

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146550.5 - phase: 0
(337 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
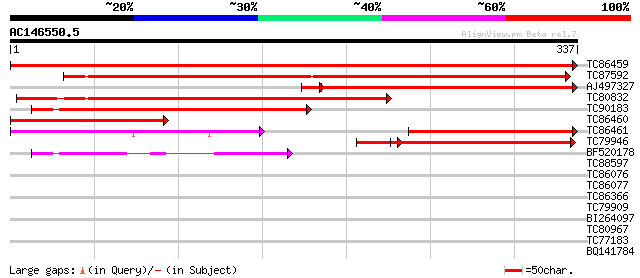
Score E
Sequences producing significant alignments: (bits) Value
TC86459 similar to GP|20257477|gb|AAM15908.1 purple acid phospha... 608 e-175
TC87592 similar to GP|7331193|gb|AAF60315.1| putative purple aci... 398 e-111
AJ497327 weakly similar to GP|20257479|gb purple acid phosphatas... 318 7e-91
TC80832 similar to GP|7331195|gb|AAF60316.1| putative purple aci... 281 3e-76
TC90183 similar to GP|7331197|gb|AAF60317.1| putative purple aci... 214 3e-56
TC86460 similar to GP|20257477|gb|AAM15908.1 purple acid phospha... 196 1e-50
TC86461 similar to GP|20257477|gb|AAM15908.1 purple acid phospha... 171 3e-43
TC79946 similar to GP|21536860|gb|AAM61192.1 acid phosphatase ty... 131 7e-34
BF520178 similar to GP|7331197|gb|A putative purple acid phospha... 103 7e-23
TC88597 weakly similar to PIR|B96700|B96700 protein F12A21.17 [i... 31 0.59
TC86076 similar to GP|7208777|emb|CAB76911.1 putative PTS protei... 31 0.77
TC86077 similar to GP|7208777|emb|CAB76911.1 putative PTS protei... 30 1.7
TC86366 similar to PIR|T01857|T01857 hypothetical protein F9D12.... 28 5.0
TC79909 similar to GP|8101442|gb|AAF72555.1| cryptochrome 1 {Lyc... 28 5.0
BI264097 similar to GP|19570986|dbj diphosphonucleotide phosphat... 28 6.5
TC80967 weakly similar to GP|21593303|gb|AAM65252.1 unknown {Ara... 28 6.5
TC77183 homologue to SP|P43282|METM_LYCES S-adenosylmethionine s... 28 6.5
BQ141784 homologue to GP|23172168|gb| CG13615-PA {Drosophila mel... 27 8.5
>TC86459 similar to GP|20257477|gb|AAM15908.1 purple acid phosphatase
{Arabidopsis thaliana}, partial (80%)
Length = 1205
Score = 608 bits (1569), Expect = e-175
Identities = 286/337 (84%), Positives = 313/337 (92%)
Frame = +3
Query: 1 MAWCLNQHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG 60
MAWCLNQ MLSSVI +SV LCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG
Sbjct: 36 MAWCLNQRMLSSVIFLLSVSLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG 215
Query: 61 NYNQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIW 120
NYNQSLVAHQMGIVG+ LNI+FV+STGDNFY+DGLKGVDDPAFYESF +IYTAPSLQ++W
Sbjct: 216 NYNQSLVAHQMGIVGEKLNIDFVISTGDNFYEDGLKGVDDPAFYESFANIYTAPSLQKVW 395
Query: 121 YNVLGNHDYRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPG 180
YNVLGNHDYRGDVEAQLSPILR KD+RWVCLRSFILD VEFFFVDTTPFVE+YFTDP
Sbjct: 396 YNVLGNHDYRGDVEAQLSPILRLKDSRWVCLRSFILDGGIVEFFFVDTTPFVEKYFTDPE 575
Query: 181 KHTYDWKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLL 240
+HTYDW GVLP ESYRA+LLK+V+S+LVQS AKWKIVV HH IK+AG HGNTQELEEQLL
Sbjct: 576 EHTYDWNGVLPRESYRAKLLKDVNSSLVQSKAKWKIVVGHHTIKTAGHHGNTQELEEQLL 755
Query: 241 PILKSNNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQ 300
PILKSNN+DAYINGHDHCLEHIIDKESGI FFTSGGGSKAW GDI+PWDPKELKLYHDGQ
Sbjct: 756 PILKSNNIDAYINGHDHCLEHIIDKESGIPFFTSGGGSKAWRGDIRPWDPKELKLYHDGQ 935
Query: 301 GFVSMQISKTNASIVFYDVFGKVLHTWSMSKKLKEAA 337
GF+S+QI++ NA IVFYDVFGKVLH W+++K++ AA
Sbjct: 936 GFMSVQITENNADIVFYDVFGKVLHRWNITKEMSAAA 1046
>TC87592 similar to GP|7331193|gb|AAF60315.1| putative purple acid
phosphatase precursor {Ipomoea batatas}, partial (87%)
Length = 1247
Score = 398 bits (1022), Expect = e-111
Identities = 195/301 (64%), Positives = 237/301 (77%)
Frame = +3
Query: 33 KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGKNLNIEFVVSTGDNFYD 92
+L RF H K SLNFLV+GDWGR+G YNQS VA QMG+VG+ L+I+FVVSTGDNFYD
Sbjct: 111 QLQRFTHTPKTDG-SLNFLVIGDWGRRGKYNQSEVAFQMGVVGEKLDIDFVVSTGDNFYD 287
Query: 93 DGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGDVEAQLSPILRQKDNRWVCLR 152
DGL G DP F ESF IY A SLQ+ WY+VLGNHDYRGDVEAQLSP LRQ D+RW+CLR
Sbjct: 288 DGLTGEHDPNFEESFSKIYAAKSLQKQWYSVLGNHDYRGDVEAQLSPFLRQIDSRWLCLR 467
Query: 153 SFILDTDNVEFFFVDTTPFVEEYFTDPGKHTYDWKGVLPLESYRAELLKEVDSALVQSTA 212
SFI+D++ VE FFVDTTPFVEEYFT P +H YDW GV P ++Y A LLK+V+ AL +STA
Sbjct: 468 SFIVDSELVEIFFVDTTPFVEEYFTVP-EHHYDWNGVNPPQTYIANLLKDVEMALRESTA 644
Query: 213 KWKIVVAHHPIKSAGPHGNTQELEEQLLPILKSNNVDAYINGHDHCLEHIIDKESGIHFF 272
KWKIVV HH I+SAG HG+T+EL LLPIL++N+VD Y+NGHDHCLEHI D S I F
Sbjct: 645 KWKIVVGHHAIRSAGHHGDTKELVNLLLPILQANHVDFYMNGHDHCLEHISDLSSPIQFL 824
Query: 273 TSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSMQISKTNASIVFYDVFGKVLHTWSMSKK 332
SG GSKAW GD++ + ++DGQGF+S+Q+++T+A+IVFYDV G+VLH + SK+
Sbjct: 825 VSGAGSKAWRGDVRETTGNGVNFFYDGQGFMSVQLTRTDANIVFYDVSGQVLHKTTSSKQ 1004
Query: 333 L 333
L
Sbjct: 1005L 1007
>AJ497327 weakly similar to GP|20257479|gb purple acid phosphatase
{Arabidopsis thaliana}, partial (35%)
Length = 682
Score = 318 bits (815), Expect(2) = 7e-91
Identities = 152/152 (100%), Positives = 152/152 (100%)
Frame = +1
Query: 186 WKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKS 245
WKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKS
Sbjct: 37 WKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAGPHGNTQELEEQLLPILKS 216
Query: 246 NNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSM 305
NNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSM
Sbjct: 217 NNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQGFVSM 396
Query: 306 QISKTNASIVFYDVFGKVLHTWSMSKKLKEAA 337
QISKTNASIVFYDVFGKVLHTWSMSKKLKEAA
Sbjct: 397 QISKTNASIVFYDVFGKVLHTWSMSKKLKEAA 492
Score = 33.9 bits (76), Expect(2) = 7e-91
Identities = 13/14 (92%), Positives = 13/14 (92%)
Frame = +2
Query: 174 EYFTDPGKHTYDWK 187
EYFTDPGKHTYD K
Sbjct: 2 EYFTDPGKHTYDGK 43
>TC80832 similar to GP|7331195|gb|AAF60316.1| putative purple acid
phosphatase precursor {Glycine max}, partial (58%)
Length = 686
Score = 281 bits (719), Expect = 3e-76
Identities = 132/223 (59%), Positives = 170/223 (76%)
Frame = +1
Query: 5 LNQHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQ 64
+ +L V+ ++ CL+ + + +L FEH KP SL+FLV+GDWGR+G YNQ
Sbjct: 31 ITMSLLVLVVFIATITQCLVYSSA----ELQSFEHAPKPDG-SLSFLVIGDWGRRGGYNQ 195
Query: 65 SLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVL 124
S VA QMG +G+ L+I+FV+STGDNFYD+GLKG+DD +F+ SF IYTAPSLQ+ WYNVL
Sbjct: 196 SQVALQMGYIGEQLDIDFVISTGDNFYDNGLKGIDDTSFHHSFTKIYTAPSLQKQWYNVL 375
Query: 125 GNHDYRGDVEAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDPGKHTY 184
GNHDYRG+VEAQLSP+L DNRW C RS++++T+ VEFFFVDTTPFV++YFT+P H Y
Sbjct: 376 GNHDYRGNVEAQLSPVLTNLDNRWFCSRSYVVNTEFVEFFFVDTTPFVDKYFTEPEDHVY 555
Query: 185 DWKGVLPLESYRAELLKEVDSALVQSTAKWKIVVAHHPIKSAG 227
DW+G P + Y + LLK++D AL QS AKWKIVV HH I+SAG
Sbjct: 556 DWRGTWPRKQYISNLLKDLDLALKQSNAKWKIVVGHHTIRSAG 684
>TC90183 similar to GP|7331197|gb|AAF60317.1| putative purple acid
phosphatase precursor {Phaseolus vulgaris}, partial
(41%)
Length = 595
Score = 214 bits (546), Expect = 3e-56
Identities = 106/166 (63%), Positives = 123/166 (73%)
Frame = +3
Query: 14 IIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGI 73
I V V++C+ N ++ +L R EH +L+FLV+GDWGRKG YNQS VA QMG
Sbjct: 99 ICVVVVVVCVSVN---VSAELQRIEHPAVKADATLSFLVIGDWGRKGTYNQSQVAFQMGR 269
Query: 74 VGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGDV 133
V LNI+FVVSTGDNFYDDGL GV DPAF SF DIYTA SLQ+ WYNVLGNHDYRGDV
Sbjct: 270 VADKLNIDFVVSTGDNFYDDGLTGVHDPAFQYSFSDIYTANSLQKQWYNVLGNHDYRGDV 449
Query: 134 EAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDTTPFVEEYFTDP 179
EAQLSP L+ D+RW C RSF + T+ EFFFVDTTPFV++YF P
Sbjct: 450 EAQLSPFLQNIDHRWFCQRSFFVHTEIAEFFFVDTTPFVDKYFLKP 587
>TC86460 similar to GP|20257477|gb|AAM15908.1 purple acid phosphatase
{Arabidopsis thaliana}, partial (14%)
Length = 349
Score = 196 bits (497), Expect = 1e-50
Identities = 94/94 (100%), Positives = 94/94 (100%)
Frame = +3
Query: 1 MAWCLNQHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG 60
MAWCLNQHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG
Sbjct: 66 MAWCLNQHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG 245
Query: 61 NYNQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDG 94
NYNQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDG
Sbjct: 246 NYNQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDG 347
>TC86461 similar to GP|20257477|gb|AAM15908.1 purple acid phosphatase
{Arabidopsis thaliana}, partial (28%)
Length = 636
Score = 171 bits (434), Expect = 3e-43
Identities = 77/100 (77%), Positives = 89/100 (89%)
Frame = +2
Query: 238 QLLPILKSNNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYH 297
Q L +SNN+DAYINGHDHCLEHIIDKESGI FFTSGGGSKAW GDI+PWDPKELKLYH
Sbjct: 209 QSLVAHQSNNIDAYINGHDHCLEHIIDKESGIPFFTSGGGSKAWRGDIRPWDPKELKLYH 388
Query: 298 DGQGFVSMQISKTNASIVFYDVFGKVLHTWSMSKKLKEAA 337
DGQGF+S+QI++ NA IVFYDVFGKVLH W+++K++ AA
Sbjct: 389 DGQGFMSVQITENNADIVFYDVFGKVLHRWNITKEMSAAA 508
Score = 136 bits (342), Expect = 1e-32
Identities = 81/173 (46%), Positives = 102/173 (58%), Gaps = 22/173 (12%)
Frame = +2
Query: 1 MAWCLNQHMLSSVIIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG 60
MAWCLNQ MLSSVI +SV LCLL+NHSSI EKLPRFEHHLKPQQQSLNFLVVGDWGRKG
Sbjct: 20 MAWCLNQRMLSSVIFLLSVSLCLLTNHSSIVEKLPRFEHHLKPQQQSLNFLVVGDWGRKG 199
Query: 61 NYNQSLVAHQMG---------------IVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYE 105
NYNQSLVAHQ I+ K I F S G + G DP +
Sbjct: 200 NYNQSLVAHQSNNIDAYINGHDHCLEHIIDKESGIPFFTSGGGSKAWRGDIRPWDPKELK 379
Query: 106 SFVDIYTAPSLQ-------QIWYNVLGNHDYRGDVEAQLSPILRQKDNRWVCL 151
+ D S+Q ++Y+V G +R ++ ++S + ++++C+
Sbjct: 380 LYHDGQGFMSVQITENNADIVFYDVFGKVLHRWNITKEMSAAA*IQ*SKYICI 538
>TC79946 similar to GP|21536860|gb|AAM61192.1 acid phosphatase type 5
{Arabidopsis thaliana}, partial (23%)
Length = 740
Score = 131 bits (330), Expect(2) = 7e-34
Identities = 60/110 (54%), Positives = 80/110 (72%)
Frame = +3
Query: 227 GPHGNTQELEEQLLPILKSNNVDAYINGHDHCLEHIIDKESGIHFFTSGGGSKAWGGDIK 286
G G+T+EL LLPIL++NNVD Y+NGHDHCLEHI +S + F TSGGGSKAW GD+
Sbjct: 63 GTTGDTKELLTHLLPILEANNVDMYMNGHDHCLEHISSTDSQMQFLTSGGGSKAWKGDVD 242
Query: 287 PWDPKELKLYHDGQGFVSMQISKTNASIVFYDVFGKVLHTWSMSKKLKEA 336
+K Y+DGQGF+S+++ + NA +V+YD+FG VLH ++SK L A
Sbjct: 243 KNRRDGVKFYYDGQGFMSVELKQMNAKVVYYDIFGNVLHVLNLSKALHYA 392
Score = 30.0 bits (66), Expect(2) = 7e-34
Identities = 13/27 (48%), Positives = 17/27 (62%)
Frame = +2
Query: 207 LVQSTAKWKIVVAHHPIKSAGPHGNTQ 233
LV + + KIVV HHP++S G H Q
Sbjct: 2 LVPNRHEGKIVVGHHPVRSIGHHXRHQ 82
>BF520178 similar to GP|7331197|gb|A putative purple acid phosphatase
precursor {Phaseolus vulgaris}, partial (21%)
Length = 612
Score = 103 bits (258), Expect = 7e-23
Identities = 63/155 (40%), Positives = 79/155 (50%)
Frame = +1
Query: 14 IIAVSVLLCLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGI 73
I V V++C+ N ++ +L R EH +L+FLV+GDWGRKG YNQS VA Q
Sbjct: 280 ICVVVVVVCVSVN---VSAELQRIEHPAVKADATLSFLVIGDWGRKGTYNQSQVAFQ--- 441
Query: 74 VGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQIWYNVLGNHDYRGDV 133
+ST + + NVLGNHDYRGDV
Sbjct: 442 ----------ISTPPTAFKN----------------------------NVLGNHDYRGDV 507
Query: 134 EAQLSPILRQKDNRWVCLRSFILDTDNVEFFFVDT 168
EAQLSP L+ D+RW C RSF + T+ EFFFVDT
Sbjct: 508 EAQLSPFLQNIDHRWFCQRSFFVHTEIAEFFFVDT 612
>TC88597 weakly similar to PIR|B96700|B96700 protein F12A21.17 [imported] -
Arabidopsis thaliana, partial (56%)
Length = 1459
Score = 31.2 bits (69), Expect = 0.59
Identities = 16/51 (31%), Positives = 25/51 (48%), Gaps = 2/51 (3%)
Frame = -3
Query: 106 SFVDIYTAPSLQQIWYNVLGNHDY--RGDVEAQLSPILRQKDNRWVCLRSF 154
SFV SLQQ WY+ HD+ R + ++P+ +K +C+ F
Sbjct: 1199 SFVLSNQEGSLQQYWYSTFRKHDW*CRNKFSSLVNPVSMKKQKNPICIYCF 1047
>TC86076 similar to GP|7208777|emb|CAB76911.1 putative PTS protein {Cicer
arietinum}, partial (51%)
Length = 677
Score = 30.8 bits (68), Expect = 0.77
Identities = 23/76 (30%), Positives = 36/76 (47%), Gaps = 2/76 (2%)
Frame = +2
Query: 63 NQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQI--W 120
N + H+M I+ + N+ +V TGDN Y D + +D AP+++ W
Sbjct: 302 NTTAFIHRM-ILAEKPNL--IVFTGDNIYGH------DSSDSAKSMDAAFAPAIESNIPW 454
Query: 121 YNVLGNHDYRGDVEAQ 136
VLGNHD G + +
Sbjct: 455 VAVLGNHDQEGSLSRE 502
>TC86077 similar to GP|7208777|emb|CAB76911.1 putative PTS protein {Cicer
arietinum}, partial (98%)
Length = 1877
Score = 29.6 bits (65), Expect = 1.7
Identities = 24/76 (31%), Positives = 37/76 (48%), Gaps = 2/76 (2%)
Frame = +2
Query: 63 NQSLVAHQMGIVGKNLNIEFVVSTGDNFYDDGLKGVDDPAFYESFVDIYTAPSLQQI--W 120
N + H+M I+ + N+ +V TGDN Y G D +S + AP+++ W
Sbjct: 575 NTTAFIHRM-ILAEKPNL--IVFTGDNIY-----GYDSSDSAKSMNAAF-APAIESNIPW 727
Query: 121 YNVLGNHDYRGDVEAQ 136
VLGNHD G + +
Sbjct: 728 VAVLGNHDQEGSLSRE 775
>TC86366 similar to PIR|T01857|T01857 hypothetical protein F9D12.1 -
Arabidopsis thaliana, partial (52%)
Length = 1722
Score = 28.1 bits (61), Expect = 5.0
Identities = 22/92 (23%), Positives = 43/92 (45%), Gaps = 9/92 (9%)
Frame = +1
Query: 30 IAEKLPRFEH-HLKPQQQSLNFLVVGDWGRKGNYNQ---SLVAHQMGIVGKNLNIEFVV- 84
+ ++LP H ++ +Q+ + F+ D + + + S V + G NL + V
Sbjct: 46 LKQELPHLSHLGVRARQEKVVFVTCEDDEKIADIQRLIGSCVRLEASAAGVNLTLASSVD 225
Query: 85 ----STGDNFYDDGLKGVDDPAFYESFVDIYT 112
S+ ++ +DD + GVD PAF + Y+
Sbjct: 226 LDGNSSVESAFDDNISGVDVPAFSAGRISKYS 321
>TC79909 similar to GP|8101442|gb|AAF72555.1| cryptochrome 1 {Lycopersicon
esculentum}, partial (27%)
Length = 1058
Score = 28.1 bits (61), Expect = 5.0
Identities = 11/29 (37%), Positives = 20/29 (68%)
Frame = +1
Query: 220 HHPIKSAGPHGNTQELEEQLLPILKSNNV 248
H PI++ PHG T+ ++Q++P + S+ V
Sbjct: 316 HEPIRNNAPHG-TRRYQDQMVPSMTSSRV 399
>BI264097 similar to GP|19570986|dbj diphosphonucleotide phosphatase -like
protein {Oryza sativa (japonica cultivar-group)},
partial (10%)
Length = 656
Score = 27.7 bits (60), Expect = 6.5
Identities = 15/49 (30%), Positives = 23/49 (46%), Gaps = 2/49 (4%)
Frame = +3
Query: 214 WKIVVAHHPIKSA--GPHGNTQELEEQLLPILKSNNVDAYINGHDHCLE 260
W I + H P+ ++ G Q+ + P+L N VD + GH H E
Sbjct: 81 WLIFMGHRPMYTSNNGFSSKDQKFINAVEPLLLQNKVDLVLFGHVHNYE 227
>TC80967 weakly similar to GP|21593303|gb|AAM65252.1 unknown {Arabidopsis
thaliana}, partial (24%)
Length = 1065
Score = 27.7 bits (60), Expect = 6.5
Identities = 31/141 (21%), Positives = 59/141 (40%), Gaps = 12/141 (8%)
Frame = +1
Query: 207 LVQSTAKWKIVVAHHPIKSAGPH-GNTQELE--EQLLPILKSNNVDAYINGHDHCLEHII 263
L +++ W+IVV + P+ G + NT++ + E L + VD Y++G D C +
Sbjct: 502 LQANSSNWRIVVGYQPLLICGQNKENTKKTKAFEYLHHLFTKFAVDVYLSGQD-CTIQVP 678
Query: 264 DKESGIHFFTSGGGSKAWGGDIKPWDPKELKLYHDGQ---------GFVSMQISKTNASI 314
D A+ G+ + K ++ +G+ GF+ Q+S T
Sbjct: 679 DNR------------VAYIGNPGLNEKKSYSVFLNGKSVFSRELANGFLLHQVSSTQIVT 822
Query: 315 VFYDVFGKVLHTWSMSKKLKE 335
+ + G+V + +K E
Sbjct: 823 YYVSLAGEVAFKTVLQEKSTE 885
>TC77183 homologue to SP|P43282|METM_LYCES S-adenosylmethionine synthetase 3
(EC 2.5.1.6) (Methionine adenosyltransferase 3),
complete
Length = 1660
Score = 27.7 bits (60), Expect = 6.5
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = -3
Query: 110 IYTAPSLQQIWYNVLGNHDYRGDVEAQL 137
+Y APS+ Q W+ G+ RGD+ Q+
Sbjct: 248 LYLAPSMHQGWHLKPGHRSCRGDLRLQI 165
>BQ141784 homologue to GP|23172168|gb| CG13615-PA {Drosophila melanogaster},
partial (7%)
Length = 1282
Score = 27.3 bits (59), Expect = 8.5
Identities = 11/24 (45%), Positives = 16/24 (65%)
Frame = +1
Query: 177 TDPGKHTYDWKGVLPLESYRAELL 200
T P +HT+D KG+ YR++LL
Sbjct: 736 TRPHRHTFDRKGINAQSRYRSQLL 807
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.137 0.427
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,192,915
Number of Sequences: 36976
Number of extensions: 184072
Number of successful extensions: 840
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 835
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 838
length of query: 337
length of database: 9,014,727
effective HSP length: 97
effective length of query: 240
effective length of database: 5,428,055
effective search space: 1302733200
effective search space used: 1302733200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146550.5


