+ L+ +LLP +HD+YH +LRFLKAR +I K +W +ML WRKE+G DTI+EDF
Sbjct: 80 LEFRRKLLERDLLPPRHDEYHTLLRFLKARDLNIEKTTQLWEEMLRWRKEYGTDTILEDF 139
Query: 172 EFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMH 231
+F EL EV++Y P GYHGVDKEGRPV+IER K +KLM++TTIDRY+KYH Q E
Sbjct: 140 DFEELEEVLQYYPQGYHGVDKEGRPVYIERLGKAHPSKLMRITTIDRYLKYHVQEFERAL 199
Query: 232 AIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFI 291
KFPAC+IA+KR I S+ TILD+QG+G N ++ KI YP+T + +I
Sbjct: 200 QEKFPACSIAAKRRICSTTTILDVQGLGIKNFTPTAANLVAAMSKIDNSYYPETLHRMYI 259
Query: 292 INVGLELRS-LRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTC-AN 349
+N G + L + F+D K +K+HV+ + KL +VID+S+LP FLGG+C+C +
Sbjct: 260 VNAGTGFKKMLWPAAQKFLDAKTIAKIHVLEPKSLFKLHEVIDSSQLPEFLGGSCSCFGD 319
Query: 350 QGGCLRSDKGPWNNPEILKV--KGSDTLTAESSSEAED 385
GGCLRS+KGPWN+PEI+K+ G +L +S+ + D
Sbjct: 320 GGGCLRSNKGPWNDPEIMKLIYHGESSLFRQSTRKLTD 357
>At5g56160 putative protein
Length = 592
Score = 278 bits (712), Expect = 3e-75
Identities = 138/313 (44%), Positives = 199/313 (63%), Gaps = 11/313 (3%)
Query: 68 DENRKERRSDFKKKLLNAFSKFKHSFK----KKRVD-ELQAVNVV-----PAVVDAFRKL 117
+E R+ R + KKK + +K H K K+++D ++ + V +V R+
Sbjct: 34 EEPRRSRIGNLKKKAFSCSTKLTHPLKMRKGKRKIDFQIPLIEDVRDEKEEKLVSKLRQQ 93
Query: 118 LIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELN 177
L+ +LLP HDDYHM+LRFLK +F I K W +ML+WRKEFG D I++DF F EL+
Sbjct: 94 LLQKDLLPPVHDDYHMLLRFLKTMEFKIEKTVTAWEEMLKWRKEFGTDRIIQDFNFKELD 153
Query: 178 EVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEEMHAIKFPA 237
EV ++ P GYHGVDK+GRP++IER K KLM+VTTI+RY+KYH Q E K PA
Sbjct: 154 EVTRHYPQGYHGVDKDGRPIYIERLGKAHPGKLMEVTTIERYLKYHVQEFERTLQEKLPA 213
Query: 238 CTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLE 297
C++A+KR + ++ TILD++G+G N ++ K+ + YP+T + FI+N G+
Sbjct: 214 CSVAAKRRVTTTTTILDVEGLGMKNFTPTAANLLATIAKVDCNYYPETLHRMFIVNAGIG 273
Query: 298 LRS-LRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCANQGGCLRS 356
RS L + +DP +K+ V+ R KLL+ ID+S+LP FLGG C C N+GGCLRS
Sbjct: 274 FRSFLWPAAQKLLDPMTIAKIQVLEPRSLSKLLEAIDSSQLPEFLGGLCKCPNEGGCLRS 333
Query: 357 DKGPWNNPEILKV 369
+KGPWN+PEI+++
Sbjct: 334 NKGPWNDPEIVEL 346
>At5g47510 putative protein
Length = 403
Score = 246 bits (628), Expect = 2e-65
Identities = 116/278 (41%), Positives = 175/278 (62%)
Query: 110 VVDAFRKLLIMDELLPQKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIME 169
+V+AFR LL++ LP KH D++ + RFLK R FD+ K+K + + ++WR ++ D I +
Sbjct: 27 MVEAFRNLLLLHGHLPDKHGDHNTLRRFLKMRDFDLEKSKEAFLNYMKWRVDYKVDLISQ 86
Query: 170 DFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQRCEE 229
F+F E EV K+ PHG+H VDK GRP++IER D N ++ TTI+RYV YH + E+
Sbjct: 87 KFKFEEYGEVKKHYPHGFHKVDKTGRPIYIERLGMTDLNAFLKATTIERYVNYHIKEQEK 146
Query: 230 MHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQS 289
++++PAC+IAS +H+ S+ TILD+ G+G N + R + KI + YP+T +
Sbjct: 147 TMSLRYPACSIASDKHVSSTTTILDVSGVGMSNFSKPARSLFMEIQKIDSNYYPETLHRL 206
Query: 290 FIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTFLGGTCTCAN 349
F++N R L + F+D + +KV V+G Y +LL+ I+ S LPTFLGG CTC++
Sbjct: 207 FVVNASSGFRMLWLALKTFLDARTLAKVQVLGPNYLGELLEAIEPSNLPTFLGGNCTCSD 266
Query: 350 QGGCLRSDKGPWNNPEILKVKGSDTLTAESSSEAEDNV 387
GGCL SD+GPWN+P I + + ++ SE D V
Sbjct: 267 HGGCLFSDEGPWNDPGIKEKIEEPSTIEDAHSETMDKV 304
>At1g72160 cytosolic factor, putative
Length = 490
Score = 71.2 bits (173), Expect = 1e-12
Identities = 69/268 (25%), Positives = 121/268 (44%), Gaps = 25/268 (9%)
Query: 94 KKKRVDELQAVNVVPAVVDAFRKLLIMDEL----LPQKHDDYH--MMLRFLKARKFDIGK 147
+KK +DEL+ ++V +D + +E+ +P DD ++L+FL+AR+F +
Sbjct: 123 EKKSLDELK--HLVREALDNHQFTNTPEEVKIWGIPLLEDDRSDVVLLKFLRAREFKVKD 180
Query: 148 AKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPHGYHGVDKEGRPVFIERFEKLDR 207
+ M + ++WRKEF D ++E+ ++L++V+ HG D+EG PV + +
Sbjct: 181 SFAMLKNTIKWRKEFKIDELVEEDLVDDLDKVV-----FMHGHDREGHPVCYNVYGEFQN 235
Query: 208 NKLMQVTTIDRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQ---GIGFCNLE 264
+L T D + H R + S + + + D++ G+G L
Sbjct: 236 KELYNKTFSDEEKRKHFLRTRIQFLERSIRKLDFSSGGVSTIFQVNDMKNSPGLGKKEL- 294
Query: 265 EADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIG-DR 323
R K+ +++L DNYP+ + INV ++ FM P+ SK+ G R
Sbjct: 295 ---RSATKQAVELLQDNYPEFVFKQAFINVPWWYLVFYTVIGPFMTPRSKSKLVFAGPSR 351
Query: 324 YQRKLLKVIDASELPTFLGG----TCTC 347
L K I ++P GG C C
Sbjct: 352 SAETLFKYISPEQVPVQYGGLSVDPCDC 379
>At5g47730 putative protein
Length = 341
Score = 68.9 bits (167), Expect = 5e-12
Identities = 60/244 (24%), Positives = 113/244 (45%), Gaps = 25/244 (10%)
Query: 111 VDAFRKLLI-MDELLPQKHDDYHM------MLRFLKARKFDIGKAKHMWADMLEWRKEFG 163
+D F++L+ ++E L + ++ H + RFLKAR +++ KA M + L WR +
Sbjct: 9 IDEFQELMDQVEEPLKKTYERVHQGYLRENLGRFLKARDWNVCKAHTMLVECLRWRVDNE 68
Query: 164 ADTIM-EDFEFNELNEVIKYNPH-GYHGVDKEGRPVF-----IERFEKLDRNKLMQVTTI 216
D+I+ + EL ++ + G G KEG PVF + F+K ++
Sbjct: 69 IDSILSKPIVPTELYRDVRDSQLIGMSGYTKEGLPVFAIGVGLSTFDK---------ASV 119
Query: 217 DRYVKYHAQRCEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLK 276
YV+ H Q E + P+ + + R I + + +LD+ G+ L + + +
Sbjct: 120 HYYVQSHIQINEYRDRVLLPSISKKNGRPITTCVKVLDMTGLKLSALSQIKLVTIISTID 179
Query: 277 ILIDNYPQTGGQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASE 336
L NYP+ +++N + + + + + KVHV+ + +LLK++D +
Sbjct: 180 DL--NYPEKTNTYYVVNAPYIFSACWKVVKPLLQERTRKKVHVLSGCGRDELLKIMDFTS 237
Query: 337 LPTF 340
LP F
Sbjct: 238 LPHF 241
>At4g09160 putative protein
Length = 723
Score = 64.3 bits (155), Expect = 1e-10
Identities = 72/293 (24%), Positives = 122/293 (41%), Gaps = 38/293 (12%)
Query: 66 GSDENRKERRSDFKKKLLNAFSKFKHSFKKKRVDELQAVNVVPAVVDAFRKLLIMDELLP 125
GS + + SD + LNA + +H + + ++ VP LL
Sbjct: 289 GSFKEETNKISDLSETELNALQELRHLLQVSQDSSKTSIWGVP--------------LLK 334
Query: 126 QKHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEFNELNEVIKYNPH 185
D ++L+FL+AR F +A M L+WR +F + ++++ ++L++V+
Sbjct: 335 DDRTDV-VLLKFLRARDFKPQEAYSMLNKTLQWRIDFNIEELLDENLGDDLDKVV----- 388
Query: 186 GYHGVDKEGRPVFIERFEKLDRNKLMQVTTID-----RYVKYHAQRCEE-MHAIKFPACT 239
G DKE PV + + L Q T D R++++ Q E+ + + F A
Sbjct: 389 FMQGQDKENHPVCYNVYGEFQNKDLYQKTFSDEEKRERFLRWRIQFLEKSIRNLDFVAGG 448
Query: 240 IASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELR 299
+++ ++ + + G G L R K+ L +L DNYP+ + INV
Sbjct: 449 VSTICQVND---LKNSPGPGKTEL----RLATKQALHLLQDNYPEFVSKQIFINVPWWYL 501
Query: 300 SLRSICEYFMDPKVASKVHVIG-DRYQRKLLKVIDASELPTFLGG----TCTC 347
+ I FM + SK+ G R LLK I +P GG C C
Sbjct: 502 AFYRIISPFMSQRSKSKLVFAGPSRSAETLLKYISPEHVPVQYGGLSVDNCEC 554
>At1g55840 polyphosphoinositide binding protein, putative
Length = 325
Score = 62.8 bits (151), Expect = 3e-10
Identities = 56/234 (23%), Positives = 99/234 (41%), Gaps = 24/234 (10%)
Query: 120 MDELLPQKHDDYHM------MLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMED--F 171
+D+ L + + + H +LRFLKAR ++ KA M + LEWR + D I+
Sbjct: 19 VDDSLRESYRNIHQGYPTENLLRFLKARDGNVQKAHKMLLECLEWRTQNEIDKILTKPIV 78
Query: 172 EFNELNEVIKYNPHGYHGVDKEGRPVF-----IERFEKLDRNKLMQVTTIDRYVKYHAQR 226
+ + G G KEG PV + ++K ++ YV+ H Q
Sbjct: 79 PVDLYRGIRDTQLVGVSGYSKEGLPVIAIGVGLSTYDK---------ASVHYYVQSHIQM 129
Query: 227 CEEMHAIKFPACTIASKRHIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTG 286
E + P+ + R I + + ILD+ G+ L + ++M I NYP+
Sbjct: 130 NEYRDRVVLPSASKKQGRPICTCLKILDMSGLKLSALSQI--KLMTAITTIDDLNYPEKT 187
Query: 287 GQSFIINVGLELRSLRSICEYFMDPKVASKVHVIGDRYQRKLLKVIDASELPTF 340
+++NV + + + + K+ V+ + +LLK++D LP F
Sbjct: 188 ETYYVVNVPYIFSACWKTIKPLLQERTKKKIQVLKGCGKDELLKIMDYESLPHF 241
>At3g51670 unknown protein
Length = 409
Score = 61.6 bits (148), Expect = 8e-10
Identities = 59/254 (23%), Positives = 104/254 (40%), Gaps = 28/254 (11%)
Query: 133 MMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIM-EDFEFNELNEVIKYNPHGYHGVD 191
++L+FL+AR F + + M LEWR+EF A+ + ED F +L + Y G D
Sbjct: 84 ILLKFLRARDFKVADSLRMLEKCLEWREEFKAEKLTEEDLGFKDLEGKVAY----MRGYD 139
Query: 192 KEGRPVFIERFEKLDRNKLMQ-----VTTIDRYVKYHAQRCEE-MHAIKFPACTIASKRH 245
KEG PV + ++ + +++++++ Q E + + F
Sbjct: 140 KEGHPVCYNAYGVFKEKEMYERVFGDEEKLNKFLRWRVQVLERGVKMLHF------KPGG 193
Query: 246 IDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSIC 305
++S I + DL+ + L A +I+ F DNYP+ INV + S+
Sbjct: 194 VNSIIQVTDLKDMPKRELRVASNQILSLFQ----DNYPELVATKIFINVPWYFSVIYSMF 249
Query: 306 EYFMDPKVASKVHVIGD-RYQRKLLKVIDASELPTFLGGTCTCANQGGCLRSDKGPWNNP 364
F+ + SK + + L K I ++P GG + S GP
Sbjct: 250 SPFLTQRTKSKFVMSKEGNAAETLYKFIRPEDIPVQYGGLSRPTD------SQNGPPKPA 303
Query: 365 EILKVKGSDTLTAE 378
+KG + + +
Sbjct: 304 SEFSIKGGEKVNIQ 317
>At1g01630 polyphosphoinositide binding protein, putative
Length = 255
Score = 57.4 bits (137), Expect = 1e-08
Identities = 61/220 (27%), Positives = 94/220 (42%), Gaps = 23/220 (10%)
Query: 127 KHDDYHMMLRFLKARKFDIGKAKHMWADMLEWRKEFGADTIMEDFEF-NELNEVIKYNPH 185
K D M+ RFL+AR DI KA M+ + L W++ + + E N+L+ +N
Sbjct: 46 KEVDDLMIRRFLRARDLDIEKASTMFLNYLTWKRSMLPKGHIPEAEIANDLS----HNKM 101
Query: 186 GYHGVDKEGRPVFIERFEKLDRNKLMQVTTIDRYVKYHAQR-CEEMHAIKFPACTIASKR 244
G DK GRP+ + + + +K R+V Y ++ C M R
Sbjct: 102 CMQGHDKMGRPIAVAIGNRHNPSK-GNPDEFKRFVVYTLEKICARM------------PR 148
Query: 245 HIDSSITILDLQGIGFCNLEEADREIMKRFLKILIDNYPQTGGQSFIINVGLELRSLRSI 304
+ + I DLQG G+ N D L L D YP+ G+ +I++ + +
Sbjct: 149 GQEKFVAIGDLQGWGYSN---CDIRGYLAALSTLQDCYPERLGKLYIVHAPYIFMTAWKV 205
Query: 305 CEYFMDPKVASK-VHVIGDRYQRKLLKVIDASELPTFLGG 343
F+D K V V + LL+ ID S+LP GG
Sbjct: 206 IYPFIDANTKKKIVFVENKKLTPTLLEDIDESQLPDIYGG 245
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.324 0.140 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,251,108
Number of Sequences: 26719
Number of extensions: 395107
Number of successful extensions: 1333
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 36
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1253
Number of HSP's gapped (non-prelim): 53
length of query: 418
length of database: 11,318,596
effective HSP length: 102
effective length of query: 316
effective length of database: 8,593,258
effective search space: 2715469528
effective search space used: 2715469528
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC121244.7


