

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148218.5 - phase: 0
(164 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
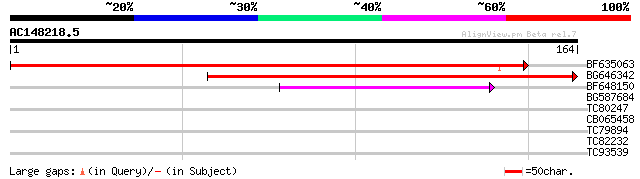
Score E
Sequences producing significant alignments: (bits) Value
BF635063 weakly similar to PIR|F84486|F84 probable retroelement ... 222 6e-59
BG646342 weakly similar to PIR|F84486|F84 probable retroelement ... 183 3e-47
BF648150 similar to GP|14586969|gb| pol polyprotein {Citrus x pa... 40 5e-04
BG587684 similar to GP|12858190|dbj evidence:NAS~putative~unclas... 32 0.15
TC80247 similar to PIR|C86476|C86476 protein F15O4.45 [imported]... 28 2.2
CB065458 27 3.7
TC79894 similar to GP|21104712|dbj|BAB93301. hypothetical protei... 27 4.8
TC82232 similar to GP|10177316|dbj|BAB10642. kinesin-like protei... 26 6.3
TC93539 weakly similar to GP|17063165|gb|AAL32979.1 At1g12970/F1... 26 8.2
>BF635063 weakly similar to PIR|F84486|F84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(4%)
Length = 677
Score = 222 bits (565), Expect = 6e-59
Identities = 115/151 (76%), Positives = 127/151 (83%), Gaps = 1/151 (0%)
Frame = -2
Query: 1 MGSKWDIEKFTGDNDFRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARS 60
MGSK DIEKFTGDNDF LWKVKMEAVLIQQKCEK LKGE SLPVTMSRAEKTEMVDKARS
Sbjct: 568 MGSKRDIEKFTGDNDFGLWKVKMEAVLIQQKCEKALKGEVSLPVTMSRAEKTEMVDKARS 389
Query: 61 AIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVEL 120
AIVLCLG+K LRE KE TA SMW+K LYMTKSLAHRQ LKQQLYSF+MV+SK ++E
Sbjct: 388 AIVLCLGDKVLREVAKERTAASMWAKL*SLYMTKSLAHRQFLKQQLYSFRMVESKAIMEQ 209
Query: 121 LVEFNKIIGDFENIKLHLED-AGALMVWCCL 150
L EFNKI+ D ENI++ LED A+++ C L
Sbjct: 208 LTEFNKILDDLENIEVQLEDEEKAILLLCAL 116
>BG646342 weakly similar to PIR|F84486|F84 probable retroelement pol
polyprotein [imported] - Arabidopsis thaliana, partial
(4%)
Length = 599
Score = 183 bits (464), Expect = 3e-47
Identities = 90/107 (84%), Positives = 97/107 (90%)
Frame = +2
Query: 58 ARSAIVLCLGNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVV 117
ARSAIVLCLG+K LRE KE TATSM +K EYLYMTKSLAHRQ LKQQLYSFKMV+SK +
Sbjct: 2 ARSAIVLCLGDKVLREVAKEPTATSMCAKLEYLYMTKSLAHRQFLKQQLYSFKMVESKAI 181
Query: 118 VELLVEFNKIIGDFENIKLHLEDAGALMVWCCLEDEEGDVSHLGSDA 164
ELLVEFNKIIGD ENI++HLEDAGALMVWCCLEDEEGDVSHLG+DA
Sbjct: 182 TELLVEFNKIIGDLENIEVHLEDAGALMVWCCLEDEEGDVSHLGNDA 322
>BF648150 similar to GP|14586969|gb| pol polyprotein {Citrus x paradisi},
partial (3%)
Length = 658
Score = 39.7 bits (91), Expect = 5e-04
Identities = 16/62 (25%), Positives = 35/62 (55%)
Frame = +2
Query: 79 TATSMWSKFEYLYMTKSLAHRQVLKQQLYSFKMVKSKVVVELLVEFNKIIGDFENIKLHL 138
+A +W K E YM + ++ L ++KMV +K V+E L E +I+ +++ +++
Sbjct: 263 SAKDLWDKLETRYMREDATSKKFLVSHFNNYKMVDNKSVMEQLYEIERILNNYKQHNMNM 442
Query: 139 ED 140
++
Sbjct: 443 DE 448
>BG587684 similar to GP|12858190|dbj evidence:NAS~putative~unclassifiable
{Mus musculus}, partial (14%)
Length = 819
Score = 31.6 bits (70), Expect = 0.15
Identities = 12/18 (66%), Positives = 15/18 (82%)
Frame = +3
Query: 1 MGSKWDIEKFTGDNDFRL 18
M +KWDIE F G+NDFR+
Sbjct: 735 MDTKWDIEIFIGENDFRV 788
>TC80247 similar to PIR|C86476|C86476 protein F15O4.45 [imported] -
Arabidopsis thaliana, partial (29%)
Length = 1225
Score = 27.7 bits (60), Expect = 2.2
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = +2
Query: 67 GNKDLREATKETTATSMWSKFEYLYMTKSLAHRQVL 102
G K ++ + ++S WSKF YL+M+ R +L
Sbjct: 326 GGKSSSSSSFSSKSSSSWSKFVYLFMSAIFRRRGLL 433
>CB065458
Length = 847
Score = 26.9 bits (58), Expect = 3.7
Identities = 8/20 (40%), Positives = 16/20 (80%)
Frame = -3
Query: 5 WDIEKFTGDNDFRLWKVKME 24
W++E TG++ FR+W++ +E
Sbjct: 368 WNLEFLTGNSLFRVWRIYLE 309
>TC79894 similar to GP|21104712|dbj|BAB93301. hypothetical protein {Oryza
sativa (japonica cultivar-group)}, partial (28%)
Length = 1034
Score = 26.6 bits (57), Expect = 4.8
Identities = 16/43 (37%), Positives = 25/43 (57%), Gaps = 5/43 (11%)
Frame = -2
Query: 70 DLREATKETTATSMWSKF-----EYLYMTKSLAHRQVLKQQLY 107
DL ATK+ + +S + K E Y ++ AH+++ KQQLY
Sbjct: 826 DLHLATKDFS*SSNFGKDSEPWKEKPYNHQTFAHKEICKQQLY 698
>TC82232 similar to GP|10177316|dbj|BAB10642. kinesin-like protein
{Arabidopsis thaliana}, partial (21%)
Length = 1181
Score = 26.2 bits (56), Expect = 6.3
Identities = 21/67 (31%), Positives = 31/67 (45%)
Frame = +3
Query: 16 FRLWKVKMEAVLIQQKCEKTLKGESSLPVTMSRAEKTEMVDKARSAIVLCLGNKDLREAT 75
FRLWK E ++Q K E K E ++ ++T+MV R L K L+E
Sbjct: 597 FRLWKASREKEVLQLKKEGR-KNEYERHKLLALNQRTKMV-LQRKTEEASLATKRLKELL 770
Query: 76 KETTATS 82
+ A+S
Sbjct: 771 ESRKASS 791
>TC93539 weakly similar to GP|17063165|gb|AAL32979.1 At1g12970/F13K23_18
{Arabidopsis thaliana}, partial (29%)
Length = 852
Score = 25.8 bits (55), Expect = 8.2
Identities = 10/21 (47%), Positives = 16/21 (75%)
Frame = -1
Query: 88 EYLYMTKSLAHRQVLKQQLYS 108
EY+Y+++ L HR +LK+ L S
Sbjct: 774 EYMYISQPL*HRHLLKENLMS 712
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.317 0.131 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,763,896
Number of Sequences: 36976
Number of extensions: 48424
Number of successful extensions: 248
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 248
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 248
length of query: 164
length of database: 9,014,727
effective HSP length: 89
effective length of query: 75
effective length of database: 5,723,863
effective search space: 429289725
effective search space used: 429289725
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 54 (25.4 bits)
Medicago: description of AC148218.5


