

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
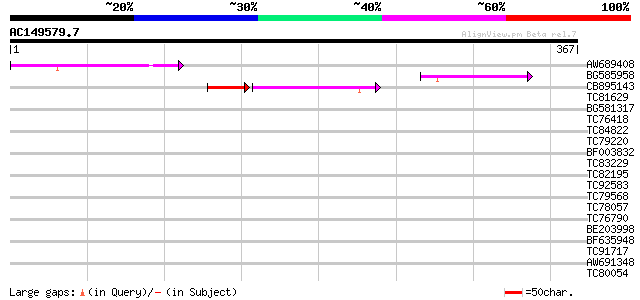
Query= AC149579.7 - phase: 0 /pseudo
(367 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW689408 130 1e-30
BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (... 66 2e-11
CB895143 62 4e-10
TC81629 similar to GP|15081654|gb|AAK82482.1 AT4g26750/F10M23_90... 39 0.004
BG581317 homologue to OMNI|NTL01EC003 IS2 hypothetical protein {... 38 0.005
TC76418 similar to GP|16226228|gb|AAL16109.1 At1g20110/T20H2_10 ... 38 0.005
TC84822 similar to GP|21554249|gb|AAM63324.1 unknown {Arabidopsi... 38 0.007
TC79220 similar to GP|2149570|gb|AAB58577.1| MAP kinase kinase p... 35 0.046
BF003832 similar to GP|21524577|emb unnamed protein product {Ara... 35 0.046
TC83229 35 0.060
TC82195 homologue to GP|19310475|gb|AAL84972.1 AT3g26744/MLJ15_1... 34 0.079
TC92583 similar to GP|7291175|gb|AAF46608.1| CG9817 gene product... 33 0.13
TC79568 weakly similar to GP|21689747|gb|AAM67517.1 unknown prot... 33 0.23
TC78057 weakly similar to PIR|T03962|T03962 r40g3 protein - rice... 33 0.23
TC76790 similar to PIR|T06817|T06817 RNA-binding protein - garde... 32 0.51
BE203998 similar to GP|10177034|dbj SCARECROW transcriptional re... 32 0.51
BF635948 similar to GP|21622312|emb putative protein {Neurospora... 31 0.67
TC91717 weakly similar to GP|6691125|gb|AAF24497.1| FH protein N... 31 0.87
AW691348 weakly similar to GP|12744989|gb| putative cyclin {Arab... 31 0.87
TC80054 SP|Q28009|FUS_BOVIN RNA-binding protein FUS (Pigpen prot... 30 1.5
>AW689408
Length = 583
Score = 130 bits (326), Expect = 1e-30
Identities = 70/135 (51%), Positives = 82/135 (59%), Gaps = 23/135 (17%)
Frame = +2
Query: 1 MDPNNNHFNNQNYSNYPFNYENPNNYPNP----------------------NQFFNQRPQ 38
MDPNNN FN QN S+Y F+Y+NPNNY +P NQF NQ PQ
Sbjct: 185 MDPNNNQFNTQNSSHYTFSYQNPNNYQHPNQFPNQHLLNPNQFSN*HPQNSNQFPNQHPQ 364
Query: 39 NIPNFGFPPNFNQSSSVP-NFHPYYGSMMKYPSQTPPFNGSMPMENDNFHNVGATQYPEF 97
N+ NFGF NFNQSSS P NF+PY+ SMM YPSQTP FNG M M N NF +VG +YP F
Sbjct: 365 NMSNFGFALNFNQSSSXPNNFNPYHRSMMGYPSQTPQFNGYMSMXNANFXSVG--EYPXF 538
Query: 98 STQITPSGMAIADEV 112
+ GM A+E+
Sbjct: 539 PXK*IGXGMTRANEI 583
>BG585958 weakly similar to GP|22830897|dbj ORF-B {Oryza sativa (japonica
cultivar-group)}, partial (13%)
Length = 769
Score = 66.2 bits (160), Expect = 2e-11
Identities = 37/75 (49%), Positives = 44/75 (58%), Gaps = 3/75 (4%)
Frame = +2
Query: 267 QIEEGSGSR---SRKYLKRDHAGANQRLIDDYFANEPTYDDAMFRRRYRMQKHVFLRIVG 323
Q E S SR R+ ++R+ +RL +DYF+ P Y D FRRRYRM KHVFLRIV
Sbjct: 440 QEEFASSSRPRSQRRNIERNREEGYKRLFNDYFSEAPVYMDE*FRRRYRMHKHVFLRIVE 619
Query: 324 DLSSSDNYFTQRVDA 338
L D YF VDA
Sbjct: 620 ALGQHDEYFQLTVDA 664
>CB895143
Length = 770
Score = 62.0 bits (149), Expect = 4e-10
Identities = 44/89 (49%), Positives = 53/89 (59%), Gaps = 6/89 (6%)
Frame = +1
Query: 158 R*AYARKRLVGR*CFDKSAGIVCRWEEYSIYFE*RMARSP*STTLW*SDGRKCWVRE*WI 217
R*AYA KR++ *C KS I+ +W E + +F RM P*STTL *S RK W+ +*WI
Sbjct: 430 R*AYATKRVIED*CIGKST*IIFKW*E*TFHFNIRMVCCP*STTL**SGRRKYWLWK*WI 609
Query: 218 *EISRGLC------RI*CSSNG*GGS*KK 240
*E + C RI* NG G S*KK
Sbjct: 610 *ESPQE*CK*FQLRRI*FLPNGEGCS*KK 696
Score = 51.2 bits (121), Expect = 6e-07
Identities = 21/27 (77%), Positives = 24/27 (88%)
Frame = +2
Query: 129 WNTQQNLVLISAWIKYGTSSVVGRNQR 155
WN QNL+LIS WIKYGT+SVVGRNQ+
Sbjct: 209 WNIDQNLILISGWIKYGTNSVVGRNQK 289
>TC81629 similar to GP|15081654|gb|AAK82482.1 AT4g26750/F10M23_90
{Arabidopsis thaliana}, partial (19%)
Length = 824
Score = 38.5 bits (88), Expect = 0.004
Identities = 32/97 (32%), Positives = 46/97 (46%), Gaps = 5/97 (5%)
Frame = +2
Query: 8 FNNQNYSNYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSS--SVPNFHPYY-GS 64
+N+Q YS PN YP+ P ++P+F P+F +SS SVP H YY G
Sbjct: 215 YNHQQYSPDQSQNLGPN-YPSHE---TPPPYSLPHFQSYPSFTESSLPSVPVNHTYYQGP 382
Query: 65 MMKYPSQTPPF--NGSMPMENDNFHNVGATQYPEFST 99
Y SQ+ P N S+ +N + G P+ +T
Sbjct: 383 DASYSSQSAPLTTNHSLNTQNSSSSRNGTVPEPKPTT 493
>BG581317 homologue to OMNI|NTL01EC003 IS2 hypothetical protein {Escherichia
coli K12-MG1655}, partial (55%)
Length = 808
Score = 38.1 bits (87), Expect = 0.005
Identities = 30/102 (29%), Positives = 44/102 (42%), Gaps = 9/102 (8%)
Frame = +2
Query: 8 FNNQNYSNYPFNYENPNNYPNPNQFFNQRPQN--IPNF-----GFPPNFNQSSSVPNFHP 60
+ + Y +P +NPN PNP PQ+ IPN F PN++ SS+ + P
Sbjct: 86 YTSAPYYQFPHMQQNPNPIPNPTP-----PQSDPIPNHYASAPPFTPNYDYSSTYSPYPP 250
Query: 61 YYGSMMKYPSQTPPFNGSMPMENDN--FHNVGATQYPEFSTQ 100
+ + PS P FN P E+ N + YP + Q
Sbjct: 251 HNPDHV--PSSNPSFN-PPPFESSNPLYQQPSQPYYPPYDQQ 367
>TC76418 similar to GP|16226228|gb|AAL16109.1 At1g20110/T20H2_10
{Arabidopsis thaliana}, partial (54%)
Length = 2020
Score = 38.1 bits (87), Expect = 0.005
Identities = 30/102 (29%), Positives = 44/102 (42%), Gaps = 9/102 (8%)
Frame = +3
Query: 8 FNNQNYSNYPFNYENPNNYPNPNQFFNQRPQN--IPNF-----GFPPNFNQSSSVPNFHP 60
+ + Y +P +NPN PNP PQ+ IPN F PN++ SS+ + P
Sbjct: 42 YTSAPYYQFPHMQQNPNPIPNPTP-----PQSDPIPNHYASAPPFTPNYDYSSTYSPYPP 206
Query: 61 YYGSMMKYPSQTPPFNGSMPMENDN--FHNVGATQYPEFSTQ 100
+ + PS P FN P E+ N + YP + Q
Sbjct: 207 HNPDHV--PSSNPSFN-PPPFESSNPLYQQPSQPYYPPYDQQ 323
>TC84822 similar to GP|21554249|gb|AAM63324.1 unknown {Arabidopsis
thaliana}, partial (47%)
Length = 636
Score = 37.7 bits (86), Expect = 0.007
Identities = 16/28 (57%), Positives = 22/28 (78%)
Frame = +1
Query: 244 ETYFQKRDAEDTYMVNRFIQRRKQIEEG 271
E Y +K + E+ Y++NRF +RRKQIEEG
Sbjct: 70 EAYKRKSEIEERYILNRFRERRKQIEEG 153
>TC79220 similar to GP|2149570|gb|AAB58577.1| MAP kinase kinase protein
DdMEK1 {Dictyostelium discoideum}, partial (6%)
Length = 1162
Score = 35.0 bits (79), Expect = 0.046
Identities = 23/81 (28%), Positives = 35/81 (42%), Gaps = 5/81 (6%)
Frame = +2
Query: 19 NYENPNNYPNPNQFFNQRPQNIPNFG---FPPNFNQSSSVPNFHPYYGSMMKYP--SQTP 73
N+E +NYPN N+++N N + F N N+ S Y SMM+ +Q
Sbjct: 302 NFEETSNYPNNNKYYNNDAYNTKYYNKDTFGNNQNELSDTKYNEEGYNSMMEKQNNNQEY 481
Query: 74 PFNGSMPMENDNFHNVGATQY 94
FN + +FH+ Y
Sbjct: 482 YFNNNNAANERSFHSNNYNNY 544
Score = 30.8 bits (68), Expect = 0.87
Identities = 16/32 (50%), Positives = 19/32 (59%)
Frame = +2
Query: 4 NNNHFNNQNYSNYPFNYENPNNYPNPNQFFNQ 35
NNN +NN N NY N E NN+ N N + NQ
Sbjct: 689 NNNWYNNNN--NYGNNNEYQNNHGNYNGYKNQ 778
Score = 28.5 bits (62), Expect = 4.3
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +2
Query: 4 NNNHFNNQNYSNYPFNYENPNNYPNPNQFFNQR 36
NNN+ NN Y N N+ N N Y N + N++
Sbjct: 710 NNNYGNNNEYQN---NHGNYNGYKNQEELKNEQ 799
>BF003832 similar to GP|21524577|emb unnamed protein product {Arabidopsis
thaliana}, partial (4%)
Length = 577
Score = 35.0 bits (79), Expect = 0.046
Identities = 25/88 (28%), Positives = 40/88 (45%)
Frame = +2
Query: 3 PNNNHFNNQNYSNYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYY 62
PNNNH Q +N P +++ NN P P Q+P N + SS V + +P +
Sbjct: 206 PNNNHHPFQQPNNNPQHFQQSNNNPQP----FQQPNN--------SIALSSQVNSANPQH 349
Query: 63 GSMMKYPSQTPPFNGSMPMENDNFHNVG 90
+M+Y + N + P++ VG
Sbjct: 350 QPVMQYTAD--QVNSNPPIQQHPAFGVG 427
>TC83229
Length = 692
Score = 34.7 bits (78), Expect = 0.060
Identities = 23/64 (35%), Positives = 32/64 (49%), Gaps = 12/64 (18%)
Frame = +1
Query: 3 PNNNHFNNQNYSN---YPFNYENPN----NYPNPNQFFNQRPQNIP-----NFGFPPNFN 50
P+ N YS+ YP NY NP +YP + Q P+ P N G+PPNF+
Sbjct: 4 PHKQAQTNVPYSHAVPYPTNYSNPYVGGPSYPAASGPSYQPPRTPPPPPTSNVGYPPNFH 183
Query: 51 QSSS 54
QS++
Sbjct: 184QSNN 195
>TC82195 homologue to GP|19310475|gb|AAL84972.1 AT3g26744/MLJ15_15
{Arabidopsis thaliana}, partial (17%)
Length = 1189
Score = 34.3 bits (77), Expect = 0.079
Identities = 45/170 (26%), Positives = 64/170 (37%), Gaps = 39/170 (22%)
Frame = +2
Query: 2 DPNNNHFNNQNYSN------YPFNY------------ENPNNY--PNPNQFFNQRPQNIP 41
+P+NN +NN N N P N+ NP+++ N N FFN N P
Sbjct: 302 EPDNNFYNNNNVVNVNNVIPLPDNFLMHQNTNTIDSISNPHSFFHNNNNYFFNNNNTNNP 481
Query: 42 -NFGFPPNF----NQSSSVPNFHPYYGSMMKYP-------SQTPPFNG--SMPME----- 82
GF F N +S+ P F S ++P + PPF+ SMP+E
Sbjct: 482 FEMGFENGFFMGNNNTSTSPVFMGGSLSASEFPPSLELDAAPVPPFSASFSMPLELAQPQ 661
Query: 83 NDNFHNVGATQYPEFSTQITPSGMAIADEVTPEDSTPKSKRSKEPAWNTQ 132
H+ Q P Q + I T + K KR E +W +
Sbjct: 662 PQQHHHQQQQQQPTTLFQKRRGALEIPRLETVGN---KKKRKVEKSWEEE 802
>TC92583 similar to GP|7291175|gb|AAF46608.1| CG9817 gene product
{Drosophila melanogaster}, partial (1%)
Length = 1015
Score = 33.5 bits (75), Expect = 0.13
Identities = 28/117 (23%), Positives = 50/117 (41%), Gaps = 16/117 (13%)
Frame = +2
Query: 25 NYPNPNQFFNQRPQNIPNFGFPPNFNQSSSV----------PNFHPYYGSMMKYPSQTPP 74
++P P+Q + P N+ + P +F SS+ P P +G P++TPP
Sbjct: 38 SHPIPSQATHPVPSNVHHSHEPVSFPFLSSLLPYAEQRTIHPPMLPTFGRRRYLPTRTPP 217
Query: 75 FNGSMPMENDNFHNVGATQYP--EFSTQITPSGMAIADEVTPEDST----PKSKRSK 125
+ + + H Q+P + S + P+ E+TP PKS++S+
Sbjct: 218 NASTSLEDTSSCHTTWLMQHP*ADTSGEALPATRTQNPEITPPPHPLPVHPKSRKSE 388
>TC79568 weakly similar to GP|21689747|gb|AAM67517.1 unknown protein
{Arabidopsis thaliana}, partial (25%)
Length = 1192
Score = 32.7 bits (73), Expect = 0.23
Identities = 33/120 (27%), Positives = 45/120 (37%), Gaps = 3/120 (2%)
Frame = +1
Query: 19 NYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKYPSQTPPFNGS 78
N N NN + N+ N PN P N Q + H GS + S P + S
Sbjct: 43 NQINNNNNKSKNKNKNNNTPTKPN-PKPKNHQQHNLNKPTHYLSGSTSLFQSLL*PSSSS 219
Query: 79 MPMENDNF--HNVGATQYPEFSTQITPSGMAIADEVTPEDSTPKSKRSKE-PAWNTQQNL 135
+ HN G++ YP S TP + TP ++ S S P + T Q L
Sbjct: 220 SLLHPFLLPHHNNGSSLYPPLSVNTTPMAAPSKSKFTPTNNPSTSSPSNSTPPFPTLQKL 399
>TC78057 weakly similar to PIR|T03962|T03962 r40g3 protein - rice, partial
(50%)
Length = 878
Score = 32.7 bits (73), Expect = 0.23
Identities = 29/108 (26%), Positives = 39/108 (35%), Gaps = 12/108 (11%)
Frame = +2
Query: 10 NQNYSNYPFNYE-------NPNNYPNPNQ----FFNQRPQNIPNFGFPPNFNQSSSVPN- 57
N N SN + N +YP P FNQ P PP+ Q PN
Sbjct: 5 NHNTSNVTHTHHHRRDDDNNEQHYPPPGHNNLSSFNQPPP-------PPHHQQPPFYPNP 163
Query: 58 FHPYYGSMMKYPSQTPPFNGSMPMENDNFHNVGATQYPEFSTQITPSG 105
+P + +T F+ NDNF+N P TQ+ +G
Sbjct: 164 SYPPPPQQQPHQPETQVFHTGHVSRNDNFNNYPQQPQPHQETQVFHTG 307
>TC76790 similar to PIR|T06817|T06817 RNA-binding protein - garden pea,
partial (79%)
Length = 1141
Score = 31.6 bits (70), Expect = 0.51
Identities = 22/67 (32%), Positives = 32/67 (46%), Gaps = 10/67 (14%)
Frame = +2
Query: 17 PFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFN----------QSSSVPNFHPYYGSMM 66
P + NP +PNP N+P+F FPPN + QS S+P PY S++
Sbjct: 53 PLHSSNP*PWPNPASSHPNPNPNLPSFPFPPNPSISISPSKPTAQSQSLP--LPY--SLL 220
Query: 67 KYPSQTP 73
K + +P
Sbjct: 221 KKATHSP 241
>BE203998 similar to GP|10177034|dbj SCARECROW transcriptional regulator-like
{Arabidopsis thaliana}, partial (6%)
Length = 594
Score = 31.6 bits (70), Expect = 0.51
Identities = 25/87 (28%), Positives = 33/87 (37%), Gaps = 8/87 (9%)
Frame = +1
Query: 7 HFNNQNYSNYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYYGSMM 66
H NN N N P ++ P Q P + P +QS+ PNFH Y S
Sbjct: 205 HANNNNTHNTPLSFSLPE--------LQQYPYHQTQRLGVPLLHQSTINPNFHHYRNSNS 360
Query: 67 KYPS--------QTPPFNGSMPMENDN 85
K QT P +G + N+N
Sbjct: 361 KLGQITNSIQTVQTIPVSGKVEENNNN 441
>BF635948 similar to GP|21622312|emb putative protein {Neurospora crassa},
partial (6%)
Length = 538
Score = 31.2 bits (69), Expect = 0.67
Identities = 21/49 (42%), Positives = 26/49 (52%), Gaps = 5/49 (10%)
Frame = +1
Query: 18 FNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVP-----NFHPY 61
FN NPN PNPN +Q P + P+ PP + SSS P N +PY
Sbjct: 160 FNPPNPNPNPNPN-ISSQFPSSAPS---PPPPSSSSSYPFPHNNNNYPY 294
>TC91717 weakly similar to GP|6691125|gb|AAF24497.1| FH protein NFH2
{Nicotiana tabacum}, partial (6%)
Length = 477
Score = 30.8 bits (68), Expect = 0.87
Identities = 30/91 (32%), Positives = 39/91 (41%)
Frame = +2
Query: 9 NNQNYSNYPFNYENPNNYPNPNQFFNQRPQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKY 68
NN + S+ P +P PNP F+ P PN + SSS P F P Y S
Sbjct: 179 NNLSPSSPP---SSPPPSPNPKYPFSTTP---------PNTSSSSSTPPFFPTYPSTPPP 322
Query: 69 PSQTPPFNGSMPMENDNFHNVGATQYPEFST 99
PS P S P N + + TQ P+ S+
Sbjct: 323 PS--PSSFASFP-ANISSLTIPQTQKPKSSS 406
>AW691348 weakly similar to GP|12744989|gb| putative cyclin {Arabidopsis
thaliana}, partial (9%)
Length = 618
Score = 30.8 bits (68), Expect = 0.87
Identities = 21/92 (22%), Positives = 38/92 (40%), Gaps = 17/92 (18%)
Frame = +2
Query: 4 NNNHFNNQNYSNYPFNYENPN----------------NYPN-PNQFFNQRPQNIPNFGFP 46
+ NH+N +NY+NY N+ N NY N + +R + P G
Sbjct: 197 SRNHYNQRNYNNYYDNHNQANFGYYGGNFNQCNADYSNYANVASSSLKKRKYSAPVRGES 376
Query: 47 PNFNQSSSVPNFHPYYGSMMKYPSQTPPFNGS 78
F ++V + P + YP+++ +N +
Sbjct: 377 QKFTLPATVYDSIPSSRNFQAYPARSIAYNST 472
>TC80054 SP|Q28009|FUS_BOVIN RNA-binding protein FUS (Pigpen protein).
[Bovine] {Bos taurus}, partial (4%)
Length = 988
Score = 30.0 bits (66), Expect = 1.5
Identities = 19/44 (43%), Positives = 20/44 (45%)
Frame = +1
Query: 37 PQNIPNFGFPPNFNQSSSVPNFHPYYGSMMKYPSQTPPFNGSMP 80
PQ G PP + QS S PN YG PSQ PP S P
Sbjct: 484 PQTQKPSGTPPAYGQSQS-PNAAGGYGQPGYPPSQPPPSGYSQP 612
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.333 0.143 0.475
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,797,999
Number of Sequences: 36976
Number of extensions: 191702
Number of successful extensions: 1878
Number of sequences better than 10.0: 81
Number of HSP's better than 10.0 without gapping: 1804
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1854
length of query: 367
length of database: 9,014,727
effective HSP length: 97
effective length of query: 270
effective length of database: 5,428,055
effective search space: 1465574850
effective search space used: 1465574850
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 59 (27.3 bits)
Medicago: description of AC149579.7


