

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146746.4 - phase: 0
(472 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
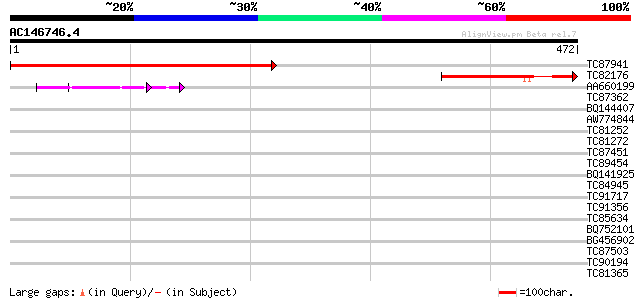
Sequences producing significant alignments: (bits) Value
TC87941 similar to PIR|T05269|T05269 adenylyl cyclase-associated... 436 e-122
TC82176 similar to GP|4126473|dbj|BAA36585.1 adenylyl cyclase as... 138 4e-33
AA660199 61 8e-10
TC87362 weakly similar to GP|21689861|gb|AAM67491.1 unknown prot... 32 0.40
BQ144407 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 32 0.40
AW774844 GP|178014|gb|A acrosin {Homo sapiens}, partial (7%) 32 0.53
TC81252 similar to PIR|T52624|T52624 b-Zip DNA binding protein [... 25 0.53
TC81272 similar to PIR|T05225|T05225 extensin homolog F17I5.160 ... 32 0.69
TC87451 similar to SP|Q42783|BCCP_SOYBN Biotin carboxyl carrier ... 32 0.69
TC89454 similar to GP|15983460|gb|AAL11598.1 At1g55000/F14C21_4 ... 31 1.2
BQ141925 homologue to GP|10727150|gb| Rm62 gene product {alt 1} ... 30 1.5
TC84945 similar to GP|20259551|gb|AAM13895.1 unknown protein {Ar... 30 1.5
TC91717 weakly similar to GP|6691125|gb|AAF24497.1| FH protein N... 30 1.5
TC91356 similar to GP|20151585|gb|AAM11152.1 LD24077p {Drosophil... 30 1.5
TC85634 similar to GP|21555308|gb|AAM63830.1 Photosystem I react... 30 2.0
BQ752101 similar to PIR|S57177|S57 branched-chain-amino-acid tra... 30 2.0
BG456902 30 2.0
TC87503 similar to SP|Q43111|PME3_PHAVU Pectinesterase 3 precurs... 30 2.0
TC90194 similar to GP|19570030|gb|AAL92326.1 hypothetical protei... 30 2.6
TC81365 similar to GP|14579399|gb|AAK69274.1 unknown {Glycine ma... 30 2.6
>TC87941 similar to PIR|T05269|T05269 adenylyl cyclase-associated protein
[imported] - Arabidopsis thaliana, partial (43%)
Length = 906
Score = 436 bits (1120), Expect = e-122
Identities = 221/222 (99%), Positives = 221/222 (99%)
Frame = +2
Query: 1 MDETLIKRLESAVTRLEALSTGIHHSSSASDASDAASDPSVVAFVDLIDQYVSRLSKAAD 60
MDETLIKRLESAVTRLEALSTGIHHSSSASDASDAASDPSVVAFVDLIDQYVSRLSKAAD
Sbjct: 191 MDETLIKRLESAVTRLEALSTGIHHSSSASDASDAASDPSVVAFVDLIDQYVSRLSKAAD 370
Query: 61 IIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGRR 120
IIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGRR
Sbjct: 371 IIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEGRR 550
Query: 121 SDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPN 180
SDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPN
Sbjct: 551 SDFFNHLKAAVDSLSALAWIAFTGKDCGMSMPIAHVEESWQMAEFYSNKVLVEYRNKDPN 730
Query: 181 HVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFAPSKV 222
HVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFA SKV
Sbjct: 731 HVEWVKALKELYLPGLRDYVKSFHPLGPVWSQTGKVFASSKV 856
Score = 31.2 bits (69), Expect = 0.90
Identities = 20/48 (41%), Positives = 23/48 (47%)
Frame = +1
Query: 192 YLPGLRDYVKSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPA 239
+ GL + + SF P K K NASAAPAAP AP PA
Sbjct: 772 WFEGLCEELSSFRPC---MESNWKSICFIKSNASAAPAAPFAPSSSPA 906
>TC82176 similar to GP|4126473|dbj|BAA36585.1 adenylyl cyclase associated
protein {Gossypium hirsutum}, partial (19%)
Length = 696
Score = 138 bits (348), Expect = 4e-33
Identities = 81/128 (63%), Positives = 86/128 (66%), Gaps = 15/128 (11%)
Frame = +1
Query: 360 KVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGCQIYLSKDSL 419
KVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGCQIYLSKD +
Sbjct: 1 KVNNITIDNCKKTGVVFKDVVAAFEVVNSNGVEVQCQGSAPTISVDNTSGCQIYLSKDFI 180
Query: 420 ETSISTAK-------SSEI--------NVLVPNVESDGDWVEHSLPQQYIHLFKEGRFET 464
+K SS+ N+L N +LFKEGRFET
Sbjct: 181 RNIHIYSKVQ*DQCISSKC*VWMVIGWNILYHN--------------STFNLFKEGRFET 318
Query: 465 TPASHSGG 472
TPASHSGG
Sbjct: 319 TPASHSGG 342
>AA660199
Length = 434
Score = 61.2 bits (147), Expect = 8e-10
Identities = 48/97 (49%), Positives = 54/97 (55%), Gaps = 1/97 (1%)
Frame = +2
Query: 23 IHHSSSASDASDAASD-PSVVAFVDLIDQYVSRLSKAADIIGGQVLDVTNRVKEAFSVQK 81
IHHSSSASDA D PSVVAFVDLI S + I+ DVTNRVKEAF QK
Sbjct: 2 IHHSSSASDAXDDGFPIPSVVAFVDLIXN-TS*XFRKLPILLVDGADVTNRVKEAFLXQK 178
Query: 82 ELLIKLKQTQKPDPAGLAEFLKPLNEVIMKASSLTEG 118
E + + + P + LNEV MKA LTEG
Sbjct: 179 EX*LXSNKLRN-XPCWAGGISETLNEVFMKA-LLTEG 283
Score = 49.3 bits (116), Expect = 3e-06
Identities = 41/96 (42%), Positives = 52/96 (53%)
Frame = +1
Query: 50 QYVSRLSKAADIIGGQVLDVTNRVKEAFSVQKELLIKLKQTQKPDPAGLAEFLKPLNEVI 109
QYV LSKAADIIGG+ K +FSV + LIK KQTQK G F E
Sbjct: 85 QYVIGLSKAADIIGGR-CGCHQ*GKRSFSVXERXLIKXKQTQKXTLLGWRNF*N--XE*S 255
Query: 110 MKASSLTEGRRSDFFNHLKAAVDSLSALAWIAFTGK 145
+ SSL + RRS FFNH + +S+ +W+ G+
Sbjct: 256 VHESSL-D*RRSLFFNH*RXV--EVSSSSWLHLLGR 354
>TC87362 weakly similar to GP|21689861|gb|AAM67491.1 unknown protein
{Arabidopsis thaliana}, partial (46%)
Length = 1996
Score = 32.3 bits (72), Expect = 0.40
Identities = 14/39 (35%), Positives = 25/39 (63%), Gaps = 4/39 (10%)
Frame = +1
Query: 219 PSKVNASAAPAA----PSAPPPPPASLFSSESSQASSSK 253
PS+ + ++ P + P+ PPPPP +++ +S ASS+K
Sbjct: 232 PSRSSPTSCPVSHP*PPNPPPPPPTPNYTTSNSHASSTK 348
>BQ144407 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (23%)
Length = 1217
Score = 32.3 bits (72), Expect = 0.40
Identities = 19/52 (36%), Positives = 24/52 (45%)
Frame = -3
Query: 204 HPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPK 255
+P+ P S + APS +S P S PPP P S S S S S P+
Sbjct: 645 YPISPPPSLDISITAPSTRPSSLTPTPASPPPPIPRSPHPSSFSPHSRSPPR 490
>AW774844 GP|178014|gb|A acrosin {Homo sapiens}, partial (7%)
Length = 367
Score = 32.0 bits (71), Expect = 0.53
Identities = 23/74 (31%), Positives = 32/74 (43%), Gaps = 2/74 (2%)
Frame = -1
Query: 228 PAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRKVTDD--MKTKN 285
P +P PPPPPAS S S+ +KP + + T T VT+ +T+
Sbjct: 361 PPSPPPPPPPPASRSCSPSTSLKPTKPSSAPGSTL--VTTAGST*NR*AVTNGGCTRTRG 188
Query: 286 RADRSGVVGNSVKE 299
R S +G S E
Sbjct: 187 RER*SSEIGESESE 146
>TC81252 similar to PIR|T52624|T52624 b-Zip DNA binding protein [imported] -
Arabidopsis thaliana, partial (48%)
Length = 1221
Score = 25.4 bits (54), Expect(2) = 0.53
Identities = 12/39 (30%), Positives = 19/39 (47%)
Frame = +3
Query: 201 KSFHPLGPVWSQTGKVFAPSKVNASAAPAAPSAPPPPPA 239
K F W K+ +S+A A+ ++PPPPP+
Sbjct: 30 KKFREEENKWRHKCKIPMHRLPTSSSAAASTASPPPPPS 146
Score = 25.0 bits (53), Expect(2) = 0.53
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = +3
Query: 231 PSAPPPPPASLFSSESSQASS 251
P+A PPPPAS S + +SS
Sbjct: 243 PAADPPPPASTRSDPKTISSS 305
>TC81272 similar to PIR|T05225|T05225 extensin homolog F17I5.160 -
Arabidopsis thaliana, partial (3%)
Length = 757
Score = 31.6 bits (70), Expect = 0.69
Identities = 24/60 (40%), Positives = 30/60 (50%), Gaps = 14/60 (23%)
Frame = +1
Query: 228 PAAPSAPPPPPASL------FSSESSQASSSKPKVGMSA------VFQEIGTG--NVTAG 273
PA P++PPP P SL S SS ASSS K G + +F +G G NV+ G
Sbjct: 430 PAPPTSPPPAPVSLSPASAPLPSASSPASSSFSKGGCKSRGRFPFLFSRLGHGLVNVSNG 609
>TC87451 similar to SP|Q42783|BCCP_SOYBN Biotin carboxyl carrier protein of
acetyl-CoA carboxylase chloroplast precursor (BCCP).
[Soybean], partial (74%)
Length = 1282
Score = 31.6 bits (70), Expect = 0.69
Identities = 28/99 (28%), Positives = 40/99 (40%), Gaps = 10/99 (10%)
Frame = +1
Query: 211 SQTGKVFAPSKVNASAAPAAPSAPPP--------PPASLFSSESSQASSSKPKVGMSAVF 262
SQT APS A AP+ P +PPP PP + + S + + V
Sbjct: 583 SQTAPPVAPSHTLALPAPSTPISPPPVNLTTSSRPPLKSPMAGTFYRSPAPGEPPFVKVG 762
Query: 263 QEIGTGNVTAGLRKVTDDMKTKN--RADRSGVVGNSVKE 299
E+ G V + + MK N AD+SG + + E
Sbjct: 763 DEVKKGQVVC----IIEAMKLMNEIEADKSGTIVEIIAE 867
>TC89454 similar to GP|15983460|gb|AAL11598.1 At1g55000/F14C21_4
{Arabidopsis thaliana}, partial (85%)
Length = 854
Score = 30.8 bits (68), Expect = 1.2
Identities = 12/44 (27%), Positives = 26/44 (58%)
Frame = +1
Query: 222 VNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEI 265
+N+S++ ++P PPPP +++ S +S S+ + +FQ +
Sbjct: 55 MNSSSSTSSPPPPPPPTSTIISPMNSHFSALSSTDTLQIIFQNL 186
>BQ141925 homologue to GP|10727150|gb| Rm62 gene product {alt 1} [Drosophila
melanogaster], partial (3%)
Length = 1059
Score = 30.4 bits (67), Expect = 1.5
Identities = 13/24 (54%), Positives = 16/24 (66%)
Frame = +2
Query: 228 PAAPSAPPPPPASLFSSESSQASS 251
P AP PPPPP+ LFS+ S+ S
Sbjct: 17 PPAPPPPPPPPSFLFSNPFSKFPS 88
>TC84945 similar to GP|20259551|gb|AAM13895.1 unknown protein {Arabidopsis
thaliana}, partial (18%)
Length = 785
Score = 30.4 bits (67), Expect = 1.5
Identities = 12/19 (63%), Positives = 13/19 (68%)
Frame = +1
Query: 223 NASAAPAAPSAPPPPPASL 241
N+SA P P PPPPP SL
Sbjct: 88 NSSATPPLPPPPPPPPQSL 144
>TC91717 weakly similar to GP|6691125|gb|AAF24497.1| FH protein NFH2
{Nicotiana tabacum}, partial (6%)
Length = 477
Score = 30.4 bits (67), Expect = 1.5
Identities = 20/59 (33%), Positives = 27/59 (44%), Gaps = 14/59 (23%)
Frame = +2
Query: 228 PAAPSAPPPPPASLFSSESSQASS------SKPK--------VGMSAVFQEIGTGNVTA 272
P PS PPPP S F+S + SS KPK V ++AV + G ++A
Sbjct: 296 PTYPSTPPPPSPSSFASFPANISSLTIPQTQKPKSSSSKLLAVAITAVIAAVAVGAISA 472
>TC91356 similar to GP|20151585|gb|AAM11152.1 LD24077p {Drosophila
melanogaster}, partial (71%)
Length = 698
Score = 30.4 bits (67), Expect = 1.5
Identities = 14/36 (38%), Positives = 20/36 (54%)
Frame = +3
Query: 219 PSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKP 254
PS +S PA P + PPPP+S+ SS + + P
Sbjct: 582 PSPSASSPPPAVPVSSPPPPSSVSSSSLVSRTPTPP 689
>TC85634 similar to GP|21555308|gb|AAM63830.1 Photosystem I reaction center
subunit IV B chloroplast precursor (PSI-E B)
{Arabidopsis thaliana}, partial (67%)
Length = 752
Score = 30.0 bits (66), Expect = 2.0
Identities = 24/67 (35%), Positives = 30/67 (43%)
Frame = +3
Query: 218 APSKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGNVTAGLRKV 277
AP+ A AAPAA +AP P P SK K+ + GTG+V A V
Sbjct: 279 APAAPAAPAAPAADAAPTPKPK---PPPIGPKRGSKVKILRKESYWYKGTGSVVA----V 437
Query: 278 TDDMKTK 284
D KT+
Sbjct: 438 DQDPKTR 458
>BQ752101 similar to PIR|S57177|S57 branched-chain-amino-acid transaminase
(EC 2.6.1.42) BAT2 cytosolic - yeast, partial (46%)
Length = 720
Score = 30.0 bits (66), Expect = 2.0
Identities = 17/45 (37%), Positives = 26/45 (57%), Gaps = 1/45 (2%)
Frame = -1
Query: 218 APSKVNASAAPAAPS-APPPPPASLFSSESSQASSSKPKVGMSAV 261
AP + +ASA PA P PPP++ + +S+ S S+P SA+
Sbjct: 711 APKRPSASALPAPPCFTSSPPPSAPTTLPASKPSPSRPPTTPSAL 577
>BG456902
Length = 680
Score = 30.0 bits (66), Expect = 2.0
Identities = 20/60 (33%), Positives = 30/60 (49%), Gaps = 4/60 (6%)
Frame = +1
Query: 211 SQTGKVFAPSKVNASAAPAAPSAPPPPPASLFSSESSQAS----SSKPKVGMSAVFQEIG 266
S T K P++ +A +PAA PP P + +E+S A+ S K K+ SA + G
Sbjct: 157 STTPKKTPPARKSAGESPAAKRTPPSKPKNAKKTEASPATDPDQSQKKKLLASATTAKSG 336
>TC87503 similar to SP|Q43111|PME3_PHAVU Pectinesterase 3 precursor (EC
3.1.1.11) (Pectin methylesterase 3) (PE 3)., partial
(93%)
Length = 2088
Score = 30.0 bits (66), Expect = 2.0
Identities = 21/54 (38%), Positives = 29/54 (52%), Gaps = 7/54 (12%)
Frame = -2
Query: 208 PVWSQTGKVFAPSKVNASAAPAAPS-------APPPPPASLFSSESSQASSSKP 254
P+ S T ++ A S + ++ P S +PPP PA LF S SQA+SS P
Sbjct: 986 PLSSHTPQISADSSSSPTSTPRLLSYTVHFHPSPPPHPA-LFHSPPSQAASSLP 828
>TC90194 similar to GP|19570030|gb|AAL92326.1 hypothetical protein
{Dictyostelium discoideum}, partial (3%)
Length = 775
Score = 29.6 bits (65), Expect = 2.6
Identities = 16/50 (32%), Positives = 26/50 (52%)
Frame = +3
Query: 220 SKVNASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSAVFQEIGTGN 269
S + ++ ++PS+ PPP +S S+ SS+ +V V GTGN
Sbjct: 396 SNQRSRSSQSSPSSQPPPASSASSTMVPHTFSSRIRVNP*PVSTSTGTGN 545
>TC81365 similar to GP|14579399|gb|AAK69274.1 unknown {Glycine max}, partial
(38%)
Length = 920
Score = 29.6 bits (65), Expect = 2.6
Identities = 16/38 (42%), Positives = 21/38 (55%)
Frame = -1
Query: 223 NASAAPAAPSAPPPPPASLFSSESSQASSSKPKVGMSA 260
++S+ P P PP P +S SSESS SSS S+
Sbjct: 401 SSSSCPFEP*PPPSPDSSSSSSESSSHSSSSSSSSSSS 288
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.128 0.363
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,246,117
Number of Sequences: 36976
Number of extensions: 181524
Number of successful extensions: 2992
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 1779
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2531
length of query: 472
length of database: 9,014,727
effective HSP length: 100
effective length of query: 372
effective length of database: 5,317,127
effective search space: 1977971244
effective search space used: 1977971244
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146746.4


