

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147429.6 - phase: 0 /pseudo
(974 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
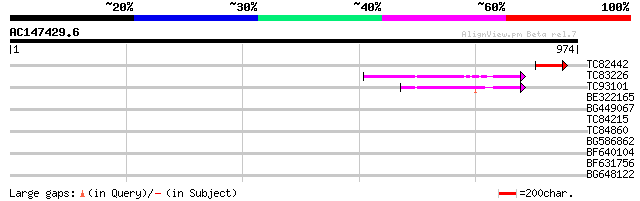
Score E
Sequences producing significant alignments: (bits) Value
TC82442 58 2e-08
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 54 4e-07
TC93101 51 2e-06
BE322165 40 0.004
BG449067 39 0.013
TC84215 weakly similar to GP|17351914|dbj|BAB78588. alcohol acet... 38 0.022
TC84860 37 0.048
BG586862 35 0.11
BF640104 34 0.24
BF631756 32 0.91
BG648122 similar to GP|22535590|dbj hypothetical protein {Oryza ... 29 7.7
>TC82442
Length = 578
Score = 57.8 bits (138), Expect = 2e-08
Identities = 28/56 (50%), Positives = 38/56 (67%), Gaps = 2/56 (3%)
Frame = -3
Query: 904 FEADNESLIQRIKVGI--DVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGNHTAH 957
FE+D E++I+++ GI D+D SYLG L EI ++ FD+C F FIPR GN AH
Sbjct: 168 FESDCEAVIRKLNRGIEKDIDISYLGSVLEEIWKISEKFDMCSFRFIPRSGNRVAH 1
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 53.5 bits (127), Expect = 4e-07
Identities = 77/278 (27%), Positives = 120/278 (42%), Gaps = 1/278 (0%)
Frame = +1
Query: 609 CGCMN*MATTLSKVVTELLGDGRIKIS*ILPPATLRLLFGRKSGDFIPSLDTKTYYEGF* 668
CGC+ + TL +V L GR+ S I +R +G++ G DT+ + *
Sbjct: 4 CGCITQLGFTLLRVDIIL*EPGRLNKSIIHQLVVMRPSYGKRFGHCTLFPDTRCCFGES* 183
Query: 669 ITLFQSGRSSTRGGLTVLYFALDAMLK*KTLHIPF*PVLISLR-FGLVLAWLLDLQTPLT 727
TL Q +G +V LDA K K L + + + +S + FGL L + L L
Sbjct: 184 TTLSQLEVL*EKGEFSVTLSVLDATPKLK-LSLTYSCLALSQKEFGLALTYA*ILTIYLI 360
Query: 728 QTASTGYMKSYPTRTNLPLFKLHLLSIIFGIQETYACSKTNTYLMLTSSSVLKDASRTFT 787
+ YM+ + R N+ K I+G+ A + YL TS S L+ +
Sbjct: 361 PIS*IDYMRLFFRRMNVSPSK*QQSYTIYGMLGI*AFWRIKRYLRWTSFSELR---IVYL 531
Query: 788 VLKKLHLLRSLILELQLHITPIPILALDRSQIDQ*QLQT*LNGRNQLRIWLK*TQMLVFE 847
++ K + R L L L+L +T P L++ + +I NG Q+ W K T M +
Sbjct: 532 IISK-QIPRLLHLWLELAMT--PGLSIVQLRIP--------NGSVQI*DW*KSTLMRTCK 678
Query: 848 LKENGVLV*LFGMTRVWLWLQEPGLPLDLKRLLKQKLL 885
ENGV WLW + G + + L + KL+
Sbjct: 679 TMENGV*ASSSETKLGWLWRRARGKQMAMIEL*RPKLM 792
>TC93101
Length = 675
Score = 51.2 bits (121), Expect = 2e-06
Identities = 70/219 (31%), Positives = 96/219 (42%), Gaps = 5/219 (2%)
Frame = -2
Query: 672 FQSGRSSTRGGLTVLYFALDAMLK*KTLHIPF-*PVLISLRFGLVLAWLLDLQTPLTQTA 730
F S +ST+ L FA D +L K L I F * V FGLVL L L +
Sbjct: 662 FLSEXNSTKEELIAHPFAQDVILIWK-LQITFS*VVREPSEFGLVLNCLFVSLIILLLIS 486
Query: 731 STGYMKSYPTRTNLPLFKLHLLSIIFGIQETYACSKTNTYLMLTSSSVLKDASRTFTVLK 790
GY+ R K L I+FG+Q T SKT++YL + S + LK S + L+
Sbjct: 485 QIGYLMLSLIRQRKL*SKFLQLPIVFGMQGTRLSSKTSSYLRILSFNKLKTVSWLMSKLQ 306
Query: 791 KLHLLRSL----ILELQLHITPIPILALDRSQIDQ*QLQT*LNGRNQLRIWLK*TQMLVF 846
+ L L+LQ+ I + + LDR G+N + K T ML+F
Sbjct: 305 SNPRIPILYYPRYLQLQIRIQLVGVATLDR------------GGKNL*ITF*KPTVMLIF 162
Query: 847 ELKENGVLV*LFGMTRVWLWLQEPGLPLDLKRLLKQKLL 885
K +GV V L M L Q G + L L++KL+
Sbjct: 161 RSKADGVWVVL*EMRMGKLRSQLLGA*MVLIVQLQRKLM 45
>BE322165
Length = 650
Score = 40.0 bits (92), Expect = 0.004
Identities = 18/38 (47%), Positives = 26/38 (68%)
Frame = -3
Query: 896 DYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEI 933
+YGF +VQFE+DNE +IQ + +V+R YLG + I
Sbjct: 117 EYGFSKVQFESDNERVIQMLNGTEEVNRLYLGSIIDSI 4
>BG449067
Length = 578
Score = 38.5 bits (88), Expect = 0.013
Identities = 17/32 (53%), Positives = 24/32 (74%)
Frame = -2
Query: 896 DYGFRQVQFEADNESLIQRIKVGIDVDRSYLG 927
+YGF +VQFE+DNE +IQ + +V+R YLG
Sbjct: 109 EYGFSKVQFESDNERVIQMLNGTEEVNRLYLG 14
>TC84215 weakly similar to GP|17351914|dbj|BAB78588. alcohol
acetyltransferase {Cucumis melo}, partial (14%)
Length = 701
Score = 37.7 bits (86), Expect = 0.022
Identities = 37/119 (31%), Positives = 60/119 (50%), Gaps = 5/119 (4%)
Frame = -3
Query: 753 SIIFGIQETYACSKTNT----YLMLTSSSVLKDASRTF-TVLKKLHLLRSLILELQLHIT 807
SI + ET + ++T+T YL LT S+VLK+A + + L L S+ + L++ ++
Sbjct: 408 SISHCMVETKSINETSTFEACYLKLTLSNVLKEAFMSS*KPTRVLTLTISIAISLRVPLS 229
Query: 808 PIPILALDRSQIDQ*QLQT*LNGRNQLRIWLK*TQMLVFELKENGVLV*LFGMTRVWLW 866
PI LNG++Q ++ + +FE K+ G LV LF +T VW W
Sbjct: 228 -YPIA---------------LNGKSQKKVSS*PIVIQIFESKDGGDLVPLFEITTVW*W 100
>TC84860
Length = 777
Score = 36.6 bits (83), Expect = 0.048
Identities = 30/102 (29%), Positives = 46/102 (44%), Gaps = 8/102 (7%)
Frame = +2
Query: 733 GYMKSYPTRTNLPL-------FKLHLLSIIFGIQE-TYACSKTNTYLMLTSSSVLKDASR 784
G + + P + N P+ L + S + I T C K Y +L VL + SR
Sbjct: 8 GVLLTCPPKRNFPVGIHLFQKLNLRMKSYVLRIYAMTIVCQKHGDYQLL--QQVLMERSR 181
Query: 785 TFTVLKKLHLLRSLILELQLHITPIPILALDRSQIDQ*QLQT 826
FT L L ++L L LH+ +A++ Q+D+ LQT
Sbjct: 182 HFTTLDFLRGFKTLSLMATLHVW*YSDIAVNVKQVDRASLQT 307
>BG586862
Length = 804
Score = 35.4 bits (80), Expect = 0.11
Identities = 51/163 (31%), Positives = 68/163 (41%), Gaps = 10/163 (6%)
Frame = -2
Query: 648 GRKSGDFIPSLDTKTYYEGF*ITLFQSGRSSTRGGLTVLYFALDAMLK*KTLHIPF*PVL 707
GRK G S DTK + I LF S +ST L VL D K*K +I F V
Sbjct: 662 GRKFGGLRLSPDTKVFSGDSYIMLFLSRMNSTNEALGVLCSVPDVSPK*KRFNIFFSIVR 483
Query: 708 ISLRFGLVLAWLLDLQTPLTQTASTGYMKSYPTRTNLPLFKLHLL----------SIIFG 757
R GL P + T +++ Y LP+ L ++ S + G
Sbjct: 482 SLRRSGL---------DPN*GSIFT-HLEYYIFMIGLPILSLKMMKKPSLP*QPCSTVSG 333
Query: 758 IQETYACSKTNTYLMLTSSSVLKDASRTFTVLKKLHLLRSLIL 800
++ET C KT YL + +V + FT K L+SLIL
Sbjct: 332 MRETKRCLKT*MYLEML*FNV---QALVFTASK---WLKSLIL 222
>BF640104
Length = 344
Score = 34.3 bits (77), Expect = 0.24
Identities = 19/45 (42%), Positives = 23/45 (50%)
Frame = -1
Query: 896 DYGFRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNF 940
D GF+ V FE DNE LI + + RSYLG + I L F
Sbjct: 140 DQGFQNVLFENDNEKLISMLSREEEGHRSYLGSIIQSIISLVPRF 6
>BF631756
Length = 622
Score = 32.3 bits (72), Expect = 0.91
Identities = 16/55 (29%), Positives = 33/55 (59%)
Frame = -1
Query: 899 FRQVQFEADNESLIQRIKVGIDVDRSYLGLFLSEIHQLQLNFDICHFSFIPRDGN 953
F++V+FE D+E+L++ ++ +D+ ++Y+G + I F++ FS R N
Sbjct: 172 FQRVEFEGDSETLMKIMRGTLDIPKTYVGNIVFGIKCRLSLFEMATFSHTGRFSN 8
>BG648122 similar to GP|22535590|dbj hypothetical protein {Oryza sativa
(japonica cultivar-group)}, partial (16%)
Length = 590
Score = 29.3 bits (64), Expect = 7.7
Identities = 14/34 (41%), Positives = 24/34 (70%)
Frame = +3
Query: 261 LWVSPTISLKLLCIVSKLLLFMLSLMVNLLRLFP 294
L SP +S+ ++ +V+++LLF S ++ LLRL P
Sbjct: 258 LGFSPWLSISVVLLVARVLLFSHSALLALLRLKP 359
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.363 0.165 0.602
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,359,722
Number of Sequences: 36976
Number of extensions: 474382
Number of successful extensions: 4869
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1570
Number of HSP's successfully gapped in prelim test: 242
Number of HSP's that attempted gapping in prelim test: 3252
Number of HSP's gapped (non-prelim): 1938
length of query: 974
length of database: 9,014,727
effective HSP length: 105
effective length of query: 869
effective length of database: 5,132,247
effective search space: 4459922643
effective search space used: 4459922643
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC147429.6


