

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
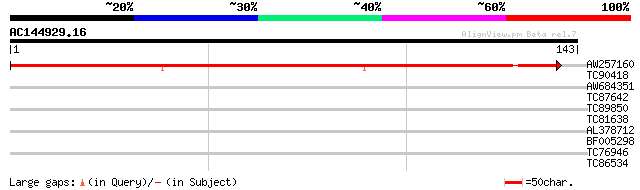
Query= AC144929.16 + phase: 0
(143 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW257160 similar to GP|9757790|dbj| gene_id:MUA22.9~unknown prot... 157 2e-39
TC90418 similar to GP|11762174|gb|AAG40365.1 At1g17620 {Arabidop... 32 0.087
AW684351 weakly similar to GP|6016719|gb|A putative cell divisio... 32 0.11
TC87642 similar to PIR|S51590|S51590 mitochondrial processing pe... 27 3.6
TC89850 similar to GP|22748338|gb|AAN05340.1 Unknown protein {Or... 26 4.7
TC81638 homologue to GP|10177686|dbj|BAB11012. emb|CAB62594.1~ge... 26 4.7
AL378712 26 6.2
BF005298 26 6.2
TC76946 homologue to PIR|T06570|T06570 superoxide dismutase (EC ... 25 8.1
TC86534 homologue to SP|P12227|RPOD_PEA DNA-directed RNA polymer... 25 8.1
>AW257160 similar to GP|9757790|dbj| gene_id:MUA22.9~unknown protein
{Arabidopsis thaliana}, partial (12%)
Length = 764
Score = 157 bits (396), Expect = 2e-39
Identities = 93/160 (58%), Positives = 109/160 (68%), Gaps = 21/160 (13%)
Frame = +3
Query: 1 MKDAAVTETQLKLISYELEKFVEADEECFYESSGRIS--FRSNVTLSRKKTDGLETEDLG 58
MKDA VTE +LKLISYELEKF+E +EE F ESSGR S SN+ +S K+TDG E ED G
Sbjct: 54 MKDADVTENELKLISYELEKFLETEEESFNESSGRNSRLSLSNIKISGKQTDGSEDEDCG 233
Query: 59 NKVICPLQEYLLGSSFEIREKTEVRIERAS-------------------VRETQVKQGRR 99
NK +CPLQ YLLGSSFEI EK +VR ERAS V++TQVKQ +
Sbjct: 234 NKDVCPLQGYLLGSSFEIPEKVQVRKERASLAELFQRTKTANEDCIETGVKDTQVKQAHK 413
Query: 100 SALHIIKKMSNMVLSSSKSCNTYGNTADHATSTNEKLCKV 139
S +HI+KKM MV SSKSCNT N AD +T+TN+KL KV
Sbjct: 414 SPMHIMKKMLKMVHISSKSCNTSENIAD-STTTNKKLSKV 530
>TC90418 similar to GP|11762174|gb|AAG40365.1 At1g17620 {Arabidopsis
thaliana}, partial (9%)
Length = 835
Score = 32.0 bits (71), Expect = 0.087
Identities = 18/41 (43%), Positives = 27/41 (64%), Gaps = 2/41 (4%)
Frame = -1
Query: 60 KVICPLQEY--LLGSSFEIREKTEVRIERASVRETQVKQGR 98
KVIC L + +L +FE+R + EVR+ S+R+ VK+GR
Sbjct: 412 KVICLLGFFAVMLTLNFELRVEEEVRLR*ESLRDVTVKEGR 290
>AW684351 weakly similar to GP|6016719|gb|A putative cell division control
protein {Arabidopsis thaliana}, partial (3%)
Length = 554
Score = 31.6 bits (70), Expect = 0.11
Identities = 21/63 (33%), Positives = 35/63 (55%)
Frame = +3
Query: 72 SSFEIREKTEVRIERASVRETQVKQGRRSALHIIKKMSNMVLSSSKSCNTYGNTADHATS 131
S E+R + R + A + E++ K R A HI K+M+ V SSS+ ++ + + A S
Sbjct: 219 SDDEVRNSSRKRRKDAMIDESEEKLQRLEARHIEKRMNTQVSSSSEEDSS--DDDEGAVS 392
Query: 132 TNE 134
T+E
Sbjct: 393 TSE 401
>TC87642 similar to PIR|S51590|S51590 mitochondrial processing peptidase (EC
3.4.24.64) alpha-II chain precursor - potato, partial
(75%)
Length = 1441
Score = 26.6 bits (57), Expect = 3.6
Identities = 12/35 (34%), Positives = 17/35 (48%)
Frame = -1
Query: 29 FYESSGRISFRSNVTLSRKKTDGLETEDLGNKVIC 63
F S +FRS+ S+ K G T+ N +IC
Sbjct: 247 FSSPSSAFTFRSSSLTSQSKKAGFLTQSTSNSIIC 143
>TC89850 similar to GP|22748338|gb|AAN05340.1 Unknown protein {Oryza sativa
(japonica cultivar-group)}, partial (34%)
Length = 1093
Score = 26.2 bits (56), Expect = 4.7
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = -3
Query: 95 KQGRRSALHIIKKMSNMVLSSSKSCNTYG 123
K GR+ ++ I +SN V+ +K+ TYG
Sbjct: 1034 KSGRKPSISISGTLSNTVIHPTKNWRTYG 948
>TC81638 homologue to GP|10177686|dbj|BAB11012.
emb|CAB62594.1~gene_id:MUL8.6~similar to unknown protein
{Arabidopsis thaliana}, partial (6%)
Length = 1585
Score = 26.2 bits (56), Expect = 4.7
Identities = 20/88 (22%), Positives = 38/88 (42%)
Frame = +3
Query: 32 SSGRISFRSNVTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRIERASVRE 91
SS R S + +L+ +K L K++ L+ E+ E +E ++
Sbjct: 1320 SSPRASLKRVSSLNSRKNKSL-------KIVSHLRNQNTARKVELDEHNNNEVEEKTLYV 1478
Query: 92 TQVKQGRRSALHIIKKMSNMVLSSSKSC 119
+ + LH+IK + M++ +S SC
Sbjct: 1479 VNM-ESENILLHLIKPQAMMMIHTSLSC 1559
>AL378712
Length = 482
Score = 25.8 bits (55), Expect = 6.2
Identities = 10/43 (23%), Positives = 22/43 (50%)
Frame = -1
Query: 42 VTLSRKKTDGLETEDLGNKVICPLQEYLLGSSFEIREKTEVRI 84
+ L RK+ D + + + CP+ EY+ + + KT++ +
Sbjct: 251 IFLKRKRNDSI*NHNYISS*FCPMVEYMKRAKIYVAHKTQLPV 123
>BF005298
Length = 512
Score = 25.8 bits (55), Expect = 6.2
Identities = 11/27 (40%), Positives = 20/27 (73%)
Frame = -2
Query: 73 SFEIREKTEVRIERASVRETQVKQGRR 99
+FE+RE +RI S +E+++++GRR
Sbjct: 406 NFEVRE---IRIGARSTKESEIRRGRR 335
>TC76946 homologue to PIR|T06570|T06570 superoxide dismutase (EC 1.15.1.1)
(Cu-Zn) - garden pea, complete
Length = 837
Score = 25.4 bits (54), Expect = 8.1
Identities = 14/37 (37%), Positives = 18/37 (47%)
Frame = -3
Query: 103 HIIKKMSNMVLSSSKSCNTYGNTADHATSTNEKLCKV 139
HII SN + S S T T+ ATS + L K+
Sbjct: 595 HIITHYSNFMHPRSSSLQTNNTTSYSATSISSSLAKL 485
>TC86534 homologue to SP|P12227|RPOD_PEA DNA-directed RNA polymerase beta'
chain (EC 2.7.7.6) (Fragment). [Garden pea] {Pisum
sativum}, partial (61%)
Length = 2441
Score = 25.4 bits (54), Expect = 8.1
Identities = 29/129 (22%), Positives = 51/129 (39%), Gaps = 39/129 (30%)
Frame = +1
Query: 32 SSGRISFRSNVTLSRKKTDG-------------LETEDLGNKVICPLQEYLL-------- 70
S+G+I F ++ + G +E++D+ + VI P + +LL
Sbjct: 730 SNGKIKFNEDLVHPTRTRHGYPAFICNIDLYVTVESDDIIHNVIIPPKSFLLVQNDQYVK 909
Query: 71 -----------GSSFEIREKTEVRIERASVRE----TQVKQGRR---SALHIIKKMSNMV 112
S+F ++E+ I S E T V S +HI+ K S++
Sbjct: 910 SEQVIAEIRAGTSTFNLKERVRKHIYSDSEGEMHWSTDVYHASEFLYSNVHILPKTSHLW 1089
Query: 113 LSSSKSCNT 121
+ S KSC +
Sbjct: 1090ILSGKSCRS 1116
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.312 0.128 0.345
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,660,675
Number of Sequences: 36976
Number of extensions: 36299
Number of successful extensions: 160
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 158
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 158
length of query: 143
length of database: 9,014,727
effective HSP length: 87
effective length of query: 56
effective length of database: 5,797,815
effective search space: 324677640
effective search space used: 324677640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC144929.16


