

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
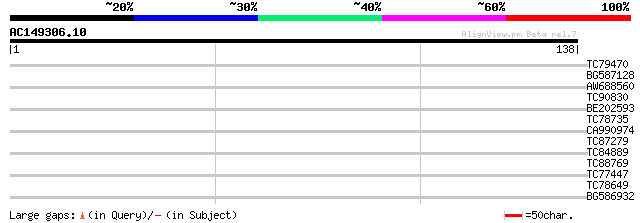
Query= AC149306.10 - phase: 0 /pseudo
(138 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC79470 similar to GP|15450980|gb|AAK96761.1 Unknown protein {Ar... 28 0.89
BG587128 28 1.2
AW688560 similar to PIR|T51529|T515 hypothetical protein T20K14_... 27 2.0
TC90830 27 2.6
BE202593 27 3.4
TC78735 similar to GP|10178184|dbj|BAB11658. 50S ribosomal prote... 27 3.4
CA990974 homologue to GP|23496034|gb|A hypothetical protein {Pla... 26 4.4
TC87279 weakly similar to GP|8843755|dbj|BAA97303.1 gene_id:MXK3... 26 4.4
TC84889 similar to GP|21537003|gb|AAM61344.1 unknown {Arabidopsi... 26 5.8
TC88769 homologue to GP|19881561|gb|AAM00962.1 hypothetical prot... 25 7.6
TC77447 similar to GP|16974501|gb|AAL31160.1 AT3g06720/F3E22_14 ... 25 7.6
TC78649 weakly similar to GP|6651029|gb|AAF22136.1| gamma-glutam... 25 9.9
BG586932 homologue to PIR|T52080|T52 multi resistance protein [i... 25 9.9
>TC79470 similar to GP|15450980|gb|AAK96761.1 Unknown protein {Arabidopsis
thaliana}, partial (47%)
Length = 1041
Score = 28.5 bits (62), Expect = 0.89
Identities = 21/60 (35%), Positives = 29/60 (48%)
Frame = +3
Query: 73 GSFEICPILSSGLLSFLFQGSYMYNLFP*IFFFFLNIRGVVKFLRKLLGSIGIQYVYLWI 132
GSF SS SF F S LFP I F + R +V + G+IG+ + +LW+
Sbjct: 210 GSFHSSTTSSS---SFPFSSSL---LFPSIHF*YQPPRSMVHLCFHVCGNIGVLHCFLWL 371
>BG587128
Length = 632
Score = 28.1 bits (61), Expect = 1.2
Identities = 13/31 (41%), Positives = 18/31 (57%)
Frame = +3
Query: 86 LSFLFQGSYMYNLFP*IFFFFLNIRGVVKFL 116
L +FQ +YN+ P FF F NI +K+L
Sbjct: 438 LMMIFQNPVIYNIHPRFFFCF*NIIRFLKYL 530
>AW688560 similar to PIR|T51529|T515 hypothetical protein T20K14_120 -
Arabidopsis thaliana, partial (8%)
Length = 601
Score = 27.3 bits (59), Expect = 2.0
Identities = 17/43 (39%), Positives = 23/43 (52%), Gaps = 3/43 (6%)
Frame = -3
Query: 88 FLFQGSYMYNL---FP*IFFFFLNIRGVVKFLRKLLGSIGIQY 127
FLF +Y+ N F +FFFF+ I ++K L L I I Y
Sbjct: 506 FLFFNNYIINFRNFFFMLFFFFIFINPILKKLFLTLSQIPISY 378
>TC90830
Length = 603
Score = 26.9 bits (58), Expect = 2.6
Identities = 14/24 (58%), Positives = 17/24 (70%)
Frame = -2
Query: 85 LLSFLFQGSYMYNLFP*IFFFFLN 108
L+ FLFQ + +F *IFFFFLN
Sbjct: 566 LIEFLFQNFF---IFL*IFFFFLN 504
>BE202593
Length = 557
Score = 26.6 bits (57), Expect = 3.4
Identities = 15/44 (34%), Positives = 22/44 (49%), Gaps = 1/44 (2%)
Frame = -1
Query: 65 ISIFWWPSGSFEIC-PILSSGLLSFLFQGSYMYNLFP*IFFFFL 107
+ I WWPS FE C +++ F S++ + *I FFL
Sbjct: 506 VCILWWPSLIFESCLQVVNGWWYKVAFVRSFLMHKVY*INTFFL 375
>TC78735 similar to GP|10178184|dbj|BAB11658. 50S ribosomal protein L29
{Arabidopsis thaliana}, partial (68%)
Length = 860
Score = 26.6 bits (57), Expect = 3.4
Identities = 23/89 (25%), Positives = 42/89 (46%)
Frame = +3
Query: 47 LETCC*KNCC*IVVVEP*ISIFWWPSGSFEICPILSSGLLSFLFQGSYMYNLFP*IFFFF 106
+E C* NCC* + + IFWW S S I +L + GS + +F ++
Sbjct: 594 IELHC*-NCC*SLFL-----IFWWMSSSRCI------NILFVVTYGSSCHKVFYVVYSVT 737
Query: 107 LNIRGVVKFLRKLLGSIGIQYVYLWIVVV 135
+ I + L+ +GI +++ +I ++
Sbjct: 738 VRIETWLSKLQFTAFKLGIHFLFCFIRIL 824
>CA990974 homologue to GP|23496034|gb|A hypothetical protein {Plasmodium
falciparum 3D7}, partial (36%)
Length = 564
Score = 26.2 bits (56), Expect = 4.4
Identities = 7/13 (53%), Positives = 10/13 (76%)
Frame = -3
Query: 27 YYSVCLFGVAYWW 39
Y+ +C+FGV WW
Sbjct: 562 YHDICIFGVP*WW 524
>TC87279 weakly similar to GP|8843755|dbj|BAA97303.1 gene_id:MXK3.14~unknown
protein {Arabidopsis thaliana}, partial (39%)
Length = 1468
Score = 26.2 bits (56), Expect = 4.4
Identities = 19/51 (37%), Positives = 25/51 (48%), Gaps = 2/51 (3%)
Frame = -3
Query: 72 SGSFEICPILSSGLLSF--LFQGSYMYNLFP*IFFFFLNIRGVVKFLRKLL 120
+G F P LS G L F LF ++ IF FF + + KF+ KLL
Sbjct: 1010 NGFFPRFPCLSIGFLCFPNLFPDKFLEG----IFLFFCQLFRLNKFISKLL 870
>TC84889 similar to GP|21537003|gb|AAM61344.1 unknown {Arabidopsis
thaliana}, partial (29%)
Length = 770
Score = 25.8 bits (55), Expect = 5.8
Identities = 14/30 (46%), Positives = 18/30 (59%)
Frame = -3
Query: 92 GSYMYNLFP*IFFFFLNIRGVVKFLRKLLG 121
GSY N FF++ +RGVV FL +L G
Sbjct: 120 GSYELN-----FFYYEGLRGVVGFLYRLEG 46
>TC88769 homologue to GP|19881561|gb|AAM00962.1 hypothetical protein {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (7%)
Length = 912
Score = 25.4 bits (54), Expect = 7.6
Identities = 9/24 (37%), Positives = 13/24 (53%)
Frame = +1
Query: 10 WYMGCQTVVFSVEL*NRYYSVCLF 33
WY C+ + SV N Y++ C F
Sbjct: 652 WYRRCRRQICSVSSTNYYFNFCFF 723
>TC77447 similar to GP|16974501|gb|AAL31160.1 AT3g06720/F3E22_14 {Arabidopsis
thaliana}, partial (96%)
Length = 2271
Score = 25.4 bits (54), Expect = 7.6
Identities = 12/35 (34%), Positives = 18/35 (51%)
Frame = +2
Query: 99 FP*IFFFFLNIRGVVKFLRKLLGSIGIQYVYLWIV 133
+P +F F ++ F+R I IQYV LW +
Sbjct: 2117 YPVLFLFLPKFTSILLFIR-----ISIQYVQLWFL 2206
>TC78649 weakly similar to GP|6651029|gb|AAF22136.1| gamma-glutamylcysteine
synthetase precursor {Phaseolus vulgaris}, partial (32%)
Length = 1052
Score = 25.0 bits (53), Expect = 9.9
Identities = 11/31 (35%), Positives = 19/31 (60%)
Frame = +3
Query: 103 FFFFLNIRGVVKFLRKLLGSIGIQYVYLWIV 133
F F NIR + ++ + G + IQYV L+++
Sbjct: 933 FLFINNIRNLCPYV*FIYGLVQIQYVQLFLL 1025
>BG586932 homologue to PIR|T52080|T52 multi resistance protein [imported] -
Arabidopsis thaliana, partial (3%)
Length = 648
Score = 25.0 bits (53), Expect = 9.9
Identities = 10/28 (35%), Positives = 16/28 (56%), Gaps = 4/28 (14%)
Frame = -2
Query: 10 WYMGCQTVVFS----VEL*NRYYSVCLF 33
W M C T++FS + + NR Y +C +
Sbjct: 509 WLMSCDTLLFS*TIHLNVINRIYELCYY 426
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.368 0.170 0.704
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,791,460
Number of Sequences: 36976
Number of extensions: 95483
Number of successful extensions: 1484
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1473
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1483
length of query: 138
length of database: 9,014,727
effective HSP length: 86
effective length of query: 52
effective length of database: 5,834,791
effective search space: 303409132
effective search space used: 303409132
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC149306.10


