

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
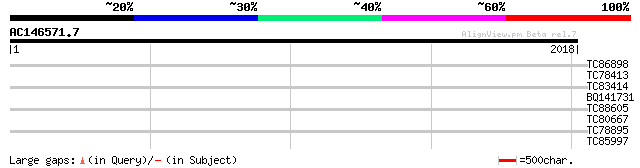
Query= AC146571.7 + phase: 0 /pseudo
(2018 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86898 similar to GP|18376007|emb|CAB91741. probable ATP citrat... 36 0.17
TC78413 similar to PIR|T06329|T06329 symbiotic ammonium transpor... 35 0.29
TC83414 weakly similar to GP|17381206|gb|AAL36415.1 unknown prot... 32 1.9
BQ141731 homologue to PIR|T05444|T054 hypothetical protein F7K2.... 31 4.3
TC88605 similar to SP|Q9SEE4|PIRL_LYCES Pirin-like protein. [Tom... 30 7.3
TC80667 similar to GP|21618284|gb|AAM67334.1 unknown {Arabidopsi... 30 7.3
TC78895 similar to GP|6056390|gb|AAF02854.1| Unknown protein {Ar... 30 7.3
TC85997 similar to PIR|JN0505|JN0505 protein kinase (EC 2.7.1.37... 30 9.5
>TC86898 similar to GP|18376007|emb|CAB91741. probable ATP citrate lyase
subunit 2 {Neurospora crassa}, complete
Length = 2389
Score = 35.8 bits (81), Expect = 0.17
Identities = 32/90 (35%), Positives = 37/90 (40%)
Frame = +2
Query: 1293 TSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHVGGRIVLNLWRLIRGEVKLN 1352
+S S P P TTS C R+ I TH GV VG LI V L+
Sbjct: 950 SSSPSFPTPKTPSTTSTSCPVRDGDWILF----THEGGVDVGDVDAKAEKLLI--PVDLS 1111
Query: 1353 LYSVEAVAEAVLRRKVPSLNHNVLTKWFSR 1382
Y A L +KVP HNVL + SR
Sbjct: 1112 QYPSNEEIAATLLKKVPKGVHNVLVDFISR 1201
>TC78413 similar to PIR|T06329|T06329 symbiotic ammonium transport protein
SAT1 - soybean, partial (30%)
Length = 1397
Score = 35.0 bits (79), Expect = 0.29
Identities = 24/73 (32%), Positives = 37/73 (49%), Gaps = 2/73 (2%)
Frame = -1
Query: 916 NLLCKTVGSEPIVDDLKSNHMKLTKVVIGNSSLVDKNLKSHLSLPTFPNN--LHLDEDDE 973
++ C + + D K++ K+TKV +GN LV+K L HL F +H+ E +
Sbjct: 326 SIFCWVFRNSFLKDKKKNSFQKITKVDVGNFPLVEKYLLCHLKETLFHQTSFVHIVETNH 147
Query: 974 MPGNALDVFLPIS 986
N L FLPI+
Sbjct: 146 ---NPLFPFLPIA 117
>TC83414 weakly similar to GP|17381206|gb|AAL36415.1 unknown protein
{Arabidopsis thaliana}, partial (28%)
Length = 1088
Score = 32.3 bits (72), Expect = 1.9
Identities = 29/117 (24%), Positives = 46/117 (38%)
Frame = -3
Query: 925 EPIVDDLKSNHMKLTKVVIGNSSLVDKNLKSHLSLPTFPNNLHLDEDDEMPGNALDVFLP 984
EP++ L +K T V + L+ + + HL +PT L +D+ LD
Sbjct: 477 EPLMKKLSFTGLKFTHV----T*LL*EKTRRHLLIPTCHRRTVLSTEDDSKNPDLD---- 322
Query: 985 ISARNSQKQKKPWNKCVTIETPRSSGTKGVSTYYQNDGSHLYLLTPNILPPSAGSVQ 1041
AR+ PWN C R ++ +D + +LT L S G+ Q
Sbjct: 321 -HARSKMSSLCPWNSCKGFTNMRKVFGGMITQVEADDFLNFSVLTNLSLSTSRGATQ 154
>BQ141731 homologue to PIR|T05444|T054 hypothetical protein F7K2.80 -
Arabidopsis thaliana, partial (4%)
Length = 1306
Score = 31.2 bits (69), Expect = 4.3
Identities = 18/56 (32%), Positives = 31/56 (55%)
Frame = +1
Query: 799 REKNTKDRTTYEKHAASDAYTPNPSLDTHLRTDDDHKVRAPERCQETDSVASGSRQ 854
R+ KDR +EKH +D TP+P+ T+ T ++ E+ ++T ++G RQ
Sbjct: 1024 RKTPRKDRRQHEKHERND--TPSPTTPTNHDTKPEN--NGDEKSRDTHKESAGQRQ 1179
>TC88605 similar to SP|Q9SEE4|PIRL_LYCES Pirin-like protein. [Tomato]
{Lycopersicon esculentum}, partial (93%)
Length = 1321
Score = 30.4 bits (67), Expect = 7.3
Identities = 20/61 (32%), Positives = 32/61 (51%), Gaps = 6/61 (9%)
Frame = -2
Query: 1224 VLNFTDEKHLLKEFTKIVSS------SDPDILMGWEIQGSSLGFLAERASHLGFGLLNDL 1277
V+NF ++KHLL + ++S + + L W QG+ L FL ++ + L LL L
Sbjct: 711 VMNFVNQKHLLLQLCMQMNSITQE*FDELEFLASW*NQGTWLVFLCKQVNSLPLILLLSL 532
Query: 1278 S 1278
S
Sbjct: 531 S 529
>TC80667 similar to GP|21618284|gb|AAM67334.1 unknown {Arabidopsis thaliana},
partial (16%)
Length = 1150
Score = 30.4 bits (67), Expect = 7.3
Identities = 27/90 (30%), Positives = 42/90 (46%), Gaps = 5/90 (5%)
Frame = +2
Query: 1277 LSRTPSNSWINSQDIKTSEKSILEPDIPDTTSLDCCARESSIIEDEWGRTHASGVHV--- 1333
L +T S IN + T K+ + ++PD +R+SS E E R SGVH
Sbjct: 845 LKQTQSQKNINLVQL-TKSKTRVSINLPDNEISMLRSRQSSFSEHEKERASTSGVHYYKH 1021
Query: 1334 --GGRIVLNLWRLIRGEVKLNLYSVEAVAE 1361
GR ++ + R + ++ L L V+A E
Sbjct: 1022 VKSGRTLVEVLREVDADI-LGLQDVKAEEE 1108
>TC78895 similar to GP|6056390|gb|AAF02854.1| Unknown protein {Arabidopsis
thaliana}, partial (28%)
Length = 2089
Score = 30.4 bits (67), Expect = 7.3
Identities = 19/69 (27%), Positives = 32/69 (45%), Gaps = 7/69 (10%)
Frame = +3
Query: 938 LTKVVIGNSSLVDKNLK---SHLSLPTFPNNL----HLDEDDEMPGNALDVFLPISARNS 990
L K V ++++++ K +HL P P+ H D+DDE P + + + +
Sbjct: 294 LVKAVATAATIIEQPPKFNNNHLEEPLNPHPFMLHHHYDDDDERPLDESEKLRRLRISKA 473
Query: 991 QKQKKPWNK 999
K PWNK
Sbjct: 474 NKGNTPWNK 500
>TC85997 similar to PIR|JN0505|JN0505 protein kinase (EC 2.7.1.37) 5 -
Arabidopsis thaliana, partial (78%)
Length = 2199
Score = 30.0 bits (66), Expect = 9.5
Identities = 27/136 (19%), Positives = 59/136 (42%)
Frame = +1
Query: 475 KMLEKANMAYESESQQECQDILDSIDDMLALDLPKKKPSRSFDHNCPIEASSDNMIPQVD 534
K L+ ++++ + + E +D+ S+ + ++ + S+ F +P
Sbjct: 271 KDLKFSSLSLQKTGKSEPEDLSKSVQNTSKQNIGEIYESKKFG------------VPSQK 414
Query: 535 GSNDDEFSSPCASLAEISSAVEINSEYKRASENHLLHNTDTSTVNTDKRNRQWGSLPFSM 594
GS+ D SS S A SS V +N+ + S+++ DK+ ++GS+ S
Sbjct: 415 GSSADSLSSKFGSGAS-SSVVNVNT------------GSQESSIDQDKKTSEYGSVKNSS 555
Query: 595 TGKVNNDGEHATSQVA 610
+DG + ++ +
Sbjct: 556 VSAKASDGASSIAKTS 603
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.318 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,996,647
Number of Sequences: 36976
Number of extensions: 941095
Number of successful extensions: 4460
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 4381
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4458
length of query: 2018
length of database: 9,014,727
effective HSP length: 111
effective length of query: 1907
effective length of database: 4,910,391
effective search space: 9364115637
effective search space used: 9364115637
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146571.7


