

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145221.10 - phase: 0
(68 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
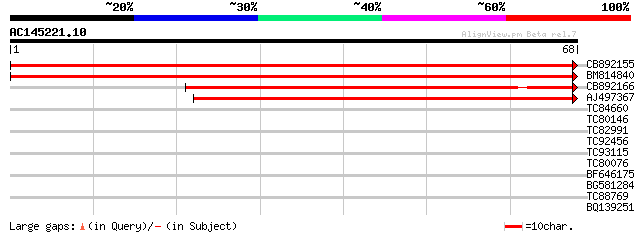
Score E
Sequences producing significant alignments: (bits) Value
CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292... 101 5e-23
BM814840 weakly similar to GP|15128241|db helicase-like protein ... 90 1e-19
CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis ... 62 6e-11
AJ497367 similar to GP|14140286|gb putative helicase {Oryza sati... 58 8e-10
TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG282... 37 0.001
TC80146 33 0.016
TC82991 similar to GP|20197706|gb|AAM15216.1 unknown protein {Ar... 27 1.5
TC92456 weakly similar to PIR|T46707|T46707 proteophosphoglycan ... 25 4.4
TC93115 similar to GP|13129508|gb|AAK13162.1 putative protein ph... 25 5.8
TC80076 similar to GP|10177459|dbj|BAB10850. gene_id:MQB2.13~unk... 25 5.8
BF646175 weakly similar to GP|10953887|gb| UDP-glucosyltransfera... 25 7.5
BG581284 similar to PIR|T48442|T484 hypothetical protein T32M21.... 25 7.5
TC88769 homologue to GP|19881561|gb|AAM00962.1 hypothetical prot... 24 9.9
BQ139251 similar to GP|396595|emb|C ACR1-protein {Saccharomyces ... 24 9.9
>CB892155 similar to PIR|D86481|D86 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (4%)
Length = 572
Score = 101 bits (252), Expect = 5e-23
Identities = 50/68 (73%), Positives = 58/68 (84%)
Frame = -2
Query: 1 MTINKSQGQSLKHDGVYLPTPVFSHGQLHVVVSKVTSREWLKILIIDEDGEDTDETSNIV 60
MTINKSQGQSLKH GVYLP+ VFSHGQL+V +S+VTSRE LKILI ++DGED TSN+V
Sbjct: 352 MTINKSQGQSLKHIGVYLPSSVFSHGQLYVALSRVTSREGLKILISNDDGEDDCVTSNVV 173
Query: 61 YEEVFQNV 68
Y EVF N+
Sbjct: 172 YREVFHNL 149
>BM814840 weakly similar to GP|15128241|db helicase-like protein {Oryza
sativa (japonica cultivar-group)}, partial (6%)
Length = 733
Score = 90.1 bits (222), Expect = 1e-19
Identities = 43/68 (63%), Positives = 53/68 (77%)
Frame = +3
Query: 1 MTINKSQGQSLKHDGVYLPTPVFSHGQLHVVVSKVTSREWLKILIIDEDGEDTDETSNIV 60
MTINKSQGQ+L G+YLP PVF+HGQL+V VS+V SR LKILI DE+G + T N+V
Sbjct: 204 MTINKSQGQTLTSVGLYLPRPVFTHGQLYVAVSRVKSRSGLKILITDENGSPSSSTVNVV 383
Query: 61 YEEVFQNV 68
Y+EVFQ +
Sbjct: 384 YQEVFQKI 407
>CB892166 similar to GP|20197614|gb unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 748
Score = 61.6 bits (148), Expect = 6e-11
Identities = 30/47 (63%), Positives = 40/47 (84%)
Frame = -1
Query: 22 VFSHGQLHVVVSKVTSREWLKILIIDEDGEDTDETSNIVYEEVFQNV 68
VFSHGQL+V +S+V+SR LKIL+IDE+G+ D T+N+VY +VFQNV
Sbjct: 274 VFSHGQLYVAISRVSSRNGLKILMIDENGDCIDNTTNVVY-KVFQNV 137
>AJ497367 similar to GP|14140286|gb putative helicase {Oryza sativa
(japonica cultivar-group)}, partial (1%)
Length = 543
Score = 57.8 bits (138), Expect = 8e-10
Identities = 27/46 (58%), Positives = 38/46 (81%)
Frame = +1
Query: 23 FSHGQLHVVVSKVTSREWLKILIIDEDGEDTDETSNIVYEEVFQNV 68
FS+G+L+V VS+VTSR+ LKIL+ EDG + TSN+VY+EVF+N+
Sbjct: 19 FSNGKLYVAVSRVTSRKGLKILLAHEDGNCMNTTSNVVYKEVFRNL 156
>TC84660 similar to PIR|D86481|D86481 hypothetical protein AAG28292.1
[imported] - Arabidopsis thaliana, partial (1%)
Length = 1009
Score = 37.0 bits (84), Expect = 0.001
Identities = 16/30 (53%), Positives = 23/30 (76%)
Frame = +2
Query: 10 SLKHDGVYLPTPVFSHGQLHVVVSKVTSRE 39
S+ G+YLP P+F HG L+V +S+VTSR+
Sbjct: 68 SMATKGMYLPQPIF*HG*LYVALSRVTSRK 157
>TC80146
Length = 476
Score = 33.5 bits (75), Expect = 0.016
Identities = 14/28 (50%), Positives = 21/28 (75%)
Frame = -3
Query: 2 TINKSQGQSLKHDGVYLPTPVFSHGQLH 29
TINKS+ QSL + +YL P+FSH +++
Sbjct: 450 TINKSR*QSLSYMKIYLSRPIFSHEEMY 367
>TC82991 similar to GP|20197706|gb|AAM15216.1 unknown protein {Arabidopsis
thaliana}, partial (5%)
Length = 769
Score = 26.9 bits (58), Expect = 1.5
Identities = 14/47 (29%), Positives = 26/47 (54%)
Frame = -1
Query: 1 MTINKSQGQSLKHDGVYLPTPVFSHGQLHVVVSKVTSREWLKILIID 47
+ ++ Q SLKH G+++PT ++ H HV+ + +K II+
Sbjct: 733 LDMSNLQFLSLKH-GLFIPTDIYRHKDFHVLTIYIPHLXIVKKKIIN 596
>TC92456 weakly similar to PIR|T46707|T46707 proteophosphoglycan
membrane-associated [imported] - Leishmania major
(fragment), partial (10%)
Length = 538
Score = 25.4 bits (54), Expect = 4.4
Identities = 13/36 (36%), Positives = 20/36 (55%), Gaps = 1/36 (2%)
Frame = -1
Query: 29 HVVVSKVTSREWLKILIIDEDGED-TDETSNIVYEE 63
H++ + V +E + ++DEDGED E N EE
Sbjct: 238 HLINNPVVPQEQEQEQVVDEDGEDAAQEDDNAAAEE 131
>TC93115 similar to GP|13129508|gb|AAK13162.1 putative protein phosphatase
2A regulatory subunit {Oryza sativa (japonica
cultivar-group)}, partial (17%)
Length = 739
Score = 25.0 bits (53), Expect = 5.8
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = +1
Query: 38 REWLKILIIDEDGEDTDETSNIVYEE 63
RE+++ L ++EDGED S V+EE
Sbjct: 214 REYIR-LSMEEDGEDMSNASGDVWEE 288
>TC80076 similar to GP|10177459|dbj|BAB10850. gene_id:MQB2.13~unknown
protein {Arabidopsis thaliana}, partial (21%)
Length = 910
Score = 25.0 bits (53), Expect = 5.8
Identities = 7/21 (33%), Positives = 15/21 (71%)
Frame = +1
Query: 38 REWLKILIIDEDGEDTDETSN 58
+ W++ +++DED E + +T N
Sbjct: 721 KSWIRKVVLDEDDEQSKKTDN 783
>BF646175 weakly similar to GP|10953887|gb| UDP-glucosyltransferase HRA25
{Phaseolus vulgaris}, partial (15%)
Length = 644
Score = 24.6 bits (52), Expect = 7.5
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = -2
Query: 20 TPVFSHGQLHVVVSKVTSREWL 41
TPVF L+ V KVT + WL
Sbjct: 202 TPVFDRSWLNSVCRKVTLQPWL 137
>BG581284 similar to PIR|T48442|T484 hypothetical protein T32M21.60 -
Arabidopsis thaliana, partial (6%)
Length = 420
Score = 24.6 bits (52), Expect = 7.5
Identities = 10/31 (32%), Positives = 19/31 (61%)
Frame = +1
Query: 36 TSREWLKILIIDEDGEDTDETSNIVYEEVFQ 66
TS EW + ++ ++DGE++ + EV+Q
Sbjct: 73 TSNEWPQNILGNDDGENSRLQEQVAASEVWQ 165
>TC88769 homologue to GP|19881561|gb|AAM00962.1 hypothetical protein {Oryza
sativa} [Oryza sativa (japonica cultivar-group)],
partial (7%)
Length = 912
Score = 24.3 bits (51), Expect = 9.9
Identities = 13/25 (52%), Positives = 15/25 (60%)
Frame = -3
Query: 10 SLKHDGVYLPTPVFSHGQLHVVVSK 34
SL DGV +PTP + G VVV K
Sbjct: 112 SLLGDGVDVPTPEGNSGGFMVVVEK 38
>BQ139251 similar to GP|396595|emb|C ACR1-protein {Saccharomyces cerevisiae},
partial (24%)
Length = 678
Score = 24.3 bits (51), Expect = 9.9
Identities = 12/43 (27%), Positives = 22/43 (50%)
Frame = +2
Query: 15 GVYLPTPVFSHGQLHVVVSKVTSREWLKILIIDEDGEDTDETS 57
G+Y G + + + TS EW K L+ D +G+ T +++
Sbjct: 509 GLYKGLGAVLTGIVPKMAIRFTSYEWYKQLLADSEGKVTSKST 637
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.314 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,035,391
Number of Sequences: 36976
Number of extensions: 22038
Number of successful extensions: 94
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 94
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 94
length of query: 68
length of database: 9,014,727
effective HSP length: 44
effective length of query: 24
effective length of database: 7,387,783
effective search space: 177306792
effective search space used: 177306792
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC145221.10


