

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
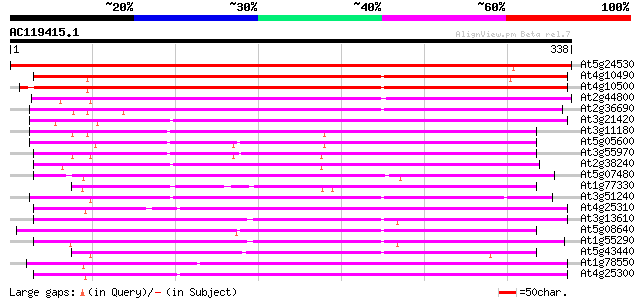
Query= AC119415.1 + phase: 0
(338 letters)
Database: ara_mips
26,719 sequences; 11,318,596 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
At5g24530 flavanone 3-hydroxylase-like protein 496 e-141
At4g10490 putative flavanone 3-beta-hydroxylase 380 e-106
At4g10500 putative Fe(II)/ascorbate oxidase 378 e-105
At2g44800 putative flavonol synthase 241 4e-64
At2g36690 putative giberellin beta-hydroxylase 239 1e-63
At3g21420 unknown protein 229 1e-60
At3g11180 putative leucoanthocyanidin dioxygenase 226 1e-59
At5g05600 leucoanthocyanidin dioxygenase-like protein 224 4e-59
At3g55970 leucoanthocyanidin dioxygenase -like protein 224 5e-59
At2g38240 putative anthocyanidin synthase 221 3e-58
At5g07480 putative protein 218 4e-57
At1g77330 unknown protein 216 2e-56
At3g51240 flavanone 3-hydroxylase (FH3) 214 5e-56
At4g25310 SRG1-like protein 208 4e-54
At3g13610 unknown protein 206 1e-53
At5g08640 flavonol synthase (FLS) (sp|Q96330) 202 3e-52
At1g55290 leucoanthocyanidin dioxygenase 2, putative 200 1e-51
At5g43440 1-aminocyclopropane-1-carboxylate oxidase 199 2e-51
At1g78550 unknown protein 198 3e-51
At4g25300 SRG1-like protein 197 9e-51
>At5g24530 flavanone 3-hydroxylase-like protein
Length = 341
Score = 496 bits (1277), Expect = e-141
Identities = 233/341 (68%), Positives = 280/341 (81%), Gaps = 3/341 (0%)
Query: 1 MDTKVLSSGIHYSKLPESYIRPESDRPCLSQVSEFENVPIIDLGSHNRTQIVQQIGEACS 60
M K++S+G ++ LPE+Y+RP SDRP LS+VS+ E+ P+IDL S +R+ ++QQI +AC+
Sbjct: 1 MAAKLISTGFRHTTLPENYVRPISDRPRLSEVSQLEDFPLIDLSSTDRSFLIQQIHQACA 60
Query: 61 SYGFFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEV 120
+GFFQV+NHGV + + + VA +FF + +EEKMKLYSDDPTKT RLSTSFNV KEEV
Sbjct: 61 RFGFFQVINHGVNKQIIDEMVSVAREFFSMSMEEKMKLYSDDPTKTTRLSTSFNVKKEEV 120
Query: 121 HNWRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDY 180
+NWRDYLRLHCYP+ YV EWPSNPPSFKE V+ Y +EVRE+G +IEE ISESLGLEKDY
Sbjct: 121 NNWRDYLRLHCYPIHKYVNEWPSNPPSFKEIVSKYSREVREVGFKIEELISESLGLEKDY 180
Query: 181 LRNALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLA 240
++ LGEQGQHMAVNYYPPCP+PELTYGLP HTDPNALTILLQD V GLQ+L DG+W A
Sbjct: 181 MKKVLGEQGQHMAVNYYPPCPEPELTYGLPAHTDPNALTILLQDTTVCGLQILIDGQWFA 240
Query: 241 INPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL 300
+NP PDAFVINIGDQLQALSNG+YKSVWHRA+ N E PRLSVASFLCP + A++ PAKPL
Sbjct: 241 VNPHPDAFVINIGDQLQALSNGVYKSVWHRAVTNTENPRLSVASFLCPADCAVMSPAKPL 300
Query: 301 TE---DGSGAVYRGFTYPEYYSKFWSRDLEKEHCLEFFKNN 338
E D + VY+ FTY EYY KFWSR+L++EHCLE F NN
Sbjct: 301 WEAEDDETKPVYKDFTYAEYYKKFWSRNLDQEHCLENFLNN 341
>At4g10490 putative flavanone 3-beta-hydroxylase
Length = 348
Score = 380 bits (977), Expect = e-106
Identities = 179/328 (54%), Positives = 237/328 (71%), Gaps = 7/328 (2%)
Query: 15 LPESYIRPESDRPCLSQV-SEFENVPIIDLGS---HNRTQIVQQIGEACSSYGFFQVVNH 70
+P +Y+RP SDRP +S+V + +++P+IDL NR I+ Q ACSS GFFQ+ NH
Sbjct: 18 VPSNYVRPVSDRPKMSEVQTSGDSIPLIDLHDLHGPNRADIINQFAHACSSCGFFQIKNH 77
Query: 71 GVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRLH 130
GVP E +KK A +FF+ E++K YS D KT RLSTSFNV+KE+V NWRD+LRLH
Sbjct: 78 GVPEETIKKMMNAAREFFRQSESERVKHYSADTKKTTRLSTSFNVSKEKVSNWRDFLRLH 137
Query: 131 CYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQGQ 190
CYP+++++ EWPS P SF+E A Y VR L L + E ISESLGL KD + N +G+ GQ
Sbjct: 138 CYPIEDFINEWPSTPISFREVTAEYATSVRALVLTLLEAISESLGLAKDRVSNTIGKHGQ 197
Query: 191 HMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPIPDAFVI 250
HMA+NYYP CPQPELTYGLPGH D N +T+LLQD V+GLQV KDGKW+A+NP+P+ F++
Sbjct: 198 HMAINYYPRCPQPELTYGLPGHKDANLITVLLQD-EVSGLQVFKDGKWIAVNPVPNTFIV 256
Query: 251 NIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL--TEDGSGAV 308
N+GDQ+Q +SN YKSV HRA+VN++ R+S+ +F CP +A+I PA+ L E+ S A+
Sbjct: 257 NLGDQMQVISNEKYKSVLHRAVVNSDMERISIPTFYCPSEDAVISPAQELINEEEDSPAI 316
Query: 309 YRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
YR FTY EY+ KFW + E C++ FK
Sbjct: 317 YRNFTYAEYFEKFWDTAFDTESCIDSFK 344
>At4g10500 putative Fe(II)/ascorbate oxidase
Length = 349
Score = 378 bits (971), Expect = e-105
Identities = 185/335 (55%), Positives = 240/335 (71%), Gaps = 9/335 (2%)
Query: 7 SSGIHYSKLPESYIRPESDRPCLSQV-SEFENVPIIDLGS---HNRTQIVQQIGEACSSY 62
+S +H +P +Y+RP SDRP LS+V S +++P+IDL NR IVQQ+ ACS+Y
Sbjct: 15 ASSVH---IPSNYVRPISDRPNLSEVESSGDSIPLIDLRDLHGPNRAVIVQQLASACSTY 71
Query: 63 GFFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHN 122
GFFQ+ NHGVP + K VA +FF P E++K YS DPTKT RLSTSFNV ++V N
Sbjct: 72 GFFQIKNHGVPDTTVNKMQTVAREFFHQPESERVKHYSADPTKTTRLSTSFNVGADKVLN 131
Query: 123 WRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLR 182
WRD+LRLHC+P+++++ EWPS+P SF+E A Y VR L LR+ E ISESLGLE D++
Sbjct: 132 WRDFLRLHCFPIEDFIEEWPSSPISFREVTAEYATSVRALVLRLLEAISESLGLESDHIS 191
Query: 183 NALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAIN 242
N LG+ QHMA NYYPPCP+PELTYGLPGH DP +T+LLQD V+GLQV KD KW+A++
Sbjct: 192 NILGKHAQHMAFNYYPPCPEPELTYGLPGHKDPTVITVLLQD-QVSGLQVFKDDKWVAVS 250
Query: 243 PIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL-T 301
PIP+ F++NIGDQ+Q +SN YKSV HRA+VN E RLS+ +F P +A+I PA L
Sbjct: 251 PIPNTFIVNIGDQMQVISNDKYKSVLHRAVVNTENERLSIPTFYFPSTDAVIGPAHELVN 310
Query: 302 EDGSGAVYRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
E S A+YR + + EY+ KFW+R L CL+ FK
Sbjct: 311 EQDSLAIYRTYPFVEYWDKFWNRSLATASCLDAFK 345
>At2g44800 putative flavonol synthase
Length = 352
Score = 241 bits (615), Expect = 4e-64
Identities = 129/332 (38%), Positives = 192/332 (56%), Gaps = 9/332 (2%)
Query: 14 KLPESYIRPESDRPCL--SQVSEFENVPIIDLGSHN----RTQIVQQIGEACSSYGFFQV 67
++P+ Y+ P S RP L S + +P+IDL + R+ + +I AC +GFFQV
Sbjct: 21 QVPDRYVLPPSQRPALGSSLGTSETTLPVIDLSLLHQPFLRSLAIHEISMACKEFGFFQV 80
Query: 68 VNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYL 127
+NHG+P + + A FF LPVEEKM L S + + +R TS N + + VH WRD++
Sbjct: 81 INHGIPSSVVNDALDAATQFFDLPVEEKMLLVSANVHEPVRYGTSLNHSTDRVHYWRDFI 140
Query: 128 RLHCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGE 187
+ + +PL ++ WPSNPP +K+ V Y + L ++ E ISESLGLEK+YL+ + E
Sbjct: 141 KHYSHPLSKWIDMWPSNPPCYKDKVGKYAEATHLLHKQLIEAISESLGLEKNYLQEEIEE 200
Query: 188 QGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGK-WLAINPIPD 246
Q MAVN YP CP+PE+ G+P H+D ++LTILLQ GLQ++ K W+ + I
Sbjct: 201 GSQVMAVNCYPACPEPEMALGMPPHSDFSSLTILLQS--SKGLQIMDCNKNWVCVPYIEG 258
Query: 247 AFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSG 306
A ++ +GDQ++ +SNG+YKSV HR VN E RLS AS I PA L +
Sbjct: 259 ALIVQLGDQVEVMSNGIYKSVIHRVTVNKEVKRLSFASLHSLPLHKKISPAPKLVNPNNA 318
Query: 307 AVYRGFTYPEYYSKFWSRDLEKEHCLEFFKNN 338
Y F++ ++ + S D +E ++ K +
Sbjct: 319 PAYGEFSFNDFLNYISSNDFIQERFIDTIKKS 350
>At2g36690 putative giberellin beta-hydroxylase
Length = 392
Score = 239 bits (611), Expect = 1e-63
Identities = 135/357 (37%), Positives = 199/357 (54%), Gaps = 37/357 (10%)
Query: 13 SKLPESYIRPESDRPCLSQVSEFEN------VPIIDLGS---HNRTQIVQQIGEACSSYG 63
+K+P YI PE DRP L++ + +P+ID NR +++ I EAC +YG
Sbjct: 30 TKVPTKYIWPEPDRPILTKSDKLIKPNKNLKLPLIDFAELLGPNRPHVLRTIAEACKTYG 89
Query: 64 FFQV--------------------------VNHGVPLEELKKTAEVAYDFFKLPVEEKMK 97
FFQV VNHG+ + K +V FF+LP EE+ K
Sbjct: 90 FFQVGSLNPKAHAVISRFLAKQVSYGNGQVVNHGMEGDVSKNMIDVCKRFFELPYEERSK 149
Query: 98 LYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRLHCYPLDNYVPEWPSNPPSFKETVANYCK 157
S D + +R TSFN K+ V WRD+L+L+ +PL +Y+P WPS+P F+ + A Y K
Sbjct: 150 YMSSDMSAPVRYGTSFNQIKDNVFCWRDFLKLYAHPLPDYLPHWPSSPSDFRSSAATYAK 209
Query: 158 EVRELGLRIEEYISESLGLE-KDYLRNALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPN 216
E +E+ + + I ESL ++ D L E Q + VN YPPCP+PELT G+P H+D
Sbjct: 210 ETKEMFEMMVKAILESLEIDGSDEAAKELEEGSQVVVVNCYPPCPEPELTLGMPPHSDYG 269
Query: 217 ALTILLQDLHVAGLQVLKDGKWLAINPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAE 276
LT+LLQD V GLQ+L +W+ ++PIP +FV+N+GD L+ SNG YKSV HR +VN+
Sbjct: 270 FLTLLLQD-EVEGLQILYRDEWVTVDPIPGSFVVNVGDHLEIFSNGRYKSVLHRVLVNST 328
Query: 277 KPRLSVASFLCPDNEALICPAKPLTEDGSGAVYRGFTYPEYYSKFWSRDLEKEHCLE 333
KPR+SVAS +++ P+ L + + + Y + + SR+ + ++ LE
Sbjct: 329 KPRISVASLHSFPLTSVVKPSPKLVDKHNPSQYMDTDFTTFLQYITSREPKWKNFLE 385
>At3g21420 unknown protein
Length = 364
Score = 229 bits (585), Expect = 1e-60
Identities = 126/337 (37%), Positives = 194/337 (57%), Gaps = 14/337 (4%)
Query: 13 SKLPESYIRPESDR----PCLSQVSEFENVPIIDLGSHNRTQI------VQQIGEACSSY 62
+K+PE +IR E +R L +P+IDL ++ + ++ +AC +
Sbjct: 26 NKVPERFIREEYERGVVVSSLKTHHLHHQIPVIDLSKLSKPDNDDFFFEILKLSQACEDW 85
Query: 63 GFFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHN 122
GFFQV+NHG+ +E ++ EVA +FF +P+EEK K Y +P +F ++++ +
Sbjct: 86 GFFQVINHGIEVEVVEDIEEVASEFFDMPLEEKKK-YPMEPGTVQGYGQAFIFSEDQKLD 144
Query: 123 WRDYLRLHCYPLDNYVPE-WPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYL 181
W + L +P P+ WPS P F E++ Y KE+REL R+ +YI+ SLGL+++
Sbjct: 145 WCNMFALGVHPPQIRNPKLWPSKPARFSESLEGYSKEIRELCKRLLKYIAISLGLKEERF 204
Query: 182 RNALGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLH-VAGLQVLKDGKWLA 240
GE Q + +NYYPPC P+L GL H+D +ALT+L Q + GLQ+LKD W+
Sbjct: 205 EEMFGEAVQAVRMNYYPPCSSPDLVLGLSPHSDGSALTVLQQSKNSCVGLQILKDNTWVP 264
Query: 241 INPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPL 300
+ P+P+A VINIGD ++ LSNG YKSV HRA+ N EK RL++ +F P+ E I P L
Sbjct: 265 VKPLPNALVINIGDTIEVLSNGKYKSVEHRAVTNREKERLTIVTFYAPNYEVEIEPMSEL 324
Query: 301 TEDGSG-AVYRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
+D + YR + + +Y + S L+ + L+F K
Sbjct: 325 VDDETNPCKYRSYNHGDYSYHYVSNKLQGKKSLDFAK 361
>At3g11180 putative leucoanthocyanidin dioxygenase
Length = 400
Score = 226 bits (577), Expect = 1e-59
Identities = 130/315 (41%), Positives = 176/315 (55%), Gaps = 11/315 (3%)
Query: 13 SKLPESYIRPESDRPCLSQVSEFE-----NVPIIDLGS--HNRTQIVQQIGEACSSYGFF 65
+ LP+ YI+P S RP + + N+PIIDL S ++I EAC +GFF
Sbjct: 65 TSLPDRYIKPPSQRPQTTIIDHQPEVADINIPIIDLDSLFSGNEDDKKRISEACREWGFF 124
Query: 66 QVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRD 125
QV+NHGV E + E FF LPVE K ++YS+ P + V K + +W D
Sbjct: 125 QVINHGVKPELMDAARETWKSFFNLPVEAK-EVYSNSPRTYEGYGSRLGVEKGAILDWND 183
Query: 126 YLRLHCYPLD-NYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNA 184
Y LH PL +WPS P + +E Y KE+ +LG R+ +S +LGL + L+ A
Sbjct: 184 YYYLHFLPLALKDFNKWPSLPSNIREMNDEYGKELVKLGGRLMTILSSNLGLRAEQLQEA 243
Query: 185 LGEQ--GQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAIN 242
G + G + VNYYP CPQPEL GL H+DP +TILL D V GLQV W+ +N
Sbjct: 244 FGGEDVGACLRVNYYPKCPQPELALGLSPHSDPGGMTILLPDDQVVGLQVRHGDTWITVN 303
Query: 243 PIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTE 302
P+ AF++NIGDQ+Q LSN YKSV HR IVN+EK R+S+A F P ++ I P + L
Sbjct: 304 PLRHAFIVNIGDQIQILSNSKYKSVEHRVIVNSEKERVSLAFFYNPKSDIPIQPMQQLVT 363
Query: 303 DGSGAVYRGFTYPEY 317
+Y T+ +Y
Sbjct: 364 STMPPLYPPMTFDQY 378
>At5g05600 leucoanthocyanidin dioxygenase-like protein
Length = 371
Score = 224 bits (572), Expect = 4e-59
Identities = 130/316 (41%), Positives = 181/316 (57%), Gaps = 13/316 (4%)
Query: 13 SKLPESYIRPESDRPCLSQ-VSEFENVPIIDLGSHNRTQ------IVQQIGEACSSYGFF 65
S LP+ YI+P S RP ++ N+PIIDL + I+ +I EAC +GFF
Sbjct: 36 SSLPDRYIKPASLRPTTTEDAPTATNIPIIDLEGLFSEEGLSDDVIMARISEACRGWGFF 95
Query: 66 QVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRD 125
QVVNHGV E + E +FF +PV K + YS+ P + V K +W D
Sbjct: 96 QVVNHGVKPELMDAARENWREFFHMPVNAK-ETYSNSPRTYEGYGSRLGVEKGASLDWSD 154
Query: 126 YLRLHCYP--LDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRN 183
Y LH P L ++ +WPS PP+ +E + Y +E+ +L RI +S +LGL++D +
Sbjct: 155 YYFLHLLPHHLKDF-NKWPSFPPTIREVIDEYGEELVKLSGRIMRVLSTNLGLKEDKFQE 213
Query: 184 ALGEQ--GQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAI 241
A G + G + VNYYP CP+PEL GL H+DP +TILL D V GLQV KD W+ +
Sbjct: 214 AFGGENIGACLRVNYYPKCPRPELALGLSPHSDPGGMTILLPDDQVFGLQVRKDDTWITV 273
Query: 242 NPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLT 301
P P AF++NIGDQ+Q LSN YKSV HR IVN++K R+S+A F P ++ I P + L
Sbjct: 274 KPHPHAFIVNIGDQIQILSNSTYKSVEHRVIVNSDKERVSLAFFYNPKSDIPIQPLQELV 333
Query: 302 EDGSGAVYRGFTYPEY 317
+ +Y T+ +Y
Sbjct: 334 STHNPPLYPPMTFDQY 349
>At3g55970 leucoanthocyanidin dioxygenase -like protein
Length = 363
Score = 224 bits (571), Expect = 5e-59
Identities = 129/319 (40%), Positives = 182/319 (56%), Gaps = 18/319 (5%)
Query: 15 LPESYIRPESDRPCLSQVSEFE----NVPIIDLGSHN------RTQIVQQIGEACSSYGF 64
+P Y++P S RP ++ ++ +PIIDLG + + + +I +AC GF
Sbjct: 25 IPNRYVKPLSQRPNITPHNKHNPQTTTIPIIDLGRLYTDDLTLQAKTLDEISKACRELGF 84
Query: 65 FQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWR 124
FQVVNHG+ + + + +FF LP+E K ++++ P + V K + +W
Sbjct: 85 FQVVNHGMSPQLMDQAKATWREFFNLPMELK-NMHANSPKTYEGYGSRLGVEKGAILDWS 143
Query: 125 DYLRLHCYP--LDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLR 182
DY LH P L +Y +WPS P +E + +YCKE+ +L + + +S++LGL++D L+
Sbjct: 144 DYYYLHYQPSSLKDYT-KWPSLPLHCREILEDYCKEMVKLCENLMKILSKNLGLQEDRLQ 202
Query: 183 NALG---EQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVL-KDGKW 238
NA G E G + VNYYP CPQPELT G+ H+DP LTILL D VA LQV D W
Sbjct: 203 NAFGGKEESGGCLRVNYYPKCPQPELTLGISPHSDPGGLTILLPDEQVASLQVRGSDDAW 262
Query: 239 LAINPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAK 298
+ + P P AF++N+GDQ+Q LSN +YKSV HR IVN E RLS+A F P I P K
Sbjct: 263 ITVEPAPHAFIVNMGDQIQMLSNSIYKSVEHRVIVNPENERLSLAFFYNPKGNVPIEPLK 322
Query: 299 PLTEDGSGAVYRGFTYPEY 317
L S A+Y TY Y
Sbjct: 323 ELVTVDSPALYSSTTYDRY 341
>At2g38240 putative anthocyanidin synthase
Length = 353
Score = 221 bits (564), Expect = 3e-58
Identities = 121/312 (38%), Positives = 181/312 (57%), Gaps = 8/312 (2%)
Query: 15 LPESYIRPESDRPCLS--QVSEFENVPIIDLGS-HNRTQIVQQIGEACSSYGFFQVVNHG 71
+P Y++P RP + Q +P++D+ + + ++ + AC +GFFQ+VNHG
Sbjct: 23 VPNRYVKPAHQRPVFNTTQSDAGIEIPVLDMNDVWGKPEGLRLVRSACEEWGFFQMVNHG 82
Query: 72 VPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRLHC 131
V +++ +FF+LP+EEK K Y++ P + V K+ +W DY L+
Sbjct: 83 VTHSLMERVRGAWREFFELPLEEKRK-YANSPDTYEGYGSRLGVVKDAKLDWSDYFFLNY 141
Query: 132 YPLDNYVP-EWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALG---E 187
P P +WPS PP +E + Y +EVR+L R+ E +SESLGL+ + L ALG +
Sbjct: 142 LPSSIRNPSKWPSQPPKIRELIEKYGEEVRKLCERLTETLSESLGLKPNKLMQALGGGDK 201
Query: 188 QGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPIPDA 247
G + N+YP CPQP+LT GL H+DP +TILL D VAGLQV + W+ I +P+A
Sbjct: 202 VGASLRTNFYPKCPQPQLTLGLSSHSDPGGITILLPDEKVAGLQVRRGDGWVTIKSVPNA 261
Query: 248 FVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSGA 307
++NIGDQLQ LSNG+YKSV H+ IVN+ R+S+A F P ++ + P + L A
Sbjct: 262 LIVNIGDQLQILSNGIYKSVEHQVIVNSGMERVSLAFFYNPRSDIPVGPIEELVTANRPA 321
Query: 308 VYRGFTYPEYYS 319
+Y+ + EY S
Sbjct: 322 LYKPIRFDEYRS 333
>At5g07480 putative protein
Length = 356
Score = 218 bits (555), Expect = 4e-57
Identities = 115/326 (35%), Positives = 182/326 (55%), Gaps = 17/326 (5%)
Query: 15 LPESYIRPESDRPCLSQVSEFENVPIIDL-----GSHNRTQIVQQIGEACSSYGFFQVVN 69
+P+ Y+ P S +PC S VP ID+ G R ++Q++ AC GFFQ+VN
Sbjct: 24 VPDCYVVPPSSKPCDSNSGI---VPTIDVSRLKGGDDERRGVIQELSLACQHLGFFQIVN 80
Query: 70 HGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRL 129
HG+ L EVA FF+LP +EK + S+D +R STS + + WR +L+
Sbjct: 81 HGINQNILDDALEVANSFFELPAKEKKQFMSNDVYAPVRYSTSLKDGLDTIQFWRIFLKH 140
Query: 130 HCYPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQG 189
+ +PL ++ WP NPP ++E + +C+EVR+L + + I+ESLGL +DYL + + E G
Sbjct: 141 YAHPLHRWIHLWPENPPGYREKMGKFCEEVRKLSIELMGAITESLGLGRDYLSSRMDENG 200
Query: 190 -QHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLK------DGKWLAIN 242
Q M VN YPPCP PE GLP H+D + +T+LLQ+L GL++ G+W+ +
Sbjct: 201 MQVMTVNCYPPCPDPETALGLPPHSDYSCITLLLQNLD--GLKIFDPMAHGGSGRWVGVP 258
Query: 243 PIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTE 302
+ ++IGD ++ LSNGLYKS+ H+ +N EK R+S+AS + + + L
Sbjct: 259 QVTGVLKVHIGDHVEVLSNGLYKSIVHKVTLNEEKTRISLASLHSLGMDDKMSVPRELVN 318
Query: 303 DGSGAVYRGFTYPEYYSKFWSRDLEK 328
D + Y+ ++ ++ D+ +
Sbjct: 319 DENPVRYKESSFNDFLDFLVKNDISQ 344
>At1g77330 unknown protein
Length = 307
Score = 216 bits (549), Expect = 2e-56
Identities = 121/292 (41%), Positives = 167/292 (56%), Gaps = 21/292 (7%)
Query: 38 VPIID---LGSHNRTQIVQQIGEACSSYGFFQVVNHGVPLEELKKTAEVAYDFFKLPVEE 94
+P+ID L R + + +I AC +GFFQ+VNHG+PLE L K +++ D +K EE
Sbjct: 3 IPVIDFSKLNGEEREKTLSEIARACEEWGFFQLVNHGIPLELLNKVKKLSSDCYKTEREE 62
Query: 95 KMKLYSDDPTKTMRLSTSFNVNKE-EVHNWRDYLRLHCYPLDNYVPEWPSNPPSFKETVA 153
K + +P K + N ++ E +W D L LD+ EWPSN KET+
Sbjct: 63 AFK--TSNPVKLLNELVQKNSGEKLENVDWEDVFTL----LDHNQNEWPSN---IKETMG 113
Query: 154 NYCKEVRELGLRIEEYISESLGLEKDYLRNALGE---QGQHMA-----VNYYPPCPQPEL 205
Y +EVR+L ++ E + E+LGL K Y++ A E G+ A V++YPPCP PEL
Sbjct: 114 EYREEVRKLASKMMEVMDENLGLPKGYIKKAFNEGMEDGEETAFFGTKVSHYPPCPHPEL 173
Query: 206 TYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPIPDAFVINIGDQLQALSNGLYK 265
GL HTD + +L QD GLQVLKDG+W+ + P+P+A VIN GDQ++ LSNG YK
Sbjct: 174 VNGLRAHTDAGGVVLLFQDDEYDGLQVLKDGEWIDVQPLPNAIVINTGDQIEVLSNGRYK 233
Query: 266 SVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSGAVYRGFTYPEY 317
S WHR + E R S+ASF P +A I PA E+GS Y F + +Y
Sbjct: 234 SAWHRVLAREEGNRRSIASFYNPSYKAAIGPAAVAEEEGSEKKYPKFVFGDY 285
>At3g51240 flavanone 3-hydroxylase (FH3)
Length = 358
Score = 214 bits (545), Expect = 5e-56
Identities = 121/323 (37%), Positives = 181/323 (55%), Gaps = 11/323 (3%)
Query: 13 SKLPESYIRPESDRPCLSQVSEFENVPIIDLGSHN-----RTQIVQQIGEACSSYGFFQV 67
SKL ++R E +RP ++ + +P+I L + R +I +QI EAC ++G FQV
Sbjct: 13 SKLNSKFVRDEDERPKVAYNVFSDEIPVISLAGIDDVDGKRGEICRQIVEACENWGIFQV 72
Query: 68 VNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYL 127
V+HGV + +A DFF LP E+K++ + K S ++ E V +WR+ +
Sbjct: 73 VDHGVDTNLVADMTRLARDFFALPPEDKLR-FDMSGGKKGGFIVSSHLQGEAVQDWREIV 131
Query: 128 RLHCYPLDNY-VPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALG 186
YP+ N WP P + + Y + + L ++ E +SE++GLEK+ L NA
Sbjct: 132 TYFSYPVRNRDYSRWPDKPEGWVKVTEEYSERLMSLACKLLEVLSEAMGLEKESLTNACV 191
Query: 187 EQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKD-GK-WLAINPI 244
+ Q + VNYYP CPQP+LT GL HTDP +T+LLQD V GLQ +D GK W+ + P+
Sbjct: 192 DMDQKIVVNYYPKCPQPDLTLGLKRHTDPGTITLLLQD-QVGGLQATRDNGKTWITVQPV 250
Query: 245 PDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDG 304
AFV+N+GD LSNG +K+ H+A+VN+ RLS+A+F P +A + P K + E
Sbjct: 251 EGAFVVNLGDHGHFLSNGRFKNADHQAVVNSNSSRLSIATFQNPAPDATVYPLK-VREGE 309
Query: 305 SGAVYRGFTYPEYYSKFWSRDLE 327
+ T+ E Y + RDLE
Sbjct: 310 KAILEEPITFAEMYKRKMGRDLE 332
>At4g25310 SRG1-like protein
Length = 353
Score = 208 bits (529), Expect = 4e-54
Identities = 114/328 (34%), Positives = 187/328 (56%), Gaps = 10/328 (3%)
Query: 15 LPESYIRPESDRPCLSQVSEFENVPIIDLG----SHNRTQIVQQIGEACSSYGFFQVVNH 70
LP Y+R + ++ + S +PIID+ S + + ++ AC +GFFQ+VNH
Sbjct: 29 LPPRYVRSDQEKGEAAIDSGENQIPIIDMSLLSSSTSMDSEIDKLDFACKEWGFFQLVNH 88
Query: 71 GVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRLH 130
G+ L++ K + DFF LP+EEK KL+ P +F ++E+ +W D L
Sbjct: 89 GMDLDKFKSDIQ---DFFNLPMEEKKKLWQQ-PGDIEGFGQAFVFSEEQKLDWADVFFLT 144
Query: 131 CYPLDNYVPE-WPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQ- 188
P+ P +P P F++T+ Y E++ + + ++ +L ++ + + ++
Sbjct: 145 MQPVPLRKPHLFPKLPLPFRDTLDTYSAELKSIAKVLFAKLASALKIKPEEMEKLFDDEL 204
Query: 189 GQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPIPDAF 248
GQ + +NYYPPCP+P+ GL H+D LTILLQ V GLQ+ KDGKW+++ P+P+A
Sbjct: 205 GQRIRMNYYPPCPEPDKAIGLTPHSDATGLTILLQVNEVEGLQIKKDGKWVSVKPLPNAL 264
Query: 249 VINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSGAV 308
V+N+GD L+ ++NG Y+S+ HR +VN+EK RLSVASF I P + L E GA+
Sbjct: 265 VVNVGDILEIITNGTYRSIEHRGVVNSEKERLSVASFHNTGFGKEIGPMRSLVERHKGAL 324
Query: 309 YRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
++ T EY+ +SR+L+ + L+ +
Sbjct: 325 FKTLTTEEYFHGLFSRELDGKAYLDVMR 352
>At3g13610 unknown protein
Length = 361
Score = 206 bits (524), Expect = 1e-53
Identities = 117/327 (35%), Positives = 182/327 (54%), Gaps = 9/327 (2%)
Query: 15 LPESYIRPESDRPCLSQVSEF-ENVPIIDLGSHNRTQIVQQIGEACSSYGFFQVVNHGVP 73
LPE YI+P +R V+E E +P+ID+ + + ++ + + +A +GFFQV+NHGVP
Sbjct: 38 LPEQYIQPLEERLINKFVNETDEAIPVIDMSNPDEDRVAEAVCDAAEKWGFFQVINHGVP 97
Query: 74 LEELKKTAEVAYDFFKLPVEEKMKLYSDDP-TKTMRLSTSFNVNKEEVHNWRDYLRLHCY 132
LE L + FF LPVEEK K ++ + T+R TSF+ E+ W+DYL L
Sbjct: 98 LEVLDDVKAATHKFFNLPVEEKRKFTKENSLSTTVRFGTSFSPLAEQALEWKDYLSLFFV 157
Query: 133 PLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGL-EKDYLRNALGEQGQH 191
WP + Y + +++ R+ EY+ ++L + E D + +L
Sbjct: 158 SEAEAEQFWPD---ICRNETLEYINKSKKMVRRLLEYLGKNLNVKELDETKESLFMGSIR 214
Query: 192 MAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQV--LKDGKWLAINPIPDAFV 249
+ +NYYP CP P+LT G+ H+D ++LTILLQD + GL V L G W+ + P+ +FV
Sbjct: 215 VNLNYYPICPNPDLTVGVGRHSDVSSLTILLQD-QIGGLHVRSLASGNWVHVPPVAGSFV 273
Query: 250 INIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSGAVY 309
INIGD +Q +SNGLYKSV HR + N R+SV F+ P E++I P + +G +Y
Sbjct: 274 INIGDAMQIMSNGLYKSVEHRVLANGYNNRISVPIFVNPKPESVIGPLPEVIANGEEPIY 333
Query: 310 RGFTYPEYYSKFWSRDLEKEHCLEFFK 336
R Y +Y F+ + + + +++ K
Sbjct: 334 RDVLYSDYVKYFFRKAHDGKKTVDYAK 360
>At5g08640 flavonol synthase (FLS) (sp|Q96330)
Length = 336
Score = 202 bits (513), Expect = 3e-52
Identities = 111/319 (34%), Positives = 176/319 (54%), Gaps = 8/319 (2%)
Query: 5 VLSSGIHYSKLPESYIRPESDRPCLSQV-SEFENVPIIDLGSHNRTQIVQQIGEACSSYG 63
+ SS + +P +IR E ++P ++ +P++DL + + + + +A +G
Sbjct: 9 ISSSSLLTEAIPLEFIRSEKEQPAITTFRGPTPAIPVVDLSDPDEESVRRAVVKASEEWG 68
Query: 64 FFQVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMR-LSTSFNVNKEEVHN 122
FQVVNHG+P E +++ +V FF+LP EK + + +K + T + E
Sbjct: 69 LFQVVNHGIPTELIRRLQDVGRKFFELPSSEKESVAKPEDSKDIEGYGTKLQKDPEGKKA 128
Query: 123 WRDYLRLHCYPLD--NYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDY 180
W D+L +P NY WP NPP ++E Y V++L + +S+ LGL++D
Sbjct: 129 WVDHLFHRIWPPSCVNY-RFWPKNPPEYREVNEEYAVHVKKLSETLLGILSDGLGLKRDA 187
Query: 181 LRNALG-EQGQHMA-VNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKW 238
L+ LG E ++M +NYYPPCP+P+L G+P HTD + +T+L+ + V GLQV KD W
Sbjct: 188 LKEGLGGEMAEYMMKINYYPPCPRPDLALGVPAHTDLSGITLLVPN-EVPGLQVFKDDHW 246
Query: 239 LAINPIPDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAK 298
IP A +++IGDQ+ LSNG YK+V HR V+ EK R+S FL P E ++ P
Sbjct: 247 FDAEYIPSAVIVHIGDQILRLSNGRYKNVLHRTTVDKEKTRMSWPVFLEPPREKIVGPLP 306
Query: 299 PLTEDGSGAVYRGFTYPEY 317
LT D + ++ F + +Y
Sbjct: 307 ELTGDDNPPKFKPFAFKDY 325
>At1g55290 leucoanthocyanidin dioxygenase 2, putative
Length = 361
Score = 200 bits (508), Expect = 1e-51
Identities = 115/326 (35%), Positives = 177/326 (54%), Gaps = 10/326 (3%)
Query: 15 LPESYIRPESDRPCLSQVSEF--ENVPIIDLGSHNRTQIVQQIGEACSSYGFFQVVNHGV 72
LP+ YI+P +R V E E++P+ID+ + + + + + +A +GFFQV+NHGV
Sbjct: 37 LPDQYIQPFEERLINFHVKEDSDESIPVIDISNLDEKSVSKAVCDAAEEWGFFQVINHGV 96
Query: 73 PLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKT-MRLSTSFNVNKEEVHNWRDYLRLHC 131
+E L+ + FF LPVEEK K + T +R TSF+ + E+ W+DYL L
Sbjct: 97 SMEVLENMKTATHRFFGLPVEEKRKFSREKSLSTNVRFGTSFSPHAEKALEWKDYLSLFF 156
Query: 132 YPLDNYVPEWPSNPPSFKETVANYCKEVRELGLRIEEYISESLGL-EKDYLRNALGEQGQ 190
WP S + Y E + L ++ ++ E+L + E D + +
Sbjct: 157 VSEAEASQLWPD---SCRSETLEYMNETKPLVKKLLRFLGENLNVKELDKTKESFFMGST 213
Query: 191 HMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQV--LKDGKWLAINPIPDAF 248
+ +NYYP CP PELT G+ H+D ++LTILLQD + GL V L G+W+ + PI +
Sbjct: 214 RINLNYYPICPNPELTVGVGRHSDVSSLTILLQD-EIGGLHVRSLTTGRWVHVPPISGSL 272
Query: 249 VINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSGAV 308
VINIGD +Q +SNG YKSV HR + N R+SV F+ P E++I P + E+G V
Sbjct: 273 VINIGDAMQIMSNGRYKSVEHRVLANGSYNRISVPIFVSPKPESVIGPLLEVIENGEKPV 332
Query: 309 YRGFTYPEYYSKFWSRDLEKEHCLEF 334
Y+ Y +Y F+ + + + ++F
Sbjct: 333 YKDILYTDYVKHFFRKAHDGKKTIDF 358
>At5g43440 1-aminocyclopropane-1-carboxylate oxidase
Length = 365
Score = 199 bits (505), Expect = 2e-51
Identities = 113/287 (39%), Positives = 162/287 (56%), Gaps = 10/287 (3%)
Query: 38 VPIIDLGSHN----RTQIVQQIGEACSSYGFFQVVNHGVPLEELKKTAEVAYDFFKLPVE 93
VPIIDLG N R +V +I EA ++GFFQV+NHG+PL LK + F + E
Sbjct: 62 VPIIDLGDGNTSAARNVLVSKIKEAAENWGFFQVINHGIPLTVLKDIKQGVRRFHEEDPE 121
Query: 94 EKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRLHCYPLDNYVPEWPSNPPSFKETVA 153
K + ++ D +T+F+++ NW+D + P D PE P + ++ V
Sbjct: 122 VKKQYFATDFNTRFAYNTNFDIHYSSPMNWKDSFTCYTCPQDPLKPE--EIPLACRDVVI 179
Query: 154 NYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQGQHMAVNYYPPCPQPELTYGLPGHT 213
Y K V ELG + + +SE+LGL+ + L+N +G M +YYPPCPQP+LT G+ HT
Sbjct: 180 EYSKHVMELGGLLFQLLSEALGLDSEILKNMDCLKGLLMLCHYYPPCPQPDLTLGISKHT 239
Query: 214 DPNALTILLQDLHVAGLQVLKDGKWLAINPIPDAFVINIGDQLQALSNGLYKSVWHRAIV 273
D + +TILLQD + GLQVL W+ + P+P A VI+IGD +Q ++N + S+ HR
Sbjct: 240 DNSFITILLQD-QIGGLQVLHQDSWVDVTPVPGALVISIGDFMQLITNDKFLSMEHRVRA 298
Query: 274 NAEKPRLSVASFLCP---DNEALICPAKPLTEDGSGAVYRGFTYPEY 317
N + PR+SVA F+ N + P K L D + A YR T PEY
Sbjct: 299 NRDGPRISVACFVSSGVFPNSTVYGPIKELLSDENPAKYRDITIPEY 345
>At1g78550 unknown protein
Length = 356
Score = 198 bits (504), Expect = 3e-51
Identities = 108/332 (32%), Positives = 183/332 (54%), Gaps = 7/332 (2%)
Query: 11 HYSKLPESYIRPESDRP-CLSQVSEFENVPIIDL----GSHNRTQIVQQIGEACSSYGFF 65
+++ +P Y+R + ++ L+ S +P+ID+ ++++ AC +GFF
Sbjct: 25 NFTTIPPRYVRVDQEKTEILNDSSLSSEIPVIDMTRLCSVSAMDSELKKLDFACQDWGFF 84
Query: 66 QVVNHGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRD 125
Q+VNHG+ L+K +FF LP++EK KL+ + V++ + +W D
Sbjct: 85 QLVNHGIDSSFLEKLETEVQEFFNLPMKEKQKLWQRSGEFEGFGQVNI-VSENQKLDWGD 143
Query: 126 YLRLHCYPLDNYVPEWPSN-PPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNA 184
L P+ + S PP F+ET+ Y EV+ + + ++ L ++ + + +
Sbjct: 144 MFILTTEPIRSRKSHLFSKLPPPFRETLETYSSEVKSIAKILFAKMASVLEIKHEEMEDL 203
Query: 185 LGEQGQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPI 244
+ Q + +NYYPPCPQP+ GL H+D LTILLQ V GLQ+ KDGKW+ + P+
Sbjct: 204 FDDVWQSIKINYYPPCPQPDQVMGLTQHSDAAGLTILLQVNQVEGLQIKKDGKWVVVKPL 263
Query: 245 PDAFVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDG 304
DA V+N+G+ L+ ++NG Y+S+ HRA+VN+EK RLSVA F P E +I PAK L +
Sbjct: 264 RDALVVNVGEILEIITNGRYRSIEHRAVVNSEKERLSVAMFHSPGKETIIRPAKSLVDRQ 323
Query: 305 SGAVYRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
+++ + EY+ F+++ L + L+ +
Sbjct: 324 KQCLFKSMSTQEYFDAFFTQKLNGKSHLDLMR 355
>At4g25300 SRG1-like protein
Length = 356
Score = 197 bits (500), Expect = 9e-51
Identities = 112/329 (34%), Positives = 182/329 (55%), Gaps = 8/329 (2%)
Query: 15 LPESYIRPESDRPCLSQVSEFEN-VPIIDLG----SHNRTQIVQQIGEACSSYGFFQVVN 69
+P Y+R + D ++ S N +PIID+ S + + ++ AC +GFFQ+VN
Sbjct: 28 VPPRYVRSDQDVAEIAVDSGLRNQIPIIDMSLLCSSTSMDSEIDKLDSACKEWGFFQLVN 87
Query: 70 HGVPLEELKKTAEVAYDFFKLPVEEKMKLYSDDPTKTMRLSTSFNVNKEEVHNWRDYLRL 129
HG+ L K DFF LP+EEK L+ P + F V++E+ +W D L
Sbjct: 88 HGMESSFLNKVKSEVQDFFNLPMEEKKNLWQQ-PDEIEGFGQVFVVSEEQKLDWADMFFL 146
Query: 130 HCYPLDNYVPE-WPSNPPSFKETVANYCKEVRELGLRIEEYISESLGLEKDYLRNALGEQ 188
P+ P +P P F++T+ Y EV+ + + I+ +L ++ + + ++
Sbjct: 147 TMQPVRLRKPHLFPKLPLPFRDTLDMYSAEVKSIAKILLGKIAVALKIKPEEMDKLFDDE 206
Query: 189 -GQHMAVNYYPPCPQPELTYGLPGHTDPNALTILLQDLHVAGLQVLKDGKWLAINPIPDA 247
GQ + +NYYP CP+P+ GL H+D LTILLQ V GLQ+ K+ KW+++ P+P+A
Sbjct: 207 LGQRIRLNYYPRCPEPDKVIGLTPHSDSTGLTILLQANEVEGLQIKKNAKWVSVKPLPNA 266
Query: 248 FVINIGDQLQALSNGLYKSVWHRAIVNAEKPRLSVASFLCPDNEALICPAKPLTEDGSGA 307
V+N+GD L+ ++NG Y+S+ HR +VN+EK RLSVA+F I P + L E A
Sbjct: 267 LVVNVGDILEIITNGTYRSIEHRGVVNSEKERLSVAAFHNIGLGKEIGPMRSLVERHKAA 326
Query: 308 VYRGFTYPEYYSKFWSRDLEKEHCLEFFK 336
++ T EY++ +SR+L+ + L+ +
Sbjct: 327 FFKSVTTEEYFNGLFSRELDGKAYLDVMR 355
Database: ara_mips
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,978,382
Number of sequences in database: 6832
Database: /data/blast2/ara_mips_chr2
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,737,135
Number of sequences in database: 4184
Database: /data/blast2/ara_mips_chr3
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,236,886
Number of sequences in database: 5377
Database: /data/blast2/ara_mips_chr4
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 1,748,816
Number of sequences in database: 4030
Database: /data/blast2/ara_mips_chr5
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 2,569,679
Number of sequences in database: 6098
Database: /data/blast2/ara_mips_chl
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 25,951
Number of sequences in database: 85
Database: /data/blast2/ara_mips_mit
Posted date: Jul 15, 2004 10:29 AM
Number of letters in database: 21,747
Number of sequences in database: 113
Lambda K H
0.317 0.137 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,610,852
Number of Sequences: 26719
Number of extensions: 392020
Number of successful extensions: 1226
Number of sequences better than 10.0: 110
Number of HSP's better than 10.0 without gapping: 102
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 887
Number of HSP's gapped (non-prelim): 125
length of query: 338
length of database: 11,318,596
effective HSP length: 100
effective length of query: 238
effective length of database: 8,646,696
effective search space: 2057913648
effective search space used: 2057913648
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC119415.1


