

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147179.2 + phase: 0
(185 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
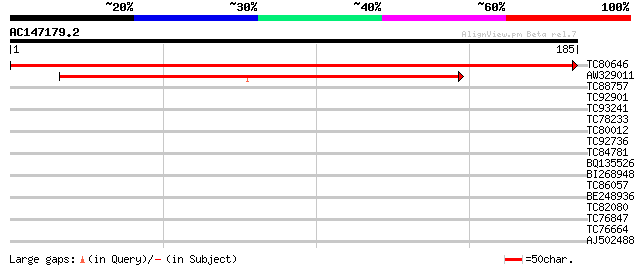
Sequences producing significant alignments: (bits) Value
TC80646 similar to GP|21553926|gb|AAM63009.1 unknown {Arabidopsi... 377 e-105
AW329011 similar to GP|21553926|gb unknown {Arabidopsis thaliana... 206 4e-54
TC88757 similar to GP|4666311|dbj|BAA77218.1 LON protease homolo... 29 1.2
TC92901 similar to GP|23306122|gb|AAN17389.1 Unknown protein {Or... 29 1.2
TC93241 similar to GP|17065634|gb|AAL33811.1 unknown protein {Ar... 28 1.6
TC78233 similar to GP|15809838|gb|AAL06847.1 AT5g51120/MWD22_6 {... 28 2.1
TC80012 GP|2598571|emb|CAA75573.1 putative {Medicago truncatula}... 28 2.7
TC92736 weakly similar to SP|Q9STK9|C71O_ARATH Cytochrome P450 7... 27 3.5
TC84781 similar to GP|15010664|gb|AAK73991.1 AT3g02690/F16B3_32 ... 27 3.5
BQ135526 27 4.6
BI268948 similar to PIR|T49951|T4995 hypothetical protein F8M21.... 27 4.6
TC86057 similar to GP|15146330|gb|AAK83648.1 At1g44800/T12C22_7 ... 27 4.6
BE248936 27 6.0
TC82080 similar to GP|9757854|dbj|BAB08488.1 contains similarity... 27 6.0
TC76847 homologue to SP|P08927|RUBB_PEA RuBisCO subunit binding-... 27 6.0
TC76664 weakly similar to PIR|T08541|T08541 hypothetical protein... 27 6.0
AJ502488 similar to GP|21536488|gb unknown {Arabidopsis thaliana... 26 7.9
>TC80646 similar to GP|21553926|gb|AAM63009.1 unknown {Arabidopsis
thaliana}, partial (86%)
Length = 841
Score = 377 bits (967), Expect = e-105
Identities = 185/185 (100%), Positives = 185/185 (100%)
Frame = +3
Query: 1 MGLSMNNPPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 60
MGLSMNNPPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI
Sbjct: 30 MGLSMNNPPATITPRHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 209
Query: 61 FTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDG 120
FTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDG
Sbjct: 210 FTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDG 389
Query: 121 SVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQQDEDLKDWP 180
SVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQQDEDLKDWP
Sbjct: 390 SVILKLSGGLAILIFCLEWVVLTLAFFLKYYACVEGGNTCRTVVLGSAKVQQDEDLKDWP 569
Query: 181 WPFQV 185
WPFQV
Sbjct: 570 WPFQV 584
>AW329011 similar to GP|21553926|gb unknown {Arabidopsis thaliana}, partial
(66%)
Length = 482
Score = 206 bits (524), Expect = 4e-54
Identities = 106/135 (78%), Positives = 112/135 (82%), Gaps = 3/135 (2%)
Frame = +3
Query: 17 YNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGAN 76
YNIHK FLFCNY+LL A+SSCIFLTLSL L PS+CG F ILLH FTIAGAVS A+
Sbjct: 78 YNIHKAFLFCNYILLSASSSCIFLTLSLNLFPSICGIFFILLHAFTIAGAVSASASSSIT 257
Query: 77 ---RWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREEDGSVILKLSGGLAIL 133
RWYSAHMV TVLTAIFQGSVSVL+FTRT DFL EL+SYVREEDGSVILKL GGL IL
Sbjct: 258 TLTRWYSAHMVVTVLTAIFQGSVSVLIFTRTEDFLVELKSYVREEDGSVILKLCGGLTIL 437
Query: 134 IFCLEWVVLTLAFFL 148
IF LEWVVL LAFFL
Sbjct: 438 IFLLEWVVLILAFFL 482
>TC88757 similar to GP|4666311|dbj|BAA77218.1 LON protease homologue
{Lithospermum erythrorhizon}, partial (84%)
Length = 1104
Score = 28.9 bits (63), Expect = 1.2
Identities = 14/39 (35%), Positives = 19/39 (47%)
Frame = +2
Query: 22 LFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 60
+F FC + L S C TLS + S+ F LI H+
Sbjct: 845 VFFFCFQLYLFNLSKCCLSTLSFVIPSSLASFVLIFCHV 961
>TC92901 similar to GP|23306122|gb|AAN17389.1 Unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (13%)
Length = 555
Score = 28.9 bits (63), Expect = 1.2
Identities = 20/64 (31%), Positives = 35/64 (54%), Gaps = 6/64 (9%)
Frame = +2
Query: 67 VSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTS-----DFLG-ELQSYVREEDG 120
+ GCA + +A M+++ + I+QGS +V F T+ + LG E Q VR+ +
Sbjct: 110 LGGCAIRDSAFASTAPMISSDIAGIWQGSKAVTTFDTTNTGIFRELLGDETQKSVRDGEN 289
Query: 121 SVIL 124
+V+L
Sbjct: 290 NVLL 301
>TC93241 similar to GP|17065634|gb|AAL33811.1 unknown protein {Arabidopsis
thaliana}, partial (5%)
Length = 644
Score = 28.5 bits (62), Expect = 1.6
Identities = 23/73 (31%), Positives = 39/73 (52%)
Frame = -3
Query: 60 IFTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQGSVSVLVFTRTSDFLGELQSYVREED 119
IF++ +V + VGA V+++LTA GS S+L+ T +S +G ++ D
Sbjct: 435 IFSVLVSVLSESLVGAGSSLLTRTVSSILTAEAAGS-SLLIGTGSSTGIG--RAAFSAGD 265
Query: 120 GSVILKLSGGLAI 132
GS +L ++G I
Sbjct: 264 GSTLLTVTGSSII 226
>TC78233 similar to GP|15809838|gb|AAL06847.1 AT5g51120/MWD22_6 {Arabidopsis
thaliana}, partial (66%)
Length = 1183
Score = 28.1 bits (61), Expect = 2.1
Identities = 23/70 (32%), Positives = 32/70 (44%), Gaps = 3/70 (4%)
Frame = -3
Query: 35 SSCIFLTLSLRLVPSVCGF---FLILLHIFTIAGAVSGCAAVGANRWYSAHMVATVLTAI 91
S+ +FL LSL F F LLHI G V GC A +S ++ + +
Sbjct: 326 SAHLFLNLSLHFAKGASFFLDLFQPLLHILQFLGGVIGCEFFIAFLMFSIYV--GIHVNL 153
Query: 92 FQGSVSVLVF 101
F G+VS + F
Sbjct: 152 FIGNVSTVDF 123
>TC80012 GP|2598571|emb|CAA75573.1 putative {Medicago truncatula}, complete
Length = 511
Score = 27.7 bits (60), Expect = 2.7
Identities = 14/38 (36%), Positives = 22/38 (57%)
Frame = -2
Query: 23 FLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHI 60
F+F ++LL FL L + L+P + F L+LLH+
Sbjct: 255 FIFLLFLLLIPNQVLHFLLLYVLLLPKLLRFLLLLLHV 142
>TC92736 weakly similar to SP|Q9STK9|C71O_ARATH Cytochrome P450 71A24 (EC
1.14.-.-). [Mouse-ear cress] {Arabidopsis thaliana},
partial (34%)
Length = 781
Score = 27.3 bits (59), Expect = 3.5
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = -1
Query: 26 CNYVLLGAASSCIFLTLSLRLVPSVCGFFLI 56
C Y +L +SS + + L L+ S C FF+I
Sbjct: 259 CKYFMLPKSSSVMCVLLPATLLTSSCNFFMI 167
>TC84781 similar to GP|15010664|gb|AAK73991.1 AT3g02690/F16B3_32
{Arabidopsis thaliana}, partial (18%)
Length = 558
Score = 27.3 bits (59), Expect = 3.5
Identities = 20/56 (35%), Positives = 24/56 (42%)
Frame = +2
Query: 39 FLTLSLRLVPSVCGFFLILLHIFTIAGAVSGCAAVGANRWYSAHMVATVLTAIFQG 94
F S RL+P+ GF L+ F SG N W S + A V A FQG
Sbjct: 398 FFVASFRLIPA--GFLLVAFAAFRGRALPSGF-----NAWLSIAIFALVDAACFQG 544
>BQ135526
Length = 899
Score = 26.9 bits (58), Expect = 4.6
Identities = 11/45 (24%), Positives = 26/45 (57%)
Frame = -2
Query: 19 IHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTI 63
IHK+ +F NY ++ +F S+++ FF++++ I+++
Sbjct: 457 IHKVVIFDNYCVI*KKYERLFRKFSIQIDKGPINFFVLIVLIYSL 323
>BI268948 similar to PIR|T49951|T4995 hypothetical protein F8M21.50 -
Arabidopsis thaliana, partial (86%)
Length = 723
Score = 26.9 bits (58), Expect = 4.6
Identities = 15/41 (36%), Positives = 21/41 (50%)
Frame = -1
Query: 15 RHYNIHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFL 55
RHY I KL + LL + + LSL+ S+C FF+
Sbjct: 579 RHYYISKLLIIAA*ALLESVEAKSCDNLSLKSSTSLCKFFM 457
>TC86057 similar to GP|15146330|gb|AAK83648.1 At1g44800/T12C22_7
{Arabidopsis thaliana}, partial (80%)
Length = 1476
Score = 26.9 bits (58), Expect = 4.6
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +1
Query: 128 GGLAILIFCLEWVVLTLAFFLKYYACVEGGNTC 160
GG+A LIF ++ V + F + ACV G C
Sbjct: 820 GGIATLIFERDFSVWAIGFDSRLLACVYSGIVC 918
>BE248936
Length = 444
Score = 26.6 bits (57), Expect = 6.0
Identities = 11/31 (35%), Positives = 16/31 (51%)
Frame = +1
Query: 24 LFCNYVLLGAASSCIFLTLSLRLVPSVCGFF 54
+FC + LG S +FL +S + GFF
Sbjct: 352 IFCEFTALGVDFSLLFLIISFEKIMLFFGFF 444
>TC82080 similar to GP|9757854|dbj|BAB08488.1 contains similarity to
apoptosis antagonizing transcription
factor~gene_id:MFB13.10, partial (37%)
Length = 908
Score = 26.6 bits (57), Expect = 6.0
Identities = 18/51 (35%), Positives = 28/51 (54%)
Frame = -2
Query: 19 IHKLFLFCNYVLLGAASSCIFLTLSLRLVPSVCGFFLILLHIFTIAGAVSG 69
IH LFL + ++ ++ + L + L L+ S+ FFL LLH+ I V G
Sbjct: 244 IHILFLKLFHFIVFPSTFLLSL-IFLFLITSLVIFFLFLLHLTIIINLVEG 95
>TC76847 homologue to SP|P08927|RUBB_PEA RuBisCO subunit binding-protein beta
subunit chloroplast precursor (60 kDa chaperonin beta
subunit), partial (96%)
Length = 2096
Score = 26.6 bits (57), Expect = 6.0
Identities = 16/49 (32%), Positives = 24/49 (48%), Gaps = 4/49 (8%)
Frame = +3
Query: 89 TAIFQGSVSVLVFTRTSDFLGEL----QSYVREEDGSVILKLSGGLAIL 133
T + GS V R S ++ Q Y +E+ I KLSGG+A++
Sbjct: 1263 TIVGDGSTQEAVTKRVSQIKNQIEAAEQDYEKEKLNERIAKLSGGVAVI 1409
>TC76664 weakly similar to PIR|T08541|T08541 hypothetical protein F27B13.40 -
Arabidopsis thaliana, partial (38%)
Length = 1860
Score = 26.6 bits (57), Expect = 6.0
Identities = 13/28 (46%), Positives = 15/28 (53%)
Frame = +3
Query: 2 GLSMNNPPATITPRHYNIHKLFLFCNYV 29
G++MNNP AT N F FCN V
Sbjct: 1281 GVAMNNPTATAVTHVLNNKHEFPFCNGV 1364
>AJ502488 similar to GP|21536488|gb unknown {Arabidopsis thaliana}, partial
(25%)
Length = 664
Score = 26.2 bits (56), Expect = 7.9
Identities = 12/40 (30%), Positives = 24/40 (60%), Gaps = 4/40 (10%)
Frame = +3
Query: 122 VILKLSGGLAILIFCLEWVV----LTLAFFLKYYACVEGG 157
V+L +S L +L + +W+V +TL ++L ++ C+ G
Sbjct: 447 VVLIMSAFLGLLGYGFQWLVIQRLITLPYYLVFFLCLIAG 566
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.328 0.142 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,229,984
Number of Sequences: 36976
Number of extensions: 114439
Number of successful extensions: 973
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 969
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 972
length of query: 185
length of database: 9,014,727
effective HSP length: 90
effective length of query: 95
effective length of database: 5,686,887
effective search space: 540254265
effective search space used: 540254265
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC147179.2


