

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148758.1 - phase: 0 /pseudo
(143 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
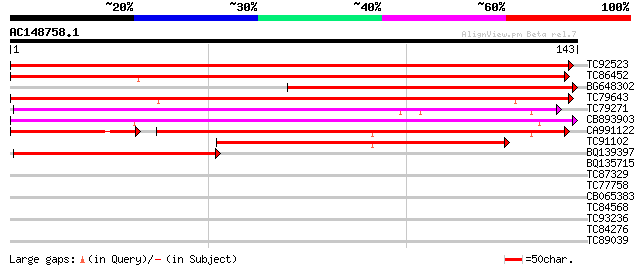
TC92523 similar to PIR|T09892|T09892 hypothetical protein T22A6.... 179 3e-46
TC86452 similar to PIR|T09892|T09892 hypothetical protein T22A6.... 163 2e-41
BG648302 similar to GP|22202725|dbj contains ESTs AU032617(S1163... 133 3e-32
TC79643 similar to PIR|C86410|C86410 protein F3M18.18 [imported]... 129 4e-31
TC79271 weakly similar to GP|20466444|gb|AAM20539.1 unknown prot... 108 7e-25
CB893903 weakly similar to GP|20466444|gb unknown protein {Arabi... 105 5e-24
CA991122 similar to PIR|C86420|C86 unknown protein 124288-12173... 87 3e-22
TC91102 homologue to PIR|C86420|C86420 unknown protein 124288-1... 69 5e-13
BQ139397 weakly similar to PIR|T09892|T098 hypothetical protein ... 57 2e-09
BQ135715 similar to GP|7025451|gb| somatostatin receptor-interac... 30 0.25
TC87329 similar to GP|21618251|gb|AAM67301.1 unknown {Arabidopsi... 27 2.8
TC77758 similar to GP|1255951|emb|CAA65634.1 PS60 {Nicotiana tab... 27 3.6
CB065383 homologue to PIR|D83571|D83 conserved hypothetical prot... 27 3.6
TC84568 similar to GP|17224922|gb|AAL37169.1 putative chloroplas... 26 4.7
TC93236 weakly similar to PIR|E96671|E96671 hypothetical protein... 26 4.7
TC84276 weakly similar to GP|23495823|dbj|BAC20033. putative NAD... 26 4.7
TC89039 homologue to GP|5230728|gb|AAD40979.1| peroxisomal coppe... 26 6.2
>TC92523 similar to PIR|T09892|T09892 hypothetical protein T22A6.120 -
Arabidopsis thaliana, partial (45%)
Length = 1327
Score = 179 bits (454), Expect = 3e-46
Identities = 87/142 (61%), Positives = 110/142 (77%)
Frame = +1
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPV 60
+E+LH+FLEFQLPRQWAPV ++ LG K + + L+FS +GP+LY+NT PVDVG RPV
Sbjct: 10 IEELHQFLEFQLPRQWAPVFSDLPLGPQWKQRSSASLQFSFMGPRLYVNTIPVDVGKRPV 189
Query: 61 TGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFSYV 120
TGLRL LEG +SNRLAIH+QHL+SLPK L D++N SY F++KV+ FS+V
Sbjct: 190 TGLRLYLEGKKSNRLAIHMQHLSSLPKIFQLEDDSNENFRRKSYDKRFYEKVQWKNFSHV 369
Query: 121 CTAPVESDDSLSIVTGAQLQVE 142
CTAPVES++ LS+VTGAQLQVE
Sbjct: 370 CTAPVESEEELSVVTGAQLQVE 435
>TC86452 similar to PIR|T09892|T09892 hypothetical protein T22A6.120 -
Arabidopsis thaliana, partial (80%)
Length = 2233
Score = 163 bits (413), Expect = 2e-41
Identities = 83/142 (58%), Positives = 107/142 (74%), Gaps = 1/142 (0%)
Frame = +3
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKH-QVNTWLRFSILGPKLYINTTPVDVGNRP 59
+E+LH+FLEFQLPRQWAPV GE+ LG RK Q ++ L+FS +GPKLY+NT+PV VG +P
Sbjct: 1128 IEELHQFLEFQLPRQWAPVFGELALGPDRKKSQSSSSLQFSFMGPKLYVNTSPVVVGMKP 1307
Query: 60 VTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFSY 119
VTGLRL LEG +SN LAIHLQHL+SLPK+ L D N +S S +++KV+ FS+
Sbjct: 1308 VTGLRLYLEGKKSNCLAIHLQHLSSLPKTFQLKDETNRNVSDASSERKYYEKVQWKSFSH 1487
Query: 120 VCTAPVESDDSLSIVTGAQLQV 141
+CTAPVES D ++VTGA +V
Sbjct: 1488 ICTAPVESYDDNAVVTGAHFEV 1553
>BG648302 similar to GP|22202725|dbj contains ESTs AU032617(S11633)
D46758(S11633) C73071(E2861)~similar to Oryza sativa
chromosome 1, partial (5%)
Length = 816
Score = 133 bits (334), Expect = 3e-32
Identities = 64/73 (87%), Positives = 68/73 (92%)
Frame = +2
Query: 71 RSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFSYVCTAPVESDDS 130
RSNRLAI+LQHL SLPKSLPLADNA AY SCDSYSC +HKK+K NCFSYVCTAP+ESDDS
Sbjct: 8 RSNRLAINLQHLVSLPKSLPLADNAPAYSSCDSYSCKYHKKLKWNCFSYVCTAPIESDDS 187
Query: 131 LSIVTGAQLQVEK 143
LSIVTGAQLQVEK
Sbjct: 188 LSIVTGAQLQVEK 226
>TC79643 similar to PIR|C86410|C86410 protein F3M18.18 [imported] -
Arabidopsis thaliana, partial (74%)
Length = 2150
Score = 129 bits (324), Expect = 4e-31
Identities = 69/149 (46%), Positives = 103/149 (68%), Gaps = 7/149 (4%)
Frame = +1
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTW--LRFSILGPKLYINTTPVDVGNR 58
+E+LH+FLEFQLPRQWAP+ G++ L K++ N L+F+++GPKLY+NT VD GNR
Sbjct: 1108 IEELHQFLEFQLPRQWAPMYGDLPLVFDHKYKRNASPSLQFTLMGPKLYVNTVKVDSGNR 1287
Query: 59 PVTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFS 118
PVTG+RL LEG ++N L+IHLQHL+ +P L ++++ + + +++ VK + FS
Sbjct: 1288 PVTGIRLYLEGKKNNHLSIHLQHLSEVPGVLEISEDHSYDPIDEPDDRGYYEPVKWSMFS 1467
Query: 119 YVCTAPVE-----SDDSLSIVTGAQLQVE 142
+V TAPV+ D+S SIVT A +V+
Sbjct: 1468 HVYTAPVQYNSSRMDESTSIVTKAWFEVK 1554
>TC79271 weakly similar to GP|20466444|gb|AAM20539.1 unknown protein
{Arabidopsis thaliana}, partial (26%)
Length = 1719
Score = 108 bits (270), Expect = 7e-25
Identities = 67/153 (43%), Positives = 92/153 (59%), Gaps = 15/153 (9%)
Frame = +2
Query: 2 EDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPVDVGNRPVT 61
EDL FLEFQ+PRQWAP+ E+ L R+ + L+F LGPKL+I++T V +PV
Sbjct: 1259 EDLQYFLEFQIPRQWAPMFCELPLRHQRRKTSSLSLQFCCLGPKLHISSTEVVSEQKPVV 1438
Query: 62 GLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANA------YLSCD--SYSCTFHKKVK 113
GLRL LEG +S+RLA+H+ HL+SLP + L+ +A+ + D S F + V+
Sbjct: 1439 GLRLYLEGKKSDRLALHINHLSSLPNKMILSSDASTPSIQSMWRGSDENESSNQFLEPVR 1618
Query: 114 RNCFSYVCTAPVESDDS-------LSIVTGAQL 139
FS VCTA V+ D + + IVTGAQL
Sbjct: 1619 WKRFSNVCTAVVKHDPNWLNDCGGVYIVTGAQL 1717
>CB893903 weakly similar to GP|20466444|gb unknown protein {Arabidopsis
thaliana}, partial (35%)
Length = 815
Score = 105 bits (263), Expect = 5e-24
Identities = 62/151 (41%), Positives = 87/151 (57%), Gaps = 8/151 (5%)
Frame = +3
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRK-HQVNTWLRFSILGPKLYINTTPVDVGNRP 59
M DL FL++Q + WAPV ++ LG ++ +L +++GPKLY+NT V VG RP
Sbjct: 48 MSDLSYFLDYQGYKIWAPVHNDLPLGPTTNISTISPFLTLNLMGPKLYVNTDKVTVGKRP 227
Query: 60 VTGLRLQLEGSRSNRLAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVKRNCFSY 119
+TG+RL LEG + NRLAIH++HL + P L S + F + + FS+
Sbjct: 228 ITGMRLFLEGMKCNRLAIHVEHLLNTPTMLSNKIEDTTIWSEEINDERFFEAISGKKFSH 407
Query: 120 VCTAPVESDDSLS-------IVTGAQLQVEK 143
VCTAPV+ + S IVTGAQL V+K
Sbjct: 408 VCTAPVKYNPKWSTEKNVAFIVTGAQLHVKK 500
>CA991122 similar to PIR|C86420|C86 unknown protein 124288-121737 [imported]
- Arabidopsis thaliana, partial (32%)
Length = 744
Score = 87.4 bits (215), Expect(2) = 3e-22
Identities = 51/117 (43%), Positives = 72/117 (60%), Gaps = 13/117 (11%)
Frame = +3
Query: 38 RFSILGPKLYINTTPVDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSLP------L 91
+FSI+G KLY++ + VG RPVTG+RL LEG++ NRL++HLQHL SLPK L +
Sbjct: 132 QFSIMGQKLYVSQEQITVGRRPVTGIRLCLEGNKQNRLSVHLQHLVSLPKILQPYWDSHV 311
Query: 92 ADNANAYLSCDSYSCTFHKKVKRNCFSYVCTAPVESDDS-------LSIVTGAQLQV 141
A A + + + + VK FS+V TAP+E+ ++ + IVTGAQL V
Sbjct: 312 AIGAPKWQGPEEQDSRWFEPVKWKNFSHVSTAPIENPETFIGDFSGVYIVTGAQLGV 482
Score = 33.1 bits (74), Expect(2) = 3e-22
Identities = 16/33 (48%), Positives = 21/33 (63%)
Frame = +1
Query: 1 MEDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQV 33
+E+L FLEFQ+PR WAP+ + G RK V
Sbjct: 22 IEELRYFLEFQIPRVWAPLHDRVP-GQQRKEPV 117
>TC91102 homologue to PIR|C86420|C86420 unknown protein 124288-121737
[imported] - Arabidopsis thaliana, partial (32%)
Length = 1061
Score = 69.3 bits (168), Expect = 5e-13
Identities = 38/80 (47%), Positives = 50/80 (62%), Gaps = 6/80 (7%)
Frame = +3
Query: 53 VDVGNRPVTGLRLQLEGSRSNRLAIHLQHLASLPKSLP------LADNANAYLSCDSYSC 106
V VG +PVTGLRL LEG++ NRLAIHLQHL SLPK+L +A A + +
Sbjct: 300 VTVGRKPVTGLRLSLEGNKQNRLAIHLQHLVSLPKNLQPHWDAHMAIGAPKWQGPEEQDS 479
Query: 107 TFHKKVKRNCFSYVCTAPVE 126
+ + +K FS+V TAP+E
Sbjct: 480 RWFEPIKWKNFSHVSTAPIE 539
>BQ139397 weakly similar to PIR|T09892|T098 hypothetical protein T22A6.120 -
Arabidopsis thaliana, partial (13%)
Length = 618
Score = 57.4 bits (137), Expect = 2e-09
Identities = 26/52 (50%), Positives = 36/52 (69%)
Frame = +3
Query: 2 EDLHRFLEFQLPRQWAPVLGEIHLGSYRKHQVNTWLRFSILGPKLYINTTPV 53
+DL FLEFQ+PR+WAP+ E+ L RK + L+FS + PKL+IN+T V
Sbjct: 282 DDLQYFLEFQIPREWAPMFSELPLRHQRKKTYSPPLQFSFMSPKLHINSTQV 437
>BQ135715 similar to GP|7025451|gb| somatostatin receptor-interacting protein
splice variant b {Homo sapiens}, partial (2%)
Length = 1165
Score = 30.4 bits (67), Expect = 0.25
Identities = 14/29 (48%), Positives = 20/29 (68%), Gaps = 1/29 (3%)
Frame = -3
Query: 78 HLQHLASL-PKSLPLADNANAYLSCDSYS 105
H+ H + L ++P AD+ + YLSCDSYS
Sbjct: 1136 HISHSSLLVTPTVPPADDTDRYLSCDSYS 1050
>TC87329 similar to GP|21618251|gb|AAM67301.1 unknown {Arabidopsis thaliana},
partial (20%)
Length = 2118
Score = 26.9 bits (58), Expect = 2.8
Identities = 10/20 (50%), Positives = 15/20 (75%)
Frame = +1
Query: 79 LQHLASLPKSLPLADNANAY 98
L HL+SLP LP+ +A++Y
Sbjct: 1039 LSHLSSLPSPLPMTSHASSY 1098
>TC77758 similar to GP|1255951|emb|CAA65634.1 PS60 {Nicotiana tabacum},
partial (96%)
Length = 2197
Score = 26.6 bits (57), Expect = 3.6
Identities = 9/12 (75%), Positives = 10/12 (83%)
Frame = +3
Query: 116 CFSYVCTAPVES 127
CF Y+C APVES
Sbjct: 1911 CFKYICCAPVES 1946
>CB065383 homologue to PIR|D83571|D83 conserved hypothetical protein PA0588
[imported] - Pseudomonas aeruginosa (strain PAO1),
partial (24%)
Length = 614
Score = 26.6 bits (57), Expect = 3.6
Identities = 22/65 (33%), Positives = 27/65 (40%), Gaps = 5/65 (7%)
Frame = +2
Query: 59 PVTGLRLQLEGSRSNR-----LAIHLQHLASLPKSLPLADNANAYLSCDSYSCTFHKKVK 113
PV G + + S R L Q A P SLPLA + L CD + +H K
Sbjct: 5 PVAGNSCKPQASSCKREQPPGLQAAAQAGALTPSSLPLAAKSLP-LYCDFLTRRYHSDSK 181
Query: 114 RNCFS 118
R C S
Sbjct: 182 RTCLS 196
>TC84568 similar to GP|17224922|gb|AAL37169.1 putative chloroplast-targeted
beta-amylase {Brassica napus}, partial (8%)
Length = 535
Score = 26.2 bits (56), Expect = 4.7
Identities = 13/30 (43%), Positives = 16/30 (53%)
Frame = +2
Query: 27 SYRKHQVNTWLRFSILGPKLYINTTPVDVG 56
SYRK + T+ I G K Y N T D+G
Sbjct: 233 SYRKRIITTYESM*ITGTKRYRNQTRSDIG 322
>TC93236 weakly similar to PIR|E96671|E96671 hypothetical protein F13O11.13
[imported] - Arabidopsis thaliana, partial (7%)
Length = 631
Score = 26.2 bits (56), Expect = 4.7
Identities = 23/77 (29%), Positives = 33/77 (41%), Gaps = 5/77 (6%)
Frame = +2
Query: 45 KLYINTTPVDVGNRPVTGLRLQLEGSRSNRLAIHLQHLAS-LPKSLPLAD--NANAY--L 99
K+YI T PV+ RL + + SN + + P+ P D ++ Y L
Sbjct: 296 KIYIGTPPVE---------RLAVADTGSNLIWVQCSPCKKCFPQDKPYFDPNKSSTYMGL 448
Query: 100 SCDSYSCTFHKKVKRNC 116
SCDS SC+ K C
Sbjct: 449 SCDSQSCSSLPLGKHRC 499
>TC84276 weakly similar to GP|23495823|dbj|BAC20033. putative NADPH HC toxin
reductase {Oryza sativa (japonica cultivar-group)},
partial (15%)
Length = 803
Score = 26.2 bits (56), Expect = 4.7
Identities = 10/26 (38%), Positives = 19/26 (72%)
Frame = -3
Query: 72 SNRLAIHLQHLASLPKSLPLADNANA 97
+N L++ + H++ PK+LPL D ++A
Sbjct: 756 NNFLSLIMMHMSLQPKNLPLIDGSSA 679
>TC89039 homologue to GP|5230728|gb|AAD40979.1| peroxisomal
copper-containing amine oxidase {Glycine max}, partial
(33%)
Length = 981
Score = 25.8 bits (55), Expect = 6.2
Identities = 10/15 (66%), Positives = 13/15 (86%)
Frame = +3
Query: 39 FSILGPKLYINTTPV 53
FS+L P +YINTTP+
Sbjct: 924 FSLLIPVVYINTTPI 968
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.321 0.136 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,183,241
Number of Sequences: 36976
Number of extensions: 78273
Number of successful extensions: 453
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 449
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 449
length of query: 143
length of database: 9,014,727
effective HSP length: 87
effective length of query: 56
effective length of database: 5,797,815
effective search space: 324677640
effective search space used: 324677640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC148758.1


