

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148758.11 + phase: 0
(358 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
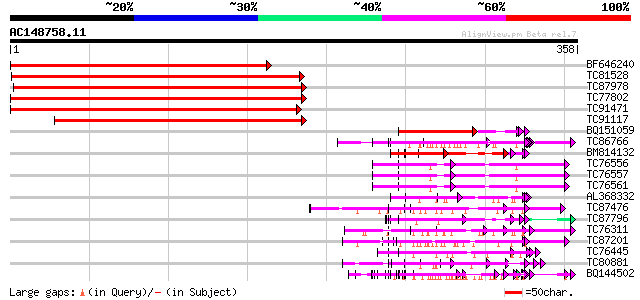
Score E
Sequences producing significant alignments: (bits) Value
BF646240 similar to GP|9294655|dbj gb|AAF34307.1~gene_id:F14O13.... 318 2e-87
TC81528 similar to GP|4803931|gb|AAD29804.1| unknown protein {Ar... 308 2e-84
TC87978 similar to GP|4803931|gb|AAD29804.1| unknown protein {Ar... 261 4e-70
TC77802 similar to GP|20161440|dbj|BAB90364. hypothetical protei... 242 1e-64
TC91471 similar to GP|8778257|gb|AAF79266.1| F12K21.21 {Arabidop... 236 1e-62
TC91117 similar to GP|21593618|gb|AAM65585.1 unknown {Arabidopsi... 200 6e-52
BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melano... 100 1e-21
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 98 6e-21
BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein ... 89 3e-18
TC76556 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 86 2e-17
TC76557 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 86 2e-17
TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 86 2e-17
AL368332 weakly similar to GP|21322711|e pherophorin-dz1 protein... 84 8e-17
TC87476 homologue to PIR|T11622|T11622 extensin class 1 precurso... 84 8e-17
TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear prot... 83 2e-16
TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer a... 81 7e-16
TC87201 similar to PIR|F86388|F86388 hypothetical protein AAG292... 80 1e-15
TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - so... 80 1e-15
TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor ... 80 2e-15
BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 78 6e-15
>BF646240 similar to GP|9294655|dbj gb|AAF34307.1~gene_id:F14O13.5~similar to
unknown protein {Arabidopsis thaliana}, partial (40%)
Length = 630
Score = 318 bits (816), Expect = 2e-87
Identities = 164/165 (99%), Positives = 164/165 (99%)
Frame = +2
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM
Sbjct: 134 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 313
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII
Sbjct: 314 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 493
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAP 165
VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITL LVLYCAP
Sbjct: 494 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLVLVLYCAP 628
>TC81528 similar to GP|4803931|gb|AAD29804.1| unknown protein {Arabidopsis
thaliana}, partial (47%)
Length = 710
Score = 308 bits (789), Expect = 2e-84
Identities = 151/185 (81%), Positives = 169/185 (90%)
Frame = +3
Query: 2 SSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTMI 61
+S+NL GF+LA++S AFIGSSFIIKKKGLQ A +NG ASVGGYGYLLQPLWW+GMVTMI
Sbjct: 150 TSSNLIGFILAVVSGAFIGSSFIIKKKGLQRAGLNGTPASVGGYGYLLQPLWWIGMVTMI 329
Query: 62 VGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIV 121
VGEIANFVAY+YAPAVLVTPLGALSIIVSAVLAHF+L EKLQKMGMLGCL+CI+GST IV
Sbjct: 330 VGEIANFVAYIYAPAVLVTPLGALSIIVSAVLAHFMLGEKLQKMGMLGCLLCIVGSTEIV 509
Query: 122 LHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSI 181
LHAPQE SL+SV +IW LA+QPAFL+YT SAIA+ FL+LYCAPRYGQ+NI VYIGICSI
Sbjct: 510 LHAPQEKSLTSVLEIWLLAVQPAFLLYTASAIAVAFFLILYCAPRYGQTNIFVYIGICSI 689
Query: 182 FGSLT 186
GSLT
Sbjct: 690 IGSLT 704
>TC87978 similar to GP|4803931|gb|AAD29804.1| unknown protein {Arabidopsis
thaliana}, partial (84%)
Length = 1653
Score = 261 bits (666), Expect = 4e-70
Identities = 126/185 (68%), Positives = 153/185 (82%)
Frame = +1
Query: 3 STNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTMIV 62
S N G +LA+ SS FIG+SFI+KKKGL+ A G A VGGY YLL+PLWWVGMVTMI
Sbjct: 133 SENYKGLILAVCSSGFIGASFILKKKGLKRAASRGTRAGVGGYTYLLEPLWWVGMVTMIT 312
Query: 63 GEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVL 122
GE ANFVAY+YAPAVLVTPLGALSIIVS+VLAHFLLKE+LQKMG+LGCL CI+GS +IV+
Sbjct: 313 GEAANFVAYIYAPAVLVTPLGALSIIVSSVLAHFLLKERLQKMGVLGCLSCIVGSIVIVI 492
Query: 123 HAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICSIF 182
HAPQE + +SVQ+IW+LA QP F++Y + +++ L L+L PRYGQ N+LVY+GICS+
Sbjct: 493 HAPQEHTPNSVQEIWELATQPEFMIYAAATVSVVLALILNFEPRYGQKNMLVYLGICSLM 672
Query: 183 GSLTV 187
GSLTV
Sbjct: 673 GSLTV 687
>TC77802 similar to GP|20161440|dbj|BAB90364. hypothetical protein~similar
to Arabidopsis thaliana chromosome 1 F17M19.5, partial
(52%)
Length = 1681
Score = 242 bits (618), Expect = 1e-64
Identities = 115/187 (61%), Positives = 144/187 (76%)
Frame = +3
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQLARVNGPSASVGGYGYLLQPLWWVGMVTM 60
MSS N+ G +LAL SS FIG+SFI+KKKGL+ A +G A GGY YL +PLWWVGM+TM
Sbjct: 375 MSSDNIKGLVLALSSSFFIGASFIVKKKGLKKAGASGIRAGSGGYSYLYEPLWWVGMITM 554
Query: 61 IVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTII 120
IVGEIANF AY +APA+LVTPLGALSII+SA LAH +L+E+L G+LGC +C++GST I
Sbjct: 555 IVGEIANFAAYAFAPAILVTPLGALSIIISAALAHIILRERLHIFGVLGCALCVVGSTTI 734
Query: 121 VLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGICS 180
VLHAPQE + SV ++W LA+ PAFL Y I T LV + P YGQ++I+VYIG+CS
Sbjct: 735 VLHAPQEREIESVPEVWDLAMDPAFLFYAALVITATFILVFHFIPLYGQTHIMVYIGVCS 914
Query: 181 IFGSLTV 187
+ GSL+V
Sbjct: 915 LVGSLSV 935
>TC91471 similar to GP|8778257|gb|AAF79266.1| F12K21.21 {Arabidopsis
thaliana}, partial (39%)
Length = 804
Score = 236 bits (602), Expect = 1e-62
Identities = 116/185 (62%), Positives = 146/185 (78%), Gaps = 1/185 (0%)
Frame = +1
Query: 1 MSSTNLTGFLLALISSAFIGSSFIIKKKGLQ-LARVNGPSASVGGYGYLLQPLWWVGMVT 59
+S+ N+TG +LAL SS FIGSSFIIKK+GL+ A G A VGGY YLL+PLWWVGM+T
Sbjct: 163 LSNENVTGLILALASSLFIGSSFIIKKQGLRRAASTYGVRAGVGGYYYLLEPLWWVGMIT 342
Query: 60 MIVGEIANFVAYMYAPAVLVTPLGALSIIVSAVLAHFLLKEKLQKMGMLGCLICILGSTI 119
MIVGE+ANF+AY +APAVLVTPLGALSIIVSAVLA +LKE+L K+G+LG ++CI GS I
Sbjct: 343 MIVGEVANFIAYAFAPAVLVTPLGALSIIVSAVLADLILKERLHKLGILGIVMCIAGSII 522
Query: 120 IVLHAPQEMSLSSVQQIWKLAIQPAFLMYTTSAIAITLFLVLYCAPRYGQSNILVYIGIC 179
IV+HAP+E ++SV +IW +A QPAFL Y S + + F+V + AP G +N+LVY GIC
Sbjct: 523 IVIHAPKEEPITSVLEIWNMATQPAFLAYVGSVVVLVFFMVFHFAPTCGHTNVLVYTGIC 702
Query: 180 SIFGS 184
S+ GS
Sbjct: 703 SLMGS 717
>TC91117 similar to GP|21593618|gb|AAM65585.1 unknown {Arabidopsis
thaliana}, partial (81%)
Length = 1196
Score = 200 bits (509), Expect = 6e-52
Identities = 91/159 (57%), Positives = 122/159 (76%)
Frame = +1
Query: 29 GLQLARVNGPSASVGGYGYLLQPLWWVGMVTMIVGEIANFVAYMYAPAVLVTPLGALSII 88
GL+ A NG A+ GG+ YL +P WW GM +MIVGEIANF AY +APA+LVTPLGALSII
Sbjct: 1 GLKKAATNGNRAATGGHSYLYEPRWWAGMTSMIVGEIANFAAYAFAPAILVTPLGALSII 180
Query: 89 VSAVLAHFLLKEKLQKMGMLGCLICILGSTIIVLHAPQEMSLSSVQQIWKLAIQPAFLMY 148
SAVLAHF+LKE+L G+LGC +C++GST IVLHAP E + SV+++W LA +P F++Y
Sbjct: 181 FSAVLAHFILKERLHIFGVLGCALCVVGSTTIVLHAPHEREIHSVKEVWHLATEPGFIVY 360
Query: 149 TTSAIAITLFLVLYCAPRYGQSNILVYIGICSIFGSLTV 187
+ +A+ L L+ A YGQ++++VY+GICS+ GS+TV
Sbjct: 361 SCLMVALVLVLIFVFARSYGQTHLVVYVGICSLTGSITV 477
>BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melanogaster},
partial (26%)
Length = 308
Score = 100 bits (248), Expect = 1e-21
Identities = 46/83 (55%), Positives = 46/83 (55%)
Frame = -1
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 296 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 117
Query: 306 PPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP PP P P PP
Sbjct: 116 PPPPPPPPPPPPPPPPPPPPPPP 48
Score = 100 bits (248), Expect = 1e-21
Identities = 46/83 (55%), Positives = 46/83 (55%)
Frame = -3
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 306 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 127
Query: 306 PPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP PP P P PP
Sbjct: 126 PPPPPPPPPPPPPPPPPPPPPPP 58
Score = 100 bits (248), Expect = 1e-21
Identities = 46/83 (55%), Positives = 46/83 (55%)
Frame = -1
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 299 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 120
Query: 306 PPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP PP P P PP
Sbjct: 119 PPPPPPPPPPPPPPPPPPPPPPP 51
Score = 100 bits (248), Expect = 1e-21
Identities = 46/83 (55%), Positives = 46/83 (55%)
Frame = -1
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 293 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 114
Query: 306 PPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP PP P P PP
Sbjct: 113 PPPPPPPPPPPPPPPPPPPPPPP 45
Score = 100 bits (248), Expect = 1e-21
Identities = 46/83 (55%), Positives = 46/83 (55%)
Frame = -1
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 302 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 123
Query: 306 PPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP PP P P PP
Sbjct: 122 PPPPPPPPPPPPPPPPPPPPPPP 54
Score = 100 bits (248), Expect = 1e-21
Identities = 46/83 (55%), Positives = 46/83 (55%)
Frame = -2
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 289 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 110
Query: 306 PPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP PP P P PP
Sbjct: 109 PPPPPPPPPPPPPPPPPPPPPPP 41
Score = 100 bits (248), Expect = 1e-21
Identities = 46/83 (55%), Positives = 46/83 (55%)
Frame = -1
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 287 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 108
Query: 306 PPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP PP P P PP
Sbjct: 107 PPPPPPPPPPPPPPPPPPPPPPP 39
Score = 100 bits (248), Expect = 1e-21
Identities = 46/83 (55%), Positives = 46/83 (55%)
Frame = -2
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 283 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 104
Query: 306 PPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP PP P P PP
Sbjct: 103 PPPPPPPPPPPPPPPPPPPPPPP 35
Score = 100 bits (248), Expect = 1e-21
Identities = 46/83 (55%), Positives = 46/83 (55%)
Frame = -2
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 304 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 125
Query: 306 PPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP PP P P PP
Sbjct: 124 PPPPPPPPPPPPPPPPPPPPPPP 56
Score = 93.6 bits (231), Expect = 1e-19
Identities = 43/78 (55%), Positives = 44/78 (56%)
Frame = -2
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 256 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 77
Query: 306 PPHLLSAPPPHRPPQLTP 323
PP PPP PP + P
Sbjct: 76 PPPPPPPPPPPPPPGVFP 23
Score = 92.4 bits (228), Expect = 2e-19
Identities = 43/79 (54%), Positives = 45/79 (56%)
Frame = -3
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP P
Sbjct: 216 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP--------PPPPPPPPPP 61
Query: 306 PPHLLSAPPPHRPPQLTPD 324
PP PPP PP +P+
Sbjct: 60 PP-----PPPPPPPGFSPE 19
Score = 76.3 bits (186), Expect = 2e-14
Identities = 31/50 (62%), Positives = 31/50 (62%)
Frame = -3
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGI 295
P PP PPPP PPPP PP P PPP P PPPPP P PPPPP GI
Sbjct: 159 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPGFSPERGI 10
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 97.8 bits (242), Expect = 6e-21
Identities = 50/98 (51%), Positives = 58/98 (59%), Gaps = 9/98 (9%)
Frame = +2
Query: 240 SPISHHPSSLPPLLPPPPI--PPPPTP---PSPHPPPSPSPPPPPRPSPPPP----PSLX 290
SP P+S PP PP P+ PPPP P SP PPP+ SPPPPP PPPP P +
Sbjct: 26 SPPPPPPNSPPPPPPPAPVFSPPPPVPYYYSSPPPPPAHSPPPPPXSPPPPPHSPTPPVY 205
Query: 291 RHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPP 328
+L PPPPP+H PPP + S PPP PP + P P PP
Sbjct: 206 PYLSPPPPPPVH-SPPPPVYSPPPPSPPPCVEPPPPPP 316
Score = 81.6 bits (200), Expect = 4e-16
Identities = 47/101 (46%), Positives = 56/101 (54%), Gaps = 12/101 (11%)
Frame = +2
Query: 241 PISHHPSSLPPLLPPPPI--PPPPTPPSPH-----PPPSPSPPPPP---RPSPPPPPSLX 290
P S P PP PPPP PPPP+ P+PH PPPSPSPPP P PSPPPP
Sbjct: 272 PPSPPPCVEPPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPPVYTS 451
Query: 291 RHLGIPPPPPLHLF--PPPHLLSAPPPHRPPQLTPDPSPPL 329
PPP P++ + PPP + S+PP PP + P PP+
Sbjct: 452 -----PPPSPVYAYPSPPPPVYSSPP---PPPVYEGPIPPV 550
Score = 81.3 bits (199), Expect = 5e-16
Identities = 47/107 (43%), Positives = 52/107 (47%), Gaps = 11/107 (10%)
Frame = +2
Query: 262 PTPP----SPHPPP--SPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPP 315
PTPP SP PPP SP PPPPP P PPP + + PPPPP H PPP PPP
Sbjct: 2 PTPPYCVRSPPPPPPNSPPPPPPPAPVFSPPPPVPYYYSSPPPPPAHSPPPPPXSPPPPP 181
Query: 316 HRP-----PQLTPDPSPPLSSHSVTFAARQLKGIIHPSTICPPTLPP 357
H P P L+P P PP+ HS P + PP PP
Sbjct: 182 HSPTPPVYPYLSPPPPPPV--HSPPPPVYSPPPPSPPPCVEPPPPPP 316
Score = 78.2 bits (191), Expect = 5e-15
Identities = 47/110 (42%), Positives = 55/110 (49%), Gaps = 19/110 (17%)
Frame = +2
Query: 241 PISHHPSSLPPLLPPPP----IPPPP--TPPSPHPPPSPSPPPPPRP---SPPPPPSLXR 291
P S P P L PPPP PPPP +PP P PPP PPPPP P PPPP S
Sbjct: 179 PHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPPPPPCVEPPPPSSPAP 358
Query: 292 HLG--------IPPPPPLHLFP--PPHLLSAPPPHRPPQLTPDPSPPLSS 331
H PPP P++ +P PP + ++PPP P P P PP+ S
Sbjct: 359 HQTPYHPPPSPSPPPSPVYAYPSPPPPVYTSPPP-SPVYAYPSPPPPVYS 505
Score = 76.6 bits (187), Expect = 1e-14
Identities = 48/104 (46%), Positives = 54/104 (51%), Gaps = 14/104 (13%)
Frame = +2
Query: 241 PISHHPSSLPP----LLPPPPIPPPPTPPSPHPPPSP---SPPPPPRPSP------PPPP 287
P+ ++ SS PP PPPP PPP P SP PP P PPPPP SP PPPP
Sbjct: 98 PVPYYYSSPPPPPAHSPPPPPXSPPPPPHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPP 277
Query: 288 SLXRHLGIPPPPPLHLFPPPHLLSAPPPHRPP-QLTPDPSPPLS 330
S + PPPPP PP S+P PH+ P P PSPP S
Sbjct: 278 SPPPCVEPPPPPPPPCVEPPP-PSSPAPHQTPYHPPPSPSPPPS 406
Score = 76.3 bits (186), Expect = 2e-14
Identities = 45/105 (42%), Positives = 53/105 (49%), Gaps = 5/105 (4%)
Frame = +2
Query: 230 PRISSVPRLLSPISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPP-----PPPRPSPP 284
P S P + SP P S PP + PPP PPPP P PP SP+P PPP PSPP
Sbjct: 233 PVHSPPPPVYSP---PPPSPPPCVEPPPPPPPPCVEPP-PPSSPAPHQTPYHPPPSPSPP 400
Query: 285 PPPSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPPL 329
P P PPPP++ PPP + A P PP + P PP+
Sbjct: 401 PSPVYAYP---SPPPPVYTSPPPSPVYAYPSPPPPVYSSPPPPPV 526
Score = 61.2 bits (147), Expect = 6e-10
Identities = 36/97 (37%), Positives = 40/97 (41%)
Frame = +2
Query: 208 HYTIYHLPILPLRTPHNFVSFRPRISSVPRLLSPISHHPSSLPPLLPPPPIPPPPTPPSP 267
H T YH P P P SP+ +PS PP+ PP P PSP
Sbjct: 359 HQTPYHPPPSPSPPP-----------------SPVYAYPSPPPPVYTSPPPSPVYAYPSP 487
Query: 268 HPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLF 304
PP SPPPPP P PP PPPPP + F
Sbjct: 488 PPPVYSSPPPPPVYEGPIPPVFGISYASPPPPPFY*F 598
>BM814132 weakly similar to OMNI|MT3615.1 PE_PGRS family protein
{Mycobacterium tuberculosis CDC1551}, partial (7%)
Length = 164
Score = 89.0 bits (219), Expect = 3e-18
Identities = 41/70 (58%), Positives = 42/70 (59%)
Frame = -2
Query: 250 PPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHL 309
PP PPPP PPPP PP P PPP P PPPPP P+PPPPP PPPPP PPP
Sbjct: 160 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPAPPPPP--------PPPPP----PPP-- 23
Query: 310 LSAPPPHRPP 319
PPP PP
Sbjct: 22 ---PPPRPPP 2
Score = 85.1 bits (209), Expect = 4e-17
Identities = 39/65 (60%), Positives = 39/65 (60%)
Frame = -3
Query: 251 PLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLL 310
P PPPP PPPP PP P PPP P PPPPP P PPPPP PPPPP PPP
Sbjct: 162 PPPPPPPPPPPPPPPPPPPPPRPPPPPPPPPPPPPPPP-------PPPPP----PPPPPP 16
Query: 311 SAPPP 315
APPP
Sbjct: 15 PAPPP 1
Score = 84.7 bits (208), Expect = 5e-17
Identities = 40/70 (57%), Positives = 41/70 (58%)
Frame = -1
Query: 250 PPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHL 309
PP PPPP PPPP PP P PPP+P PPPPP P PPPP PPPPP PPP
Sbjct: 164 PPPPPPPPPPPPPPPPPPPPPPAPPPPPPPPPPRPPPP--------PPPPP----PPP-- 27
Query: 310 LSAPPPHRPP 319
PPP PP
Sbjct: 26 ---PPPPPPP 6
Score = 83.2 bits (204), Expect = 1e-16
Identities = 43/74 (58%), Positives = 43/74 (58%)
Frame = -1
Query: 246 PSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFP 305
P PP PPPP PPPP PP P PPP P PPPPPRP PPPPP PPPPP P
Sbjct: 164 PPPPPPPPPPPPPPPPPPPPPPAPPP-PPPPPPPRPPPPPPP--------PPPPP----P 24
Query: 306 PPHLLSAPPPHRPP 319
PP PPP PP
Sbjct: 23 PP-----PPP--PP 3
Score = 82.0 bits (201), Expect = 3e-16
Identities = 40/70 (57%), Positives = 41/70 (58%)
Frame = -3
Query: 259 PPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRP 318
PPPP PP P PPP P PPPPPRP PPPPP PPPPP PPP PPP P
Sbjct: 162 PPPPPPPPPPPPPPPPPPPPPRPPPPPPPP-------PPPPPP---PPP-----PPP--P 34
Query: 319 PQLTPDPSPP 328
P P P+PP
Sbjct: 33 PPPPPPPAPP 4
Score = 79.7 bits (195), Expect = 2e-15
Identities = 38/70 (54%), Positives = 38/70 (54%)
Frame = -1
Query: 259 PPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRP 318
PPPP PP P PPP P PPPPP P PPPPP PPP P PPP PPP P
Sbjct: 161 PPPPPPPPPPPPPPPPPPPPPAPPPPPPP--------PPPRP----PPP-----PPPPPP 33
Query: 319 PQLTPDPSPP 328
P P P PP
Sbjct: 32 PPPPPPPPPP 3
Score = 74.7 bits (182), Expect = 5e-14
Identities = 37/69 (53%), Positives = 37/69 (53%)
Frame = -2
Query: 259 PPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRP 318
PPPP PP P PPP P PPPPP PPPPP PPPPP PPP PPP P
Sbjct: 163 PPPPPPPPPPPPPPPPPPPPP---PPPPP--------PPPPPPPAPPPP----PPPPPPP 29
Query: 319 PQLTPDPSP 327
P P P P
Sbjct: 28 PPPPPRPPP 2
Score = 54.7 bits (130), Expect = 5e-08
Identities = 22/37 (59%), Positives = 23/37 (61%)
Frame = -3
Query: 241 PISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPP 277
P P PP PPPP PPPP PP P PPP P+PPP
Sbjct: 111 PPPPRPPPPPPPPPPPPPPPPPPPPPPPPPPPPAPPP 1
Score = 40.4 bits (93), Expect = 0.001
Identities = 18/34 (52%), Positives = 19/34 (54%)
Frame = -3
Query: 236 PRLLSPISHHPSSLPPLLPPPPIPPPPTPPSPHP 269
PR P P PP PPPP PPPP PP+P P
Sbjct: 102 PRPPPPPPPPPPPPPPPPPPPPPPPPPPPPAPPP 1
>TC76556 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 754
Score = 85.9 bits (211), Expect = 2e-17
Identities = 52/119 (43%), Positives = 57/119 (47%), Gaps = 11/119 (9%)
Frame = -2
Query: 246 PSSLPPLL-PPPPIPPPPTP---PSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPL 301
P S PPLL PPPP PPPP P P P P SP PPPPP P PPPPP R PPPPP
Sbjct: 564 PPSPPPLLYPPPPYPPPPPPLL*PPPPPRRSPYPPPPPPPYPPPPP*PRRSPPYPPPPP- 388
Query: 302 HLFPPPHLLSAPPPHRP-------PQLTPDPSPPLSSHSVTFAARQLKGIIHPSTICPP 353
PPP PPP P L P S P + +F L + +P + P
Sbjct: 387 -P*PPPPRPPPPPPPLPRL*AWFTVMLRPSKS*PFIASMASFMDFSLANVTNPKPLDRP 214
Score = 45.8 bits (107), Expect = 3e-05
Identities = 25/52 (48%), Positives = 25/52 (48%)
Frame = -2
Query: 230 PRISSVPRLLSPISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRP 281
PR S P P P P PP P PPPP PP P PP PPPPP P
Sbjct: 483 PRRSPYPPPPPPPYPPPPP*PRRSPPYPPPPPP*PPPPRPP----PPPPPLP 340
>TC76557 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 912
Score = 85.9 bits (211), Expect = 2e-17
Identities = 52/119 (43%), Positives = 57/119 (47%), Gaps = 11/119 (9%)
Frame = -2
Query: 246 PSSLPPLL-PPPPIPPPPTP---PSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPL 301
P S PPLL PPPP PPPP P P P P SP PPPPP P PPPPP R PPPPP
Sbjct: 722 PPSPPPLLYPPPPYPPPPPPLL*PPPPPRRSPYPPPPPPPYPPPPP*PRRSPPYPPPPP- 546
Query: 302 HLFPPPHLLSAPPPHRP-------PQLTPDPSPPLSSHSVTFAARQLKGIIHPSTICPP 353
PPP PPP P L P S P + +F L + +P + P
Sbjct: 545 -P*PPPPRPPPPPPPLPRL*AWFTVMLRPSKS*PFIASMASFMDFSLANVTNPKPLDRP 372
Score = 45.8 bits (107), Expect = 3e-05
Identities = 25/52 (48%), Positives = 25/52 (48%)
Frame = -2
Query: 230 PRISSVPRLLSPISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRP 281
PR S P P P P PP P PPPP PP P PP PPPPP P
Sbjct: 641 PRRSPYPPPPPPPYPPPPP*PRRSPPYPPPPPP*PPPPRPP----PPPPPLP 498
>TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 1054
Score = 85.9 bits (211), Expect = 2e-17
Identities = 52/119 (43%), Positives = 57/119 (47%), Gaps = 11/119 (9%)
Frame = -3
Query: 246 PSSLPPLL-PPPPIPPPPTP---PSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPL 301
P S PPLL PPPP PPPP P P P P SP PPPPP P PPPPP R PPPPP
Sbjct: 818 PPSPPPLLYPPPPYPPPPPPLL*PPPPPRRSPYPPPPPPPYPPPPP*PRRSPPYPPPPP- 642
Query: 302 HLFPPPHLLSAPPPHRP-------PQLTPDPSPPLSSHSVTFAARQLKGIIHPSTICPP 353
PPP PPP P L P S P + +F L + +P + P
Sbjct: 641 -P*PPPPRPPPPPPPLPRL*AWFTVMLRPSKS*PFIASMASFMDFSLANVTNPKPLDRP 468
Score = 45.8 bits (107), Expect = 3e-05
Identities = 25/52 (48%), Positives = 25/52 (48%)
Frame = -3
Query: 230 PRISSVPRLLSPISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRP 281
PR S P P P P PP P PPPP PP P PP PPPPP P
Sbjct: 737 PRRSPYPPPPPPPYPPPPP*PRRSPPYPPPPPP*PPPPRPP----PPPPPLP 594
>AL368332 weakly similar to GP|21322711|e pherophorin-dz1 protein {Volvox
carteri f. nagariensis}, partial (10%)
Length = 478
Score = 84.0 bits (206), Expect = 8e-17
Identities = 49/103 (47%), Positives = 55/103 (52%), Gaps = 16/103 (15%)
Frame = +1
Query: 241 PISHHPSSLPPLLPPPPIPPPPTPPSPH----PPPSPS-----PPPPPRPSPPPPPSLXR 291
P++ PS P+ PPP P PP PH PPP+PS PPPPPR PPPPPS
Sbjct: 85 PLNPPPSPFHPINPPPSPFHPFNPPPPHSIKSPPPAPSHSSPPPPPPPRKXPPPPPS--- 255
Query: 292 HLGIPPPPPLHLFPPP---HLLSAPPPH----RPPQLTPDPSP 327
H PPPP H PPP H ++ PPH PP L P PSP
Sbjct: 256 HPFSPPPP--HNHPPPSPHHPIAPSPPHVRPSPPPPLPPSPSP 378
Score = 79.7 bits (195), Expect = 2e-15
Identities = 48/103 (46%), Positives = 52/103 (49%), Gaps = 24/103 (23%)
Frame = +1
Query: 251 PLLPPPP---IPPPPTPPSP-HPPPSP-------------SPPPPPRPSPPPPPSLXRHL 293
P LPPPP + PPP+P P +PPPSP SPPP P S PPPP R
Sbjct: 58 PHLPPPPSHPLNPPPSPFHPINPPPSPFHPFNPPPPHSIKSPPPAPSHSSPPPPPPPRK- 234
Query: 294 GIPPPPPLHLF--PPPHLLSAPPPHR-----PPQLTPDPSPPL 329
PPPPP H F PPPH P PH PP + P P PPL
Sbjct: 235 -XPPPPPSHPFSPPPPHNHPPPSPHHPIAPSPPHVRPSPPPPL 360
Score = 54.7 bits (130), Expect = 5e-08
Identities = 28/60 (46%), Positives = 32/60 (52%), Gaps = 2/60 (3%)
Frame = +1
Query: 271 PSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLF--PPPHLLSAPPPHRPPQLTPDPSPP 328
P P PPPP PPPS + PPP P H F PPPH + +PPP P +P P PP
Sbjct: 52 PFPHLPPPPSHPLNPPPSPFHPIN-PPPSPFHPFNPPPPHSIKSPPP-APSHSSPPPPPP 225
Score = 48.5 bits (114), Expect = 4e-06
Identities = 26/47 (55%), Positives = 26/47 (55%), Gaps = 1/47 (2%)
Frame = +1
Query: 241 PISHHPSSLPPLLPPPPIPPPPTPPS-PHPPPSPSPPPPPRPSPPPP 286
P SH S PP PPP P P PS PH PSP PP PP PSP P
Sbjct: 247 PPSHPFSPPPPHNHPPPSPHHPIAPSPPHVRPSPPPPLPPSPSPYHP 387
Score = 38.9 bits (89), Expect = 0.003
Identities = 33/95 (34%), Positives = 36/95 (37%), Gaps = 27/95 (28%)
Frame = +1
Query: 215 PILPLRTPHN-FVSFRP----RISSVPRLLSPISHHPSSLPPLLPPPP-----IPPPP-- 262
P P+ P + F F P I S P S S P P PPPP PPPP
Sbjct: 106 PFHPINPPPSPFHPFNPPPPHSIKSPPPAPSHSSPPPPPPPRKXPPPPPSHPFSPPPPHN 285
Query: 263 -TPPSPH--------------PPPSPSPPPPPRPS 282
PPSPH PPP P P P P+
Sbjct: 286 HPPPSPHHPIAPSPPHVRPSPPPPLPPSPSPYHPT 390
>TC87476 homologue to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 939
Score = 84.0 bits (206), Expect = 8e-17
Identities = 64/175 (36%), Positives = 79/175 (44%), Gaps = 36/175 (20%)
Frame = +3
Query: 190 STYLSLFQYSCMLYRLIFHYTIYHLPILP--LRTPHNFVSFRPRI--SSVPRLLSPISHH 245
ST L L Q+ +L+ + HLP+LP + T H+ P + + LL + HH
Sbjct: 159 STSLLLHQHPHLLHPIFISRHHLHLPLLPHHMFTSHHLHRLHPHHLHTFINLLLHHLLHH 338
Query: 246 -------------PSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPP--------P 279
PS PP + P PP P+PP P+ PPPSPSPPPP P
Sbjct: 339 RLHISTSHHRPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPP 518
Query: 280 RPSPPPP---PSLXRHLGIPPPPPLHLFPPPHLLSAPPPH---RPPQLTPDPSPP 328
SPPPP S PPPP + PPP S PPP+ PP P PSPP
Sbjct: 519 SASPPPPYYYKSPPPPSPSPPPPYGYKSPPPPSPSPPPPYIYKSPP--PPSPSPP 677
Score = 82.8 bits (203), Expect = 2e-16
Identities = 44/97 (45%), Positives = 52/97 (53%), Gaps = 8/97 (8%)
Frame = +2
Query: 240 SPISHHPSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPPPRPSPPPPPSLXRHLG 294
SP PS PP + P PP P+PP P+ PPPSPSPPPP PPPP+
Sbjct: 23 SPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPTPS---- 190
Query: 295 IPPPPPLHLFPPPHLLSAPPPH---RPPQLTPDPSPP 328
PPPP ++ PPP S PPP+ PP +P P PP
Sbjct: 191 -PPPPYIYKSPPPPSPSPPPPYVYKSPPPPSPSPPPP 298
Score = 82.4 bits (202), Expect = 2e-16
Identities = 44/97 (45%), Positives = 52/97 (53%), Gaps = 8/97 (8%)
Frame = +2
Query: 240 SPISHHPSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPPPRPSPPPPPSLXRHLG 294
SP PS PP + P PP P+PP P+ PPP+PSPPPP PPPPS
Sbjct: 71 SPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPTPSPPPPYIYKSPPPPSPS---- 238
Query: 295 IPPPPPLHLFPPPHLLSAPPPH---RPPQLTPDPSPP 328
PPPP ++ PPP S PPP+ PP +P P PP
Sbjct: 239 -PPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPP 346
Score = 78.2 bits (191), Expect = 5e-15
Identities = 41/90 (45%), Positives = 49/90 (53%), Gaps = 11/90 (12%)
Frame = +2
Query: 250 PPLLPPPPIPPPPTPPSPH-----PPPSPSPPPP------PRPSPPPPPSLXRHLGIPPP 298
PP + P PP P+PP P+ PPPSPSPPPP P PSP PPP ++ PP
Sbjct: 5 PPYIYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPP---YVYKSPP 175
Query: 299 PPLHLFPPPHLLSAPPPHRPPQLTPDPSPP 328
PP PPP++ +PPP P PSPP
Sbjct: 176 PPTPSPPPPYIYKSPPP-------PSPSPP 244
Score = 77.8 bits (190), Expect = 6e-15
Identities = 46/117 (39%), Positives = 58/117 (49%), Gaps = 5/117 (4%)
Frame = +2
Query: 240 SPISHHPSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPPPRPSPPPPPSLXRHLG 294
SP PS PP + P PP P+PP P+ PPPSPSPPPP PPPPS
Sbjct: 119 SPPPPSPSPPPPYVYKSPPPPTPSPPPPYIYKSPPPPSPSPPPPYVYKSPPPPSPS---- 286
Query: 295 IPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPPLSSHSVTFAARQLKGIIHPSTIC 351
PPPP ++ PPP S PPP+ +P + P ++ S+ PS+IC
Sbjct: 287 -PPPPYVYKSPPPPSPSPPPPY--IYKSPPTTIPFTTTSICL*VPSSSFSFTPSSIC 448
Score = 74.3 bits (181), Expect = 7e-14
Identities = 41/88 (46%), Positives = 45/88 (50%), Gaps = 13/88 (14%)
Frame = +2
Query: 254 PPPPIPPPPTPPSPH----------PPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHL 303
PPP I P PPSP PPPSPSPPPP PPPPS PPPP ++
Sbjct: 2 PPPYIYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPS-----PPPPYVYK 166
Query: 304 FPPPHLLSAPPPH---RPPQLTPDPSPP 328
PPP S PPP+ PP +P P PP
Sbjct: 167 SPPPPTPSPPPPYIYKSPPPPSPSPPPP 250
Score = 70.9 bits (172), Expect = 7e-13
Identities = 54/144 (37%), Positives = 67/144 (46%), Gaps = 19/144 (13%)
Frame = +3
Query: 191 TYLSLFQYSCMLYRLIFHYTIYHLPILPLRTPHNFVSFRPRISSVPR----LLSPISHHP 246
T+++L + + +RL T +H P P P P S P SP P
Sbjct: 300 TFINLLLHHLLHHRLHIS-TSHHRPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSP 476
Query: 247 SSLPPLL---PPPPI---PPP-----PTPPSPHPPP--SPSPPPPPRPSPPPPPSLXRHL 293
S PP + PPPP PPP P PPSP PPP PPPP PSPPPP ++
Sbjct: 477 SPPPPYVYKSPPPPSASPPPPYYYKSPPPPSPSPPPPYGYKSPPPPSPSPPPP-----YI 641
Query: 294 GIPPPPPLHLFPP--PHLLSAPPP 315
PPPP PP P+L ++PPP
Sbjct: 642 YKSPPPPSPSPPPYHPYLYNSPPP 713
Score = 65.9 bits (159), Expect = 2e-11
Identities = 44/123 (35%), Positives = 53/123 (42%), Gaps = 11/123 (8%)
Frame = +2
Query: 240 SPISHHPSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPPPRPSPPPPPSLXRHLG 294
SP PS PP + P PP P+PP P+ PPPSPSPPPP PPPPS
Sbjct: 167 SPPPPTPSPPPPYIYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPS------ 328
Query: 295 IPPPPPLHLFPPPHLLSAPPPHRPPQLT------PDPSPPLSSHSVTFAARQLKGIIHPS 348
P P PPP++ +PP P T P S + S+ PS
Sbjct: 329 -PSP------PPPYIYKSPPTTIPFTTTSICL*VPSSSFSFTPSSICLQISSPTISFTPS 487
Query: 349 TIC 351
T+C
Sbjct: 488 TLC 496
>TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear protein.
[strain B95-8 Human herpesvirus 4] {Epstein-barr
virus}, partial (23%)
Length = 431
Score = 82.8 bits (203), Expect = 2e-16
Identities = 44/92 (47%), Positives = 47/92 (50%), Gaps = 2/92 (2%)
Frame = -1
Query: 239 LSPISHHPSSLPPLLPP--PPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIP 296
L+P +P PP PP PP PP PP P PPP P PPPPP PPP P P
Sbjct: 266 LNPQGSYPPWPPPSPPPIPPPYAPPCAPPPPPPPPEPPPPPPPPGDPPPYPP-------P 108
Query: 297 PPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPP 328
PPPL +P P APPP PP P P PP
Sbjct: 107 VPPPLP-YPTPCAPPAPPPPPPPLPPPPPYPP 15
Score = 75.9 bits (185), Expect = 2e-14
Identities = 39/80 (48%), Positives = 41/80 (50%)
Frame = -1
Query: 240 SPISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPP 299
SP P PP PPPP PPP PP P PP P P PPP P P P P+ PP P
Sbjct: 227 SPPPIPPPYAPPCAPPPPPPPPEPPPPPPPPGDPPPYPPPVPPPLPYPTPC----APPAP 60
Query: 300 PLHLFPPPHLLSAPPPHRPP 319
P PPP L PPP+ PP
Sbjct: 59 P----PPPPPLPPPPPYPPP 12
Score = 74.7 bits (182), Expect = 5e-14
Identities = 41/83 (49%), Positives = 42/83 (50%), Gaps = 3/83 (3%)
Frame = -1
Query: 246 PSSLPPLLPP--PPIPPPPTPPSPHPPPSPSPPPPPRPSPPP-PPSLXRHLGIPPPPPLH 302
P S PP+ PP PP PPP PP P PPP P PP P P PPP PP L PP P
Sbjct: 233 PPSPPPIPPPYAPPCAPPPPPPPPEPPPPPPPPGDPPPYPPPVPPPLPYPTPCAPPAP-- 60
Query: 303 LFPPPHLLSAPPPHRPPQLTPDP 325
PPP PPP PP P P
Sbjct: 59 --PPP-----PPPLPPPPPYPPP 12
Score = 62.8 bits (151), Expect = 2e-10
Identities = 42/120 (35%), Positives = 47/120 (39%)
Frame = -1
Query: 238 LLSPISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPP 297
+L +S H +L P PP PPP PP P P P PPPP PPPP PP
Sbjct: 299 VLGVLS*H*IALNPQGSYPPWPPPSPPPIPPPYAPPCAPPPP----PPPPE-------PP 153
Query: 298 PPPLHLFPPPHLLSAPPPHRPPQLTPDPSPPLSSHSVTFAARQLKGIIHPSTICPPTLPP 357
PPP PPP PP P PPL +P+ PP PP
Sbjct: 152 PPP------------PPPGDPPPYPPPVPPPLP---------------YPTPCAPPAPPP 54
Score = 53.5 bits (127), Expect = 1e-07
Identities = 27/56 (48%), Positives = 28/56 (49%), Gaps = 7/56 (12%)
Frame = -1
Query: 241 PISHHPSSLPPLLPPPPIPPPPTPPSPHP-------PPSPSPPPPPRPSPPPPPSL 289
P P PP PPP PPP PP P+P PP P PP PP P PPP SL
Sbjct: 170 PPPEPPPPPPPPGDPPPYPPPVPPPLPYPTPCAPPAPPPPPPPLPPPPPYPPP*SL 3
>TC76311 homologue to GP|3204132|emb|CAA07235.1 extensin {Cicer arietinum},
partial (77%)
Length = 973
Score = 80.9 bits (198), Expect = 7e-16
Identities = 45/106 (42%), Positives = 51/106 (47%), Gaps = 18/106 (16%)
Frame = +2
Query: 240 SPISHHPSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPPPRPSPPPPPSLXRHLG 294
SP PS PP P PP P+PP P+ PPPSPSPPPP PPPPS H
Sbjct: 41 SPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSPTPHTP 220
Query: 295 I----------PPPPPLHLFPPPHLLSAPPPHR---PPQLTPDPSP 327
PPPP H PP S+PPP+ PP +P P+P
Sbjct: 221 YHYKSPPPPTASPPPPYHYVSPPPPTSSPPPYHYTSPPPPSPAPAP 358
Score = 75.5 bits (184), Expect = 3e-14
Identities = 41/88 (46%), Positives = 45/88 (50%), Gaps = 10/88 (11%)
Frame = +2
Query: 254 PPPPIPPPPTPPSPHPPPSPSPPP------PPRPSPPPPPSLXRHLGIPPPPPLHLFPPP 307
PP P PPPP PPPSPSPPP PP PSP PPP + PPPP PPP
Sbjct: 2 PPSPSPPPPYYYHSPPPPSPSPPPPYYYKSPPPPSPSPPPP---YYYKSPPPPSPSPPPP 172
Query: 308 HLLSAPPPHRPPQLTP----DPSPPLSS 331
+ +PPP P TP P PP +S
Sbjct: 173 YYYQSPPPPSPTPHTPYHYKSPPPPTAS 256
Score = 69.7 bits (169), Expect = 2e-12
Identities = 44/108 (40%), Positives = 48/108 (43%), Gaps = 16/108 (14%)
Frame = +2
Query: 240 SPISHHPSSLPPLL-----PPPPIPPPP----TPPSPHPPPSP----SPPPPPRPSPPPP 286
SP PS PP PP P PPPP +PP P P P PPPP SPPPP
Sbjct: 89 SPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSPTPHTPYHYKSPPPPTASPPPP 268
Query: 287 PSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLT---PDPSPPLSS 331
+ + PPPP PP H S PPP P T P PP+ S
Sbjct: 269 -----YHYVSPPPPTSSPPPYHYTSPPPPSPAPAPTYIYKSPPPPMKS 397
Score = 64.3 bits (155), Expect = 7e-11
Identities = 51/137 (37%), Positives = 64/137 (46%), Gaps = 33/137 (24%)
Frame = +2
Query: 212 YHLPILPLRTP---HNFVSFRPRISSVPR---LLSPISHHPSSLPPLL---PPPPIP--- 259
YH P P +P + + S P S P SP PS PP PPPP P
Sbjct: 35 YHSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYQSPPPPSPTPH 214
Query: 260 --------PPPT--PPSPH-----PPPSPSPPP-----PPRPSPPPPPSLXRHLGIPPPP 299
PPPT PP P+ PPP+ SPPP PP PSP P P+ ++ PPP
Sbjct: 215 TPYHYKSPPPPTASPPPPYHYVSPPPPTSSPPPYHYTSPPPPSPAPAPT---YIYKSPPP 385
Query: 300 PLHLFPPP-HLLSAPPP 315
P+ PPP ++ ++PPP
Sbjct: 386 PMKSPPPPVYIYASPPP 436
Score = 57.0 bits (136), Expect = 1e-08
Identities = 38/97 (39%), Positives = 47/97 (48%), Gaps = 6/97 (6%)
Frame = +2
Query: 212 YHLPILPLRTPH---NFVSFRPRISSVPRLLSPISHHPSSLPPLLPPPPIP-PPPTPPSP 267
Y P P TPH ++ S P +S P P H+ S PP PPP P PPSP
Sbjct: 179 YQSPPPPSPTPHTPYHYKSPPPPTASPP----PPYHYVSPPPPTSSPPPYHYTSPPPPSP 346
Query: 268 HPPPS--PSPPPPPRPSPPPPPSLXRHLGIPPPPPLH 302
P P+ PPPP SPPPP ++ PPPP++
Sbjct: 347 APAPTYIYKSPPPPMKSPPPPV----YIYASPPPPIY 445
Score = 45.4 bits (106), Expect = 3e-05
Identities = 34/97 (35%), Positives = 40/97 (41%), Gaps = 9/97 (9%)
Frame = +2
Query: 270 PPSPSPPPPPRPSPPPPPSLXRHLGIPP----PPPLHLF--PPPHLLSAPPPH---RPPQ 320
PPSPSPPPP P PP PPP + + PPP S PPP+ PP
Sbjct: 2 PPSPSPPPPYYYHSP-----------PPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPP 148
Query: 321 LTPDPSPPLSSHSVTFAARQLKGIIHPSTICPPTLPP 357
+P P PP S + H + PPT P
Sbjct: 149 PSPSPPPPYYYQSPPPPSPTPHTPYHYKSPPPPTASP 259
>TC87201 similar to PIR|F86388|F86388 hypothetical protein AAG29221.1
[imported] - Arabidopsis thaliana, partial (41%)
Length = 991
Score = 80.1 bits (196), Expect = 1e-15
Identities = 48/109 (44%), Positives = 54/109 (49%), Gaps = 25/109 (22%)
Frame = +1
Query: 245 HPSSLPPLLPPP--------------PIPPPPTPPSPH----PPPSP----SPPPPPR-- 280
+PS PP PPP P PPPP+PP P+ PPP P SPPPPP
Sbjct: 43 YPSPPPPSPPPPIRIQITTRAPPYVYPSPPPPSPPPPYIYKSPPPPPYVYKSPPPPPYVY 222
Query: 281 PSPPPP-PSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPP 328
PSPPPP P PPPPP PPP++ +PPP P PSPP
Sbjct: 223 PSPPPPSPPPPYIYKSPPPPPYVYTPPPYVYKSPPP-------PSPSPP 348
Score = 76.6 bits (187), Expect = 1e-14
Identities = 49/104 (47%), Positives = 58/104 (55%), Gaps = 20/104 (19%)
Frame = +1
Query: 245 HPSSLPPLLPPPPI----PPPP---TPP-----SPHPPPSPSPPPP-PRPSPPPPPSLXR 291
+PS PP PPP I PPPP TPP SP PPPSPSPPPP SPPPPP + +
Sbjct: 220 YPSPPPPSPPPPYIYKSPPPPPYVYTPPPYVYKSP-PPPSPSPPPPYVYKSPPPPPYVYK 396
Query: 292 HLGIPPPPPLHLFPPP--HLLSAPPP-----HRPPQLTPDPSPP 328
PPPP ++ PPP ++ +PPP PP +P P PP
Sbjct: 397 --SPPPPPYVYESPPPPQYVYKSPPPPPYVYESPPPPSPSPPPP 522
Score = 76.3 bits (186), Expect = 2e-14
Identities = 44/100 (44%), Positives = 53/100 (53%), Gaps = 16/100 (16%)
Frame = +1
Query: 245 HPSSLPPLLPPPPI----PPPP-----TPPSPHPPPSPSPPPPPRP----SPPPPPSLXR 291
+PS PP PPP I PPPP PP P+ PSP PP PP P SPPPPP +
Sbjct: 118 YPSPPPPSPPPPYIYKSPPPPPYVYKSPPPPPYVYPSPPPPSPPPPYIYKSPPPPPYVYT 297
Query: 292 ---HLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPP 328
++ PPPP PPP++ +PPP PP + P PP
Sbjct: 298 PPPYVYKSPPPPSPSPPPPYVYKSPPP--PPYVYKSPPPP 411
Score = 67.4 bits (163), Expect = 8e-12
Identities = 47/121 (38%), Positives = 54/121 (43%), Gaps = 28/121 (23%)
Frame = +1
Query: 236 PRLLSPISHHPSSLPPLL----PPPPI---PPP-----PTPPSPHPPP-----SP----- 273
P + P PS PP + PPPP PPP P PPSP PPP SP
Sbjct: 208 PPYVYPSPPPPSPPPPYIYKSPPPPPYVYTPPPYVYKSPPPPSPSPPPPYVYKSPPPPPY 387
Query: 274 ---SPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPD---PSP 327
SPPPPP PPP + PPPP ++ PPP S PPP+ + P SP
Sbjct: 388 VYKSPPPPPYVYESPPPPQYVYKSPPPPPYVYESPPPPSPSPPPPYYYKPIPPPYVYESP 567
Query: 328 P 328
P
Sbjct: 568 P 570
Score = 66.6 bits (161), Expect = 1e-11
Identities = 47/124 (37%), Positives = 54/124 (42%), Gaps = 13/124 (10%)
Frame = +1
Query: 243 SHH---PSSLPPLLPPPPIPPPP--------TPPSPHPPPSPSPPPPPR--PSPPPPPSL 289
SHH P P PPPP PPPP PP +P P P PPPP SPPPPP +
Sbjct: 16 SHHLLRPYVYPS--PPPPSPPPPIRIQITTRAPPYVYPSPPPPSPPPPYIYKSPPPPPYV 189
Query: 290 XRHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPPLSSHSVTFAARQLKGIIHPST 349
+ PP PPP++ +PPP PP SPP + T K PS
Sbjct: 190 YKS------PP----PPPYVYPSPPPPSPPPPYIYKSPPPPPYVYTPPPYVYKSPPPPSP 339
Query: 350 ICPP 353
PP
Sbjct: 340 SPPP 351
Score = 65.1 bits (157), Expect = 4e-11
Identities = 53/146 (36%), Positives = 67/146 (45%), Gaps = 29/146 (19%)
Frame = +1
Query: 211 IYHLPILP--LRTPHNFVSFRPRISSVPRLLSPISHHPSSLPPLL----PPPPI----PP 260
IY P P + TP +V P S P P + PP + PPPP PP
Sbjct: 259 IYKSPPPPPYVYTPPPYVYKSPPPPS-PSPPPPYVYKSPPPPPYVYKSPPPPPYVYESPP 435
Query: 261 PPT------PPSPH-----PPPSPSPPPPP--RPSPPP-----PPSLXRHLGIPPPPPLH 302
PP PP P+ PPPSPSPPPP +P PPP PP + + PPP P
Sbjct: 436 PPQYVYKSPPPPPYVYESPPPPSPSPPPPYYYKPIPPPYVYESPPYIYKS---PPPAPYV 606
Query: 303 LFPPPHLLSAPP-PHRPPQLTPDPSP 327
PP++ +PP P++P Q + P P
Sbjct: 607 YKSPPYVYKSPPTPYKPYQYSSPPPP 684
Score = 64.7 bits (156), Expect = 5e-11
Identities = 44/115 (38%), Positives = 51/115 (44%), Gaps = 26/115 (22%)
Frame = +1
Query: 240 SPISHHPSSLPPLL----PPPPI-----PPPP----TPPSPH-----PPPSP---SPPPP 278
SP PS PP + PPPP PPPP +PP P PPP P PPP
Sbjct: 319 SPPPPSPSPPPPYVYKSPPPPPYVYKSPPPPPYVYESPPPPQYVYKSPPPPPYVYESPPP 498
Query: 279 PRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPP-----HRPPQLTPDPSPP 328
P PSPPPP + P PPP PP++ +PPP PP + P P
Sbjct: 499 PSPSPPPP-----YYYKPIPPPYVYESPPYIYKSPPPAPYVYKSPPYVYKSPPTP 648
>TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - soybean
(fragment), partial (76%)
Length = 821
Score = 80.1 bits (196), Expect = 1e-15
Identities = 49/109 (44%), Positives = 53/109 (47%), Gaps = 19/109 (17%)
Frame = +2
Query: 246 PSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPP------PRPSPPPPPSLXRHLG 294
PS PP P PP P+PPSP+ PPPSPSPPPP P PSP PPP +
Sbjct: 2 PSPPPPYYYKSPPPPSPSPPSPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPY--YYK 175
Query: 295 IPPPPPLHLFPPPHLLSAPPP---HRPPQL-----TPDPSPPLSSHSVT 335
PPPP PP H S PPP PP P PSPP H V+
Sbjct: 176 SPPPPSPSPPPPYHYQSPPPPSPISHPPNYYKSPPPPSPSPPPPYHYVS 322
Score = 77.8 bits (190), Expect = 6e-15
Identities = 43/105 (40%), Positives = 49/105 (45%), Gaps = 15/105 (14%)
Frame = +2
Query: 240 SPISHHPSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPPPRPSPPPPPSLXRH-- 292
SP PS PP P PP P+PP P+ PPPSPSPPPP PPPPS H
Sbjct: 80 SPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYHYQSPPPPSPISHPP 259
Query: 293 --------LGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPPL 329
PPPP H PP + +PPP P + P PP+
Sbjct: 260 NYYKSPPPPSPSPPPPYHYVSPPPPVKSPPP--PAYIYASPPPPI 388
Score = 75.9 bits (185), Expect = 2e-14
Identities = 48/110 (43%), Positives = 51/110 (45%), Gaps = 11/110 (10%)
Frame = +2
Query: 233 SSVPRLLSPISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPP--------PRPSPP 284
S P SP S + PP PP P PPPP PPPSPSPPPP P PSPP
Sbjct: 32 SPPPPSPSPPSPYYYKSPP--PPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPP 205
Query: 285 PPPSLXRHLGIPPPPPLHLFPPPHLLSAPPPH---RPPQLTPDPSPPLSS 331
PP H PPPP PP + S PPP PP P PP+ S
Sbjct: 206 PP----YHYQSPPPPSPISHPPNYYKSPPPPSPSPPPPYHYVSPPPPVKS 343
Score = 61.6 bits (148), Expect = 4e-10
Identities = 39/101 (38%), Positives = 44/101 (42%), Gaps = 21/101 (20%)
Frame = +2
Query: 240 SPISHHPSSLPPLL---PPPPIPPPPTPPSPHPPPSPSP----------PPPPRPSPPPP 286
SP PS PP PPPP P PP P PP PSP PPPP PSPPPP
Sbjct: 128 SPPPPSPSPPPPYYYKSPPPPSPSPPPPYHYQSPPPPSPISHPPNYYKSPPPPSPSPPPP 307
Query: 287 PSLXRHLGIPPPPP--------LHLFPPPHLLSAPPPHRPP 319
H PPPP ++ PPP + + P + P
Sbjct: 308 ----YHYVSPPPPVKSPPPPAYIYASPPPPIYN*PHNTKSP 418
>TC80881 similar to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (34%)
Length = 597
Score = 79.7 bits (195), Expect = 2e-15
Identities = 42/95 (44%), Positives = 51/95 (53%), Gaps = 8/95 (8%)
Frame = +3
Query: 242 ISHHPSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPPPRPSPPPPPSLX---RHL 293
+ H PS PP + P PP P+PP P+ PPPSPSPPPP PPPPS ++
Sbjct: 315 LHHLPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSLPPNYV 494
Query: 294 GIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPP 328
PPPP PPP++ +PPP P PSPP
Sbjct: 495 YKSPPPPSPSPPPPYVYKSPPP-------PSPSPP 578
Score = 67.4 bits (163), Expect = 8e-12
Identities = 37/82 (45%), Positives = 43/82 (52%), Gaps = 5/82 (6%)
Frame = +3
Query: 240 SPISHHPSSLPPLLPPPPIPPPPTPPSPH-----PPPSPSPPPPPRPSPPPPPSLXRHLG 294
SP PS PP + P PP P+PP P+ PPPSPS PP PPPPS
Sbjct: 357 SPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSLPPNYVYKSPPPPSPS---- 524
Query: 295 IPPPPPLHLFPPPHLLSAPPPH 316
PPPP ++ PPP S PPP+
Sbjct: 525 -PPPPYVYKSPPPPSPSPPPPY 587
Score = 59.7 bits (143), Expect = 2e-09
Identities = 40/105 (38%), Positives = 49/105 (46%), Gaps = 11/105 (10%)
Frame = +3
Query: 211 IYHLPILPLRTPHNFVSFRPRISSVPRLLSPISHHPSSLPPLLPPPPI----PPPPTPPS 266
++HLP P P+ + S P S P P + P PPPP PPPP+P
Sbjct: 315 LHHLPSPP--PPYVYKSPPPPSPSPP---PPYVYKSPPPPSPSPPPPYVYKSPPPPSPSL 479
Query: 267 PH-------PPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLF 304
P PPPSPSPPPP PPPPS P PPP +++
Sbjct: 480 PPNYVYKSPPPPSPSPPPPYVYKSPPPPS-------PSPPPPYIY 593
Score = 45.1 bits (105), Expect(2) = 1e-09
Identities = 26/62 (41%), Positives = 29/62 (45%)
Frame = +3
Query: 273 PSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPPLSSH 332
PSPPPP PPPPS PPPP PP +PPP + P PSP L +
Sbjct: 327 PSPPPPYVYKSPPPPSPS------PPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSLPPN 488
Query: 333 SV 334
V
Sbjct: 489 YV 494
Score = 44.7 bits (104), Expect = 6e-05
Identities = 29/88 (32%), Positives = 38/88 (42%), Gaps = 9/88 (10%)
Frame = +2
Query: 260 PPPTPPS---------PHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLL 310
P PTPPS PP PPPP PS PPPP +H +PP +
Sbjct: 167 PQPTPPSHGQQPPYYFKSPPYYYKSPPPPSPS-------------PPPPYVHKYPPYYYK 307
Query: 311 SAPPPHRPPQLTPDPSPPLSSHSVTFAA 338
S PPP P T +++ S++F +
Sbjct: 308 SPPPP--SPFTTSSIRLQVTTSSISFTS 385
Score = 35.0 bits (79), Expect = 0.045
Identities = 24/58 (41%), Positives = 28/58 (47%), Gaps = 12/58 (20%)
Frame = +2
Query: 241 PISHHPSSLPPL---LPP-----PPI----PPPPTPPSPHPPPSPSPPPPPRPSPPPP 286
P +++P PP PP PP PPPP+P SP PP PP SPPPP
Sbjct: 152 PNNNYPQPTPPSHGQQPPYYFKSPPYYYKSPPPPSP-SPPPPYVHKYPPYYYKSPPPP 322
Score = 34.7 bits (78), Expect(2) = 1e-09
Identities = 16/36 (44%), Positives = 18/36 (49%), Gaps = 4/36 (11%)
Frame = +2
Query: 250 PPLLPPPPIPPPPTPPSP----HPPPSPSPPPPPRP 281
PP P PP P+PP P +PP PPPP P
Sbjct: 221 PPYYYKSPPPPSPSPPPPYVHKYPPYYYKSPPPPSP 328
>BQ144502 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (39%)
Length = 1358
Score = 77.8 bits (190), Expect = 6e-15
Identities = 44/99 (44%), Positives = 48/99 (48%), Gaps = 12/99 (12%)
Frame = -2
Query: 241 PISHHPSSLPPLL----PPPPIP--------PPPTPPSPHPPPSPSPPPPPRPSPPPPPS 288
P HP PP L PPPP P P +PP P PPP P+ PPRP PPPPP
Sbjct: 316 PALPHPPGPPPPLDRAPPPPPAPLPLASRLRPRASPPPPPPPPGPAASAPPRPPPPPPPP 137
Query: 289 LXRHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSP 327
R PPPP PPPH PP PP+ P P+P
Sbjct: 136 PQR----PPPP-----PPPHTHQPEPPTSPPRRQP-PAP 50
Score = 72.4 bits (176), Expect = 3e-13
Identities = 45/110 (40%), Positives = 47/110 (41%), Gaps = 3/110 (2%)
Frame = -2
Query: 251 PLLPPPPIPPPPTPPSPHPPPSPSPPPP---PRPSPPPPPSLXRHLGIPPPPPLHLFPPP 307
P LP PP PPPP +P PPP+P P PR SPPPPP PPP P PP
Sbjct: 316 PALPHPPGPPPPLDRAPPPPPAPLPLASRLRPRASPPPPP--------PPPGPAASAPP- 164
Query: 308 HLLSAPPPHRPPQLTPDPSPPLSSHSVTFAARQLKGIIHPSTICPPTLPP 357
PPP PPQ P P PP H PPT PP
Sbjct: 163 --RPPPPPPPPPQRPPPPPPP-----------------HTHQPEPPTSPP 71
Score = 70.1 bits (170), Expect = 1e-12
Identities = 48/148 (32%), Positives = 60/148 (40%), Gaps = 23/148 (15%)
Frame = -1
Query: 229 RPRISSVPRLLSPISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRP------S 282
+PR + VP L ++ +P P PPPP+PPPP+PP P +PPPPP P +
Sbjct: 617 QPRPALVP--LVRVATYPR---PPRPPPPVPPPPSPPRAPPQSEAAPPPPPPPRGARPRA 453
Query: 283 PPPPPSLXRHLGIP-----------------PPPPLHLFPPPHLLSAPPPHRPPQLTPDP 325
P PPS G P P H P P L PPP P P P
Sbjct: 452 PLRPPSAPDPAGHPRAPRQGVAVVPGPRSGNSPRAAHRGPEPRPLPRPPPPTRPPAAPGP 273
Query: 326 SPPLSSHSVTFAARQLKGIIHPSTICPP 353
PP + + I P+T PP
Sbjct: 272 GPPPAPGPTPLSKPPAAEGIPPATAPPP 189
Score = 69.7 bits (169), Expect = 2e-12
Identities = 37/71 (52%), Positives = 37/71 (52%)
Frame = -3
Query: 245 HPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLF 304
HP P PPPP PPPP P P PPP P PPRP PP SL PPPPP
Sbjct: 216 HP---PRHRPPPPGPPPPPPLGPRPPPHPPRSGPPRPHPPTRTSLS-----PPPPPPAAS 61
Query: 305 PPPHLLSAPPP 315
PPP APPP
Sbjct: 60 PPP---PAPPP 37
Score = 68.9 bits (167), Expect = 3e-12
Identities = 34/69 (49%), Positives = 35/69 (50%)
Frame = -2
Query: 247 SSLPPLLPPPPIPPPPTPPSPHPPPSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPP 306
S L P PPP PPPP P + PP P PPPPP PPPPP H PP P PP
Sbjct: 235 SRLRPRASPPPPPPPPGPAASAPPRPPPPPPPPPQRPPPPPPPHTHQPEPPTSPPRRQPP 56
Query: 307 PHLLSAPPP 315
APPP
Sbjct: 55 AP--RAPPP 35
Score = 66.6 bits (161), Expect = 1e-11
Identities = 40/102 (39%), Positives = 42/102 (40%), Gaps = 13/102 (12%)
Frame = -3
Query: 233 SSVPRLLSPISHHPSSLPPLLPPPPIPPPPTP---------PSPHPPPSPSPPPPPRPSP 283
S P P HP+ P PPP P P +P P H PP P PPPPP P
Sbjct: 336 SRAPSPAPPSPTHPAPRRPWTGPPPRPRPHSP*QAACGRGHPPRHRPPPPGPPPPPPLGP 157
Query: 284 PPPPSLXRHLGIPPPPPLHLF----PPPHLLSAPPPHRPPQL 321
PPP R P PP PPP S PPP PP L
Sbjct: 156 RPPPHPPRSGPPRPHPPTRTSLSPPPPPPAASPPPPAPPPLL 31
Score = 63.5 bits (153), Expect = 1e-10
Identities = 42/115 (36%), Positives = 46/115 (39%), Gaps = 29/115 (25%)
Frame = -3
Query: 238 LLSPISHHPSSLPPLLPPPPIPPPPT------------------PPSPHP-------PPS 272
L + + HPS P PPPP PPPP+ PP P PP
Sbjct: 744 LRASAAQHPSRAPRRHPPPPPPPPPSSPPGMGTRPSCQENWWAAPPGARPSRPSRYIPPP 565
Query: 273 PSPPPPPRPSPPPP--PSLXRHLGIPPPPPLHLFP--PPHLLSAPPPHRPPQLTP 323
SPPP P+P PP PS R PP PP P PP P P RPP P
Sbjct: 564 TSPPPASPPAPVPPQSPSTIRGRAPPPAPPPRRPPARPPPAPQRPRPRRPPARPP 400
Score = 63.5 bits (153), Expect = 1e-10
Identities = 57/176 (32%), Positives = 63/176 (35%), Gaps = 47/176 (26%)
Frame = -3
Query: 229 RPRISSVPRLLSPISHHPSSLPPLLP-------PPPIPPPPTPPSPHPPPSPSPPPPPRP 281
RP P P + P+ +PP P PPP PPP PP+ PPP+P P P RP
Sbjct: 591 RPSRYIPPPTSPPPASPPAPVPPQSPSTIRGRAPPPAPPPRRPPA-RPPPAPQRPRPRRP 415
Query: 282 SPPPPPSLXRH-----------LGIP--------PPPPLH-------LFPPPHL------ 309
P PPS R G P PP P H PPP
Sbjct: 414 -PARPPSGRRRGARAKVRKLPKGGTPGSRAPSPAPPSPTHPAPRRPWTGPPPRPRPHSP* 238
Query: 310 -----LSAPPPHRPPQLTPDPSPPLSSHSVTFAARQLKGIIHPST---ICPPTLPP 357
PP HRPP P P PPL R HP T + PP PP
Sbjct: 237 QAACGRGHPPRHRPPPPGPPPPPPLGPRPPPHPPRSGPPRPHPPTRTSLSPPPPPP 70
Score = 63.5 bits (153), Expect = 1e-10
Identities = 45/106 (42%), Positives = 47/106 (43%), Gaps = 23/106 (21%)
Frame = -3
Query: 249 LPPLLPPPPIPPPPTPPSP-HPPP-SPSPPPPPRPSP-----------------PPPPSL 289
LP P P P PPSP HP P P PPPRP P PPPP
Sbjct: 360 LPKGGTPGSRAPSPAPPSPTHPAPRRPWTGPPPRPRPHSP*QAACGRGHPPRHRPPPP-- 187
Query: 290 XRHLGIPPPPPLHLFPPPH-LLSAPPPHRPP---QLTPDPSPPLSS 331
G PPPPPL PPPH S PP PP L+P P PP +S
Sbjct: 186 ----GPPPPPPLGPRPPPHPPRSGPPRPHPPTRTSLSPPPPPPAAS 61
Score = 62.0 bits (149), Expect = 3e-10
Identities = 44/119 (36%), Positives = 48/119 (39%), Gaps = 20/119 (16%)
Frame = -1
Query: 230 PRISSVPRLLSPISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPP----------PP 279
PR + PR +H PL PPP PP P P PPP+P P P PP
Sbjct: 374 PRSGNSPRA----AHRGPEPRPLPRPPPPTRPPAAPGPGPPPAPGPTPLSKPPAAEGIPP 207
Query: 280 RPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAP-PPHRP---PQLTP------DPSPP 328
+PPP R P PPP PP AP PPH P P L P P PP
Sbjct: 206 ATAPPPRARRLRPPSAPAPPPT---PPAAAPPAPTPPHAPA*APHLPPPPPAPRPPRPP 39
Score = 58.2 bits (139), Expect = 5e-09
Identities = 34/70 (48%), Positives = 38/70 (53%), Gaps = 6/70 (8%)
Frame = -3
Query: 246 PSSLPPLLP-PPPIPPPPTPPSPHPP--PSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLH 302
P PPL P PPP PP PP PHPP S SPPPPP + PPPP+ PPPL
Sbjct: 183 PPPPPPLGPRPPPHPPRSGPPRPHPPTRTSLSPPPPPPAASPPPPA---------PPPLL 31
Query: 303 LFP---PPHL 309
++ PP L
Sbjct: 30 VYN**LPPSL 1
Score = 58.2 bits (139), Expect = 5e-09
Identities = 51/165 (30%), Positives = 61/165 (36%), Gaps = 26/165 (15%)
Frame = -2
Query: 219 LRTPHNFVSFRPRISSVPRLLSPISHHPSSLPPLLPPPPIP---PPPTPPSPHPPPSPSP 275
L TP + PR S PR H+P PP PPP P PP PP+P PP+
Sbjct: 580 LHTPAHLAP--PRQSPRPRPPPEPLHNPRPRPPPRPPPAAPARAPPSGPPAPPTPPATRA 407
Query: 276 PP----PPRPSPPPPPSLXRHLGI-----------PPPPPLHLFPPPHLLSAPPPHRP-- 318
PP P P RH G+ PP P PPP + PPP P
Sbjct: 406 PPVRASPWCPGQGQETPQGRHTGVQSPVPCPALPHPPGP-----PPPLDRAPPPPPAPLP 242
Query: 319 ------PQLTPDPSPPLSSHSVTFAARQLKGIIHPSTICPPTLPP 357
P+ +P P PP + + R P PP PP
Sbjct: 241 LASRLRPRASPPPPPPPPGPAASAPPRPPPPPPPPPQRPPPPPPP 107
Score = 58.2 bits (139), Expect = 5e-09
Identities = 37/92 (40%), Positives = 43/92 (46%), Gaps = 13/92 (14%)
Frame = -1
Query: 250 PPLLPPPPIPPPPTPP---SPHPPPSPSPP-----PPPRPSPPPPP----SLXRHLGIPP 297
P LP PP PP P +PH P P PPPRP PPPP L + +G P
Sbjct: 791 PSALPAPPSATPPGAPCARAPHSIPRAHPAATPLLPPPRPPPPPPEWERVPLAKKIGGQP 612
Query: 298 PPPLHLFPPPHLLSAP-PPHRPPQLTPDPSPP 328
P L P + + P PP PP + P PSPP
Sbjct: 611 RPA--LVPLVRVATYPRPPRPPPPVPPPPSPP 522
Score = 55.5 bits (132), Expect = 3e-08
Identities = 39/116 (33%), Positives = 44/116 (37%), Gaps = 29/116 (25%)
Frame = -1
Query: 240 SPISHHPSSLPPLLPPP-PIPPPPTPPSPHPP---------------------------- 270
+P + P S+P P P+ PPP PP P PP
Sbjct: 749 APCARAPHSIPRAHPAATPLLPPPRPP-PPPPEWERVPLAKKIGGQPRPALVPLVRVATY 573
Query: 271 PSPSPPPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPS 326
P P PPPP P PP PP PPPP PPP P RPP PDP+
Sbjct: 572 PRPPRPPPPVPPPPSPPRAPPQSEAAPPPP----PPPRGARPRAPLRPPS-APDPA 420
Score = 55.5 bits (132), Expect = 3e-08
Identities = 30/67 (44%), Positives = 32/67 (46%), Gaps = 6/67 (8%)
Frame = -2
Query: 229 RPRISSVPRLLSP---ISHHPSSLPPLLPPPPIPPPPTPPSPHPPPSPSPPP---PPRPS 282
RPR S P P S P PP PPP PPPP PP H P P+ PP PP P
Sbjct: 226 RPRASPPPPPPPPGPAASAPPRPPPPPPPPPQRPPPPPPPHTHQPEPPTSPPRRQPPAPR 47
Query: 283 PPPPPSL 289
PPP +
Sbjct: 46 APPPSGI 26
Score = 50.1 bits (118), Expect = 1e-06
Identities = 31/80 (38%), Positives = 37/80 (45%), Gaps = 9/80 (11%)
Frame = -1
Query: 215 PILPLRTPHNFVSFRPRISSVPRLLSPISHHPSSLPPLLP-PPPIPPPPTPPSPHPPPSP 273
P P TP +S P +P +P PP P PPP PP PP+P PP +P
Sbjct: 266 PPAPGPTP---LSKPPAAEGIPPATAPPPRARRLRPPSAPAPPPTPPAAAPPAPTPPHAP 96
Query: 274 S-----PPPPPR---PSPPP 285
+ PPPPP P PPP
Sbjct: 95 A*APHLPPPPPAPRPPRPPP 36
Score = 48.1 bits (113), Expect = 5e-06
Identities = 43/133 (32%), Positives = 47/133 (35%), Gaps = 35/133 (26%)
Frame = -2
Query: 231 RISSVPRLLSPISHHPS------SLPPLLPPPPIPPPPTPPSP----------------- 267
R S R P HP+ SL PPPP PPP P P
Sbjct: 790 RPPSPHRPAPPPQAHPARERRTASLARTPPPPPSSPPPAPLLPPRNGNASLLPRKLVGSP 611
Query: 268 -----------HPPPSPSPP-PPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPP 315
H P +PP PRP PPP P H P PPP PPP + PP
Sbjct: 610 ARRSSLSSESLHTPAHLAPPRQSPRPRPPPEP---LHNPRPRPPPR---PPPAAPARAPP 449
Query: 316 HRPPQLTPDPSPP 328
PP P+PP
Sbjct: 448 SGPP---APPTPP 419
Score = 44.3 bits (103), Expect = 7e-05
Identities = 35/115 (30%), Positives = 42/115 (36%), Gaps = 13/115 (11%)
Frame = -3
Query: 256 PPIPPPPTPPSPHPPPSPSPPPPPRPSPPP--PPSLXRH-----------LGIPPPPPLH 302
PP + HP +P PPP P PPP PP + G P P
Sbjct: 759 PPRRTLRASAAQHPSRAPRRHPPPPPPPPPSSPPGMGTRPSCQENWWAAPPGARPSRPSR 580
Query: 303 LFPPPHLLSAPPPHRPPQLTPDPSPPLSSHSVTFAARQLKGIIHPSTICPPTLPP 357
PPP ++PPP PP P PP S ++ A P PP PP
Sbjct: 579 YIPPP---TSPPPASPPA----PVPPQSPSTIRGRA--------PPPAPPPRRPP 460
Score = 38.9 bits (89), Expect = 0.003
Identities = 41/146 (28%), Positives = 45/146 (30%), Gaps = 47/146 (32%)
Frame = -2
Query: 230 PRISSVPRLLSPISHHPSSLPPLLPPPPIPPPPTPPSP----------HPPPSPSPPPPP 279
PR S VPR H + P PP P S PPSP P PP
Sbjct: 919 PRYSLVPRA------HAREIDRAPTFEHFPGPPDPASTPLDRARACCARRPPSPHRPAPP 758
Query: 280 --------------RPSPPPPPSLXRHLGIPPPPPLHLFPP--------PHLLSAPPPHR 317
+PPPPPS PPP L PP P L P R
Sbjct: 757 PQAHPARERRTASLARTPPPPPS--------SPPPAPLLPPRNGNASLLPRKLVGSPARR 602
Query: 318 ---------------PPQLTPDPSPP 328
PP+ +P P PP
Sbjct: 601 SSLSSESLHTPAHLAPPRQSPRPRPP 524
Score = 30.4 bits (67), Expect = 1.1
Identities = 29/110 (26%), Positives = 34/110 (30%), Gaps = 38/110 (34%)
Frame = -3
Query: 257 PIPPPPTPPSPHPPPSPSPPPPPRP---------------------------SPPPP-PS 288
P+P P PH P S P P P SP P P
Sbjct: 1008 PLPRSARSPRPHRPASAPSSPAPLPARDCRRGTLLFRALTRARSTARRRSSTSPAPQIPP 829
Query: 289 LXRHL-------GIPPPPPLHLFPPPHLL---SAPPPHRPPQLTPDPSPP 328
L R + + PP PP L +A P R P+ P P PP
Sbjct: 828 LRRSIVRARAARAVRPPRTAQRHPPRRTLRASAAQHPSRAPRRHPPPPPP 679
Score = 27.3 bits (59), Expect = 9.3
Identities = 19/56 (33%), Positives = 20/56 (34%)
Frame = -2
Query: 276 PPPPRPSPPPPPSLXRHLGIPPPPPLHLFPPPHLLSAPPPHRPPQLTPDPSPPLSS 331
P PP P+ P PP P PPP A R L P PP SS
Sbjct: 850 PGPPDPASTPLDRARACCARRPPSPHRPAPPPQAHPARE-RRTASLARTPPPPPSS 686
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.142 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,953,388
Number of Sequences: 36976
Number of extensions: 753503
Number of successful extensions: 70071
Number of sequences better than 10.0: 1958
Number of HSP's better than 10.0 without gapping: 8346
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 25969
length of query: 358
length of database: 9,014,727
effective HSP length: 97
effective length of query: 261
effective length of database: 5,428,055
effective search space: 1416722355
effective search space used: 1416722355
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148758.11


