

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
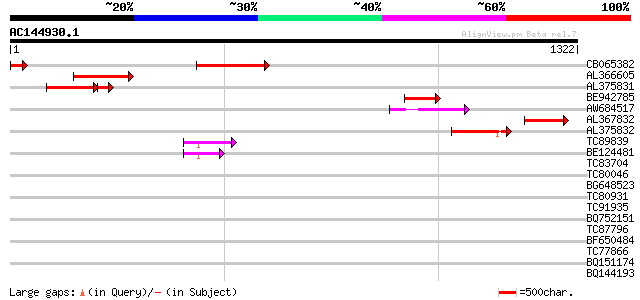
Query= AC144930.1 - phase: 0 /pseudo
(1322 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fra... 304 2e-82
AL366605 294 2e-79
AL375831 233 7e-78
BE942785 166 7e-41
AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thalia... 150 3e-36
AL367832 weakly similar to GP|14091845|gb Putative retroelement ... 150 3e-36
AL375832 142 7e-34
TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC ... 90 7e-18
BE124481 71 3e-12
TC83704 similar to PIR|T05005|T05005 hypothetical protein T19P19... 39 0.017
TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor ... 35 0.19
BG648523 similar to GP|9955540|emb| putative protein {Arabidopsi... 34 0.33
TC80931 similar to GP|14190783|gb|AAF65166.2 putative phloem tra... 34 0.33
TC91935 34 0.43
BQ752151 similar to GP|1945155|emb MN1 {Homo sapiens}, partial (1%) 33 0.56
TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear prot... 33 0.56
BF650484 similar to SP|P49730|RIR2 Ribonucleoside-diphosphate re... 33 0.73
TC77866 similar to GP|18266655|gb|AAL67601.1 putative cinnamoyl-... 33 0.95
BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xy... 33 0.95
BQ144193 weakly similar to GP|14026007|dbj putative transporter ... 33 0.95
>CB065382 similar to PIR|T09671|T096 RPE15 protein - alfalfa (fragment),
partial (9%)
Length = 624
Score = 304 bits (778), Expect = 2e-82
Identities = 146/169 (86%), Positives = 153/169 (90%)
Frame = -2
Query: 436 QQQQNQQARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGA 495
QQ+Q QQARPTFP IPMLYAE LPTLL RGHCT RQGKPPPDPLPPRFRSDLKCDFHQGA
Sbjct: 509 QQRQ*QQARPTFPLIPMLYAE*LPTLLLRGHCTIRQGKPPPDPLPPRFRSDLKCDFHQGA 330
Query: 496 LGHDVEGCYALKYIVKKLIDQGKLTFENNVPHVLDNPLPNHAAVNMIEVCEEAPRLDVRN 555
LGHDVEGCYALK+IVKKLI+QGKLTFENNVPHVLDNPLPNHAAVNMIEV EEAP LDVRN
Sbjct: 329 LGHDVEGCYALKHIVKKLINQGKLTFENNVPHVLDNPLPNHAAVNMIEVYEEAPGLDVRN 150
Query: 556 VATPLVPLHIKLCKASLFSHDHAKCLGCLHNPLGCYTVQDDIQSLMNDN 604
V TPLVPLHIKLC+ASLF HDHA C +NPLGC VQ+DIQSLMN+N
Sbjct: 149 VTTPLVPLHIKLCQASLFDHDHANCQE*FYNPLGCCVVQNDIQSLMNNN 3
Score = 63.5 bits (153), Expect = 5e-10
Identities = 32/40 (80%), Positives = 35/40 (87%)
Frame = -1
Query: 1 NAQLRTELATLREELAKANDVMTALLAAQEQSATAIPIAT 40
NAQLRTELA++REELAKAND MTALLAAQEQS+T P T
Sbjct: 618 NAQLRTELASMREELAKANDTMTALLAAQEQSSTGSPTTT 499
>AL366605
Length = 422
Score = 294 bits (752), Expect = 2e-79
Identities = 140/140 (100%), Positives = 140/140 (100%)
Frame = -3
Query: 148 HEIKANRGNADSFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 207
HEIKANRGNADSFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD
Sbjct: 420 HEIKANRGNADSFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 241
Query: 208 NDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLS 267
NDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLS
Sbjct: 240 NDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLS 61
Query: 268 QKKEESFREYAQRWRGAAAR 287
QKKEESFREYAQRWRGAAAR
Sbjct: 60 QKKEESFREYAQRWRGAAAR 1
>AL375831
Length = 467
Score = 233 bits (594), Expect(2) = 7e-78
Identities = 114/121 (94%), Positives = 117/121 (96%)
Frame = +2
Query: 87 VLIPATNAASINATLPQTTAAVTEPLVHTMPQSVNINTQHGSIPVTKTMEEMMEELAKEL 146
VLIPATNAASI ATLPQTTAAVTEPLVHT+PQ +NINTQH SIPVTKTMEEMMEELAKEL
Sbjct: 2 VLIPATNAASIIATLPQTTAAVTEPLVHTLPQGININTQHRSIPVTKTMEEMMEELAKEL 181
Query: 147 RHEIKANRGNADSFKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYK 206
RHEI+ANRGNADS KTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYK
Sbjct: 182 RHEIQANRGNADSVKTQDLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYK 361
Query: 207 D 207
D
Sbjct: 362 D 364
Score = 77.8 bits (190), Expect(2) = 7e-78
Identities = 35/36 (97%), Positives = 35/36 (97%)
Frame = +3
Query: 206 KDNDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDE 241
K NDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDE
Sbjct: 360 KTNDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDE 467
>BE942785
Length = 460
Score = 166 bits (419), Expect = 7e-41
Identities = 79/85 (92%), Positives = 79/85 (92%)
Frame = -1
Query: 920 ICKKWKACYHSWRGSIPG*PAIIFLLHRSRVCRGDRFSRINCRRYGAQEGWNCNGFFEGC 979
ICK W ACYHSWRGSI *PAIIFLLHRSRVCRGDRFSRINCRRYGAQEGWNCNGFFEGC
Sbjct: 385 ICK-WXACYHSWRGSILD*PAIIFLLHRSRVCRGDRFSRINCRRYGAQEGWNCNGFFEGC 209
Query: 980 TESRPRGSSCRLGQVNTAS*EQT*G 1004
TESRPR SSCRLGQVNTAS*EQ *G
Sbjct: 208 TESRPRESSCRLGQVNTAS*EQA*G 134
>AW684517 similar to GP|9927273|dbj Similar to Arabidopsis thaliana chromosome
II BAC F26H6; putative retroelement pol polyprotein,
partial (1%)
Length = 488
Score = 150 bits (379), Expect = 3e-36
Identities = 88/187 (47%), Positives = 105/187 (56%)
Frame = +3
Query: 885 HLSGHGH*CLLQLSLGETLDTRRGSSDLYFAPKVEICKKWKACYHSWRGSIPG*PAIIFL 944
H+SGHG+ C + LS G+TLDT R +LY P+VEIC++W
Sbjct: 3 HVSGHGYQCFV*LSPGKTLDT*RWRGNLYPTPEVEICEEW-------------------- 122
Query: 945 LHRSRVCRGDRFSRINCRRYGAQEGWNCNGFFEGCTESRPRGSSCRLGQVNTAS*EQT*G 1004
+C G+ FSR+ R AQEGW+CNGFFEG E G+ CRLGQVN *E G
Sbjct: 123 -----IC*GNCFSRVVYGRCRAQEGWSCNGFFEGRPEGCSGGTGCRLGQVNPTL*E*AKG 287
Query: 1005 RSWLLPNFRSFHRSIL*CWVCQRNHRRSNWIWPEACICYTRRHCERLGCH*HSLNHACF* 1064
RS +LPN H + *C VCQ R I A +C TRRHCERLGC * SLNHAC
Sbjct: 288 RSQILPNIGCLHGNFS*CRVCQYTC*RGGSICS*ALVCDTRRHCERLGCR*CSLNHACVX 467
Query: 1065 VIYLVA* 1071
+ LVA*
Sbjct: 468 XMCLVA* 488
>AL367832 weakly similar to GP|14091845|gb Putative retroelement {Oryza
sativa}, partial (2%)
Length = 384
Score = 150 bits (379), Expect = 3e-36
Identities = 83/103 (80%), Positives = 88/103 (84%)
Frame = -2
Query: 1200 KKRKPFSLTRKRLSLST*VPRRTSEKSRSVLL*RKGLKRRSSNSSGNIRIFLHGRMKICQ 1259
KK KPFSLTRKRLSLST*VPRRTSE+SRSVLL*RKGLK RSS+SS N IFLH RMKICQ
Sbjct: 383 KKGKPFSLTRKRLSLST*VPRRTSERSRSVLL*RKGLKGRSSSSSENTWIFLHARMKICQ 204
Query: 1260 VYIL*LWSIGSPPSLNVLPSGRN*EGLIPIWLSRLRSEVRTPL 1302
V IL LW+IGS SLNVLPSGRN*EGLI IWLSRLR ++ L
Sbjct: 203 V*ILKLWNIGSQQSLNVLPSGRN*EGLIQIWLSRLRVRFKSRL 75
>AL375832
Length = 535
Score = 142 bits (359), Expect = 7e-34
Identities = 89/149 (59%), Positives = 95/149 (63%), Gaps = 10/149 (6%)
Frame = +2
Query: 1031 RSNWIWPEACICYTRRHCERLGCH*HSLNHACF*VIYLVA*VSCFSK*QSSRSAQSESDI 1090
RSNWIW +ACICYTRRHCE LG H*HS NHACF V+ LVA CF K*QSSRSAQSESDI
Sbjct: 23 RSNWIWSQACICYTRRHCE*LGRH*HSFNHACFRVMRLVALALCFQK*QSSRSAQSESDI 202
Query: 1091 FCKGMFVSALPLINKNCRFFHD-CFF*YLFSWK*W*CQKKPKHLYFL---------INFK 1140
FCKG+FVSALP F FF* LF K +K ++ N K
Sbjct: 203 FCKGVFVSALPFDE*KLSFLSRLLFF*CLFLGKMVMPEKT*TSVFSY*FQKPA*KKKNTK 382
Query: 1141 NLHKKPPSKIIQPFYAD*TIINPLGTVIL 1169
N KK SKIIQ YA *TIINPL TVIL
Sbjct: 383 N--KKTLSKIIQSLYAG*TIINPLNTVIL 463
>TC89839 homologue to SP|P23466|CYAA_SACKL Adenylate cyclase (EC 4.6.1.1)
(ATP pyrophosphate-lyase) (Adenylyl cyclase). [Yeast],
partial (0%)
Length = 665
Score = 89.7 bits (221), Expect = 7e-18
Identities = 55/132 (41%), Positives = 67/132 (50%), Gaps = 7/132 (5%)
Frame = +3
Query: 405 PPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQ------QQNQ-QARPTFPPIPMLYAEL 457
PP Q P P QP P Q +Q +P Q QNQ Q PIP+ YA+L
Sbjct: 228 PPLVQQRPQSPYQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVKYADL 407
Query: 458 LPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHDVEGCYALKYIVKKLIDQG 517
LP LL + T P+ LPP +R DL C FHQGA GHD E CY LK V+KLI+
Sbjct: 408 LPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHDTEQCYPLKEEVQKLIENN 587
Query: 518 KLTFENNVPHVL 529
+F++ VL
Sbjct: 588 VWSFDDQDIKVL 623
>BE124481
Length = 538
Score = 70.9 bits (172), Expect = 3e-12
Identities = 43/102 (42%), Positives = 50/102 (48%), Gaps = 7/102 (6%)
Frame = +3
Query: 405 PPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQ------QQNQ-QARPTFPPIPMLYAEL 457
PP Q P P QP P Q +Q +P Q QNQ Q PIP+ YA+L
Sbjct: 231 PPLVQQRPQSPYQPMCPQKPRQQAPRQSIPQNQVPQKFIPQNQVQKASQCDPIPVKYADL 410
Query: 458 LPTLLHRGHCTTRQGKPPPDPLPPRFRSDLKCDFHQGALGHD 499
LP LL + T P+ LPP +R DL C FHQGA GHD
Sbjct: 411 LPILLKKNLIQTLPLPRVPNSLPPWYRPDLNCVFHQGAPGHD 536
>TC83704 similar to PIR|T05005|T05005 hypothetical protein T19P19.70 -
Arabidopsis thaliana, partial (15%)
Length = 716
Score = 38.5 bits (88), Expect = 0.017
Identities = 33/115 (28%), Positives = 47/115 (40%), Gaps = 11/115 (9%)
Frame = +1
Query: 379 ATVAPINAAQMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQ 438
+TV P+ +P PY QH PP P PP P++ + ++QLP
Sbjct: 262 STVPPVATTSLPEPLPY----QHKEQPPIPVTLPPPP-----PLSKLPPATREQLPPPPP 414
Query: 439 QNQ--QARPTFPPIPMLYAEL---------LPTLLHRGHCTTRQGKPPPDPLPPR 482
++ A PP P A+L LP L + R+ PPP PLP +
Sbjct: 415 LSKLPPAAREHPPPPPPSAKLPPAARERLPLPPPLSKLPPAARERLPPPSPLPEK 579
>TC80046 similar to SP|P22699|GUN6_DICDI Endoglucanase precursor (EC
3.2.1.4) (Endo-1 4-beta-glucanase) (Spore germination
protein 270-6), partial (11%)
Length = 643
Score = 35.0 bits (79), Expect = 0.19
Identities = 30/103 (29%), Positives = 37/103 (35%), Gaps = 1/103 (0%)
Frame = +1
Query: 383 PINAAQMPPSYP-YVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQ 441
P Q PP P P + P PP LP PQ Q++ Q+LP + Q
Sbjct: 286 PQRPPQRPPQKPPQRPPQKPPQRPPQRPPQKLPQRPPQKLPQRPPQKLPQKLPQKPPQRP 465
Query: 442 QARPTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRFR 484
P PP+ L P L H +P P P P R
Sbjct: 466 LPHPQKPPLRPLPHPQKPPLRPLPHPQKPPLRPLPHPQKPPLR 594
>BG648523 similar to GP|9955540|emb| putative protein {Arabidopsis thaliana},
partial (9%)
Length = 770
Score = 34.3 bits (77), Expect = 0.33
Identities = 35/112 (31%), Positives = 44/112 (39%), Gaps = 3/112 (2%)
Frame = +3
Query: 373 GAGQSMATVAP-INAAQMPPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQ 431
G G +M P + +Q P+ P PP H L P Q Q QQ Q
Sbjct: 318 GVGVAMPLSIPSFDGSQGEQKQPHPGSIGAPPLPPGPHPSLLNPNQQQP-----FQQNPQ 482
Query: 432 QLPVQQQQNQQARPTFPPIPMLYAELLPTLLHRGHCT--TRQGKPPPDPLPP 481
Q+P QQQ QQ P+PM +P + H H + Q P P P P
Sbjct: 483 QIPQHQQQLQQ---HMGPLPM--PPNMPQIQHSSHSSMLPHQHLPRPPPQMP 623
>TC80931 similar to GP|14190783|gb|AAF65166.2 putative phloem transcription
factor M1 {Apium graveolens}, partial (67%)
Length = 1193
Score = 34.3 bits (77), Expect = 0.33
Identities = 26/84 (30%), Positives = 38/84 (44%)
Frame = +1
Query: 26 LAAQEQSATAIPIATTIPMTTSMLPTASTDARFAMPAGFPYGLPPFFTPSTAAGTSGTAN 85
+ +Q+Q + I TT+ TA+T++ A FP TP+TAA T T+
Sbjct: 535 MQSQQQHLSTIAATDATTTTTATASTAATES-----AAFPDSTADSTTPATAAATPSTST 699
Query: 86 NVLIPATNAASINATLPQTTAAVT 109
A +AA+ AT TA T
Sbjct: 700 ---AAAFSAAATAATASAVTATAT 762
>TC91935
Length = 1144
Score = 33.9 bits (76), Expect = 0.43
Identities = 26/99 (26%), Positives = 45/99 (45%), Gaps = 8/99 (8%)
Frame = +2
Query: 168 VSKVDVPKKFKIPDFDRYNGLTCP--------QNHIIKYVRKMGNYKDNDSLMIHCFQDS 219
VS+V + ++ +P+F R LTCP N++I Y+ ++ N ++ C QD
Sbjct: 521 VSRVIIQSRYNVPNFSRSTALTCPTFDQEASLPNNLIAYLSRLANM----MIIFSCKQDH 688
Query: 220 LMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKP 258
+ Y L I T ++ FK + +TR +P
Sbjct: 689 I-------YVRLKNKGIITLAQV-EGFKRN---DTRFQP 772
>BQ752151 similar to GP|1945155|emb MN1 {Homo sapiens}, partial (1%)
Length = 650
Score = 33.5 bits (75), Expect = 0.56
Identities = 30/95 (31%), Positives = 38/95 (39%), Gaps = 1/95 (1%)
Frame = +3
Query: 390 PPSYPYVPYSQHPFFPPFYHQYPLPPGQPQ-VPVNAIAQQMKQQLPVQQQQNQQARPTFP 448
PPS+P P PP +P P Q +P QQ QQQ QQA P P
Sbjct: 327 PPSHPPPPKQPQQQAPPPQQAWPSPHVSAQKIPPGGHLQQQTWASAPPQQQPQQAPP--P 500
Query: 449 PIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPRF 483
P M + RG + G+P P +P R+
Sbjct: 501 PHHMSMRD------DRG--ASMYGQPHPHAMPGRY 581
>TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear protein.
[strain B95-8 Human herpesvirus 4] {Epstein-barr
virus}, partial (23%)
Length = 431
Score = 33.5 bits (75), Expect = 0.56
Identities = 27/92 (29%), Positives = 32/92 (34%)
Frame = -1
Query: 390 PPSYPYVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQARPTFPP 449
PPS P +P P+ PP P PP +P P P PP
Sbjct: 233 PPSPPPIP---PPYAPPCAPPPPPPPPEPPPP---------------PPPPGDPPPYPPP 108
Query: 450 IPMLYAELLPTLLHRGHCTTRQGKPPPDPLPP 481
+P P L + C PPP PLPP
Sbjct: 107 VP-------PPLPYPTPCAPPAPPPPPPPLPP 33
>BF650484 similar to SP|P49730|RIR2 Ribonucleoside-diphosphate reductase small
chain (EC 1.17.4.1) (Ribonucleotide reductase), partial
(60%)
Length = 660
Score = 33.1 bits (74), Expect = 0.73
Identities = 13/30 (43%), Positives = 15/30 (49%)
Frame = -3
Query: 1023 WVCQRNHRRSNWIWPEACICYTRRHCERLG 1052
WV H S W+W E + RRHC LG
Sbjct: 133 WVLDWEHTESVWVWSEKRFFWERRHCLVLG 44
>TC77866 similar to GP|18266655|gb|AAL67601.1 putative cinnamoyl-CoA
reductase {Oryza sativa}, partial (58%)
Length = 1300
Score = 32.7 bits (73), Expect = 0.95
Identities = 14/58 (24%), Positives = 31/58 (53%)
Frame = -2
Query: 558 TPLVPLHIKLCKASLFSHDHAKCLGCLHNPLGCYTVQDDIQSLMNDNFLTVSDVCVIV 615
TP+ + I+ ++F +H+ C GC HN + C+ + + ++ + VS + +I+
Sbjct: 657 TPMYFIQIRFYGFTIFPPNHSSC*GCEHNFVYCFCLSTSFKHVVCCSNFHVSHIFIII 484
>BQ151174 similar to GP|11036868|gb| PxORF73 peptide {Plutella xylostella
granulovirus}, partial (42%)
Length = 909
Score = 32.7 bits (73), Expect = 0.95
Identities = 27/98 (27%), Positives = 36/98 (36%), Gaps = 3/98 (3%)
Frame = +1
Query: 388 QMPPSYP---YVPYSQHPFFPPFYHQYPLPPGQPQVPVNAIAQQMKQQLPVQQQQNQQAR 444
Q PP+ P P + P PP P PP P P + + + P +
Sbjct: 169 QAPPTKPPTRQTPKNNTPPPPPHTPPDPTPPQPPPQPPKPPHHEKRPRTP-------EPP 327
Query: 445 PTFPPIPMLYAELLPTLLHRGHCTTRQGKPPPDPLPPR 482
PP P + PT RG R +P P P PP+
Sbjct: 328 GGRPPPPTTPKQARPTTQERG--GARGARPRPPPAPPQ 435
>BQ144193 weakly similar to GP|14026007|dbj putative transporter
{Mesorhizobium loti}, partial (4%)
Length = 1037
Score = 32.7 bits (73), Expect = 0.95
Identities = 21/67 (31%), Positives = 28/67 (41%), Gaps = 1/67 (1%)
Frame = +2
Query: 352 SKRYGNGHHKK-KETEIGMVSAGAGQSMATVAPINAAQMPPSYPYVPYSQHPFFPPFYHQ 410
+K+ NG K ET G + +S + +MPP YP HP PP H
Sbjct: 47 TKKRANGKQAKINETNAGKLKV---KSNTPAYLVGYKEMPPVYPRKSPKTHPRPPPNSHN 217
Query: 411 YPLPPGQ 417
P PP +
Sbjct: 218 RPEPPAK 238
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.332 0.143 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 45,106,210
Number of Sequences: 36976
Number of extensions: 773812
Number of successful extensions: 5790
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 4055
Number of HSP's successfully gapped in prelim test: 205
Number of HSP's that attempted gapping in prelim test: 1233
Number of HSP's gapped (non-prelim): 4714
length of query: 1322
length of database: 9,014,727
effective HSP length: 108
effective length of query: 1214
effective length of database: 5,021,319
effective search space: 6095881266
effective search space used: 6095881266
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144930.1


