

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144928.9 - phase: 2 /pseudo
(1036 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
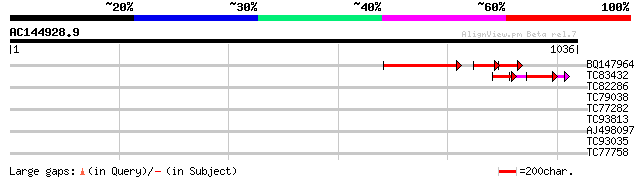
Score E
Sequences producing significant alignments: (bits) Value
BQ147964 similar to GP|7769862|gb F12M16.28 {Arabidopsis thalian... 291 e-112
TC83432 weakly similar to GP|7769862|gb|AAF69540.1| F12M16.28 {A... 108 1e-36
TC82286 weakly similar to PIR|B96544|B96544 hypothetical protein... 35 0.11
TC79038 similar to GP|15418778|gb|AAK37761.1 zinc transporter pr... 31 2.8
TC77282 homologue to GP|17065300|gb|AAL32804.1 Unknown protein {... 30 4.8
TC93813 similar to GP|11935197|gb|AAG42014.1 unknown protein {Ar... 30 6.3
AJ498097 weakly similar to PIR|T14441|T144 glycine-rich protein ... 30 6.3
TC93035 30 6.3
TC77758 similar to GP|1255951|emb|CAA65634.1 PS60 {Nicotiana tab... 29 8.2
>BQ147964 similar to GP|7769862|gb F12M16.28 {Arabidopsis thaliana}, partial
(15%)
Length = 698
Score = 291 bits (745), Expect(3) = e-112
Identities = 143/143 (100%), Positives = 143/143 (100%)
Frame = +3
Query: 683 LSRPSS*MDASQ*ISNTS*YETACSTVWYTAECKFS**LGGSGKDVRRRIME*RKKQCGT 742
LSRPSS*MDASQ*ISNTS*YETACSTVWYTAECKFS**LGGSGKDVRRRIME*RKKQCGT
Sbjct: 3 LSRPSS*MDASQ*ISNTS*YETACSTVWYTAECKFS**LGGSGKDVRRRIME*RKKQCGT 182
Query: 743 ARRKDKT*FFKIQGFIQPENSWCIQAI*VLSYTGWEAAITRSQDPGRRLSYFATCWSMLR 802
ARRKDKT*FFKIQGFIQPENSWCIQAI*VLSYTGWEAAITRSQDPGRRLSYFATCWSMLR
Sbjct: 183 ARRKDKT*FFKIQGFIQPENSWCIQAI*VLSYTGWEAAITRSQDPGRRLSYFATCWSMLR 362
Query: 803 INYQIE*SNIWCIWLHLYGDCCI 825
INYQIE*SNIWCIWLHLYGDCCI
Sbjct: 363 INYQIE*SNIWCIWLHLYGDCCI 431
Score = 90.5 bits (223), Expect(3) = e-112
Identities = 46/48 (95%), Positives = 47/48 (97%)
Frame = +2
Query: 848 LFYAK*RP*DHFLWINCTTGGRVILA*AA*LIFSRKTRWIILIQ*SNQ 895
LFYAK*RP*DHFLWINCTTGGRVILA*AA*LIFSRKTRWIILIQ*S +
Sbjct: 431 LFYAK*RP*DHFLWINCTTGGRVILA*AA*LIFSRKTRWIILIQ*STR 574
Score = 65.1 bits (157), Expect(3) = e-112
Identities = 35/43 (81%), Positives = 35/43 (81%)
Frame = +1
Query: 894 NQWCIYLCSIS*PIQDLLSQITILCCFVLYTV*LA*HMHYP*F 936
NQ IYLCSIS*PIQDLLSQI CFVL T *LA*HMHYP*F
Sbjct: 568 NQGGIYLCSIS*PIQDLLSQILYXACFVLDTG*LA*HMHYP*F 696
>TC83432 weakly similar to GP|7769862|gb|AAF69540.1| F12M16.28 {Arabidopsis
thaliana}, partial (12%)
Length = 433
Score = 108 bits (271), Expect(2) = 1e-36
Identities = 54/57 (94%), Positives = 54/57 (94%)
Frame = +3
Query: 945 TSSSCFDSHCNTTKR**NFKGYS*FMLL*VGFTSIGHCKC*KVPRSVAHNSLRFTFK 1001
TSSSCFDSHCNTTKR**NFKGYS*FMLL*VGFTSIGHCKC*KVPRSVAHNSLR K
Sbjct: 201 TSSSCFDSHCNTTKR**NFKGYS*FMLL*VGFTSIGHCKC*KVPRSVAHNSLRSLLK 371
Score = 93.2 bits (230), Expect = 5e-19
Identities = 59/111 (53%), Positives = 63/111 (56%), Gaps = 2/111 (1%)
Frame = +2
Query: 914 ITILCCFVLYTV*LA*HMHYP*FLNLAQRSCTS--SSCFDSHCNTTKR**NFKGYS*FML 971
ITILCCF LYTV*LA*HMHYP*FLNLAQRSC F K+ F
Sbjct: 101 ITILCCFGLYTV*LA*HMHYP*FLNLAQRSCGQYFFQLFRLSLQHNKKIVKF*RL*LIYA 280
Query: 972 L*VGFTSIGHCKC*KVPRSVAHNSLRFTFKKWL*SS*LESLHIHPHSYGCY 1022
G + K + + TFKKWL*SS*LESLHIHPHSYGCY
Sbjct: 281 TLSGLYKHWSLQMLKGTKECGS*LVAVTFKKWL*SS*LESLHIHPHSYGCY 433
Score = 63.9 bits (154), Expect(2) = 1e-36
Identities = 33/44 (75%), Positives = 37/44 (84%), Gaps = 1/44 (2%)
Frame = +1
Query: 883 KTRWIILIQ*SNQWCIYLCSIS*PIQDLLSQIT-ILCCFVLYTV 925
K RWIILIQ*SNQWCIYLCSIS*PIQDLLSQ I+ +++Y V
Sbjct: 7 KKRWIILIQ*SNQWCIYLCSIS*PIQDLLSQHNYIVLLWLVYCV 138
>TC82286 weakly similar to PIR|B96544|B96544 hypothetical protein F4M15.4
[imported] - Arabidopsis thaliana, partial (5%)
Length = 1782
Score = 35.4 bits (80), Expect = 0.11
Identities = 18/46 (39%), Positives = 26/46 (56%), Gaps = 7/46 (15%)
Frame = -1
Query: 145 PMQPNHTCGGANVWADFSSSSETFCSA-------GSYCPTTTTKFP 183
P+ P+ C ++ ++ FSSSS +F S+ GSY T TT FP
Sbjct: 492 PLSPSFMCSASSSFSSFSSSSSSFSSSPQSVINSGSYSSTITTFFP 355
>TC79038 similar to GP|15418778|gb|AAK37761.1 zinc transporter protein ZIP1
{Glycine max}, partial (87%)
Length = 1546
Score = 30.8 bits (68), Expect = 2.8
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +2
Query: 91 EMPARTSNCRACCEGFFCPHGITCMI 116
E A NCR CC+G CP+ +C +
Sbjct: 53 EGAAGRPNCRFCCDGHTCPNSNSCRV 130
>TC77282 homologue to GP|17065300|gb|AAL32804.1 Unknown protein {Arabidopsis
thaliana}, partial (19%)
Length = 787
Score = 30.0 bits (66), Expect = 4.8
Identities = 18/50 (36%), Positives = 26/50 (52%), Gaps = 11/50 (22%)
Frame = -1
Query: 37 EVKFYLNSLM--------EKSSSANYLKPNRNCN---LTSWVSGCEPGWA 75
+ KF LN L KSS++N +R C+ ++SW+S C P WA
Sbjct: 520 KTKFCLNHLS*NHGDENSSKSSNSNKGVGDRECSRGCISSWLSRCGPAWA 371
>TC93813 similar to GP|11935197|gb|AAG42014.1 unknown protein {Arabidopsis
thaliana}, partial (70%)
Length = 664
Score = 29.6 bits (65), Expect = 6.3
Identities = 16/53 (30%), Positives = 24/53 (45%), Gaps = 13/53 (24%)
Frame = +1
Query: 64 TSWVSGCEPG-------------WACSVPSGQKIDLKDSKEMPARTSNCRACC 103
TS++SG +PG W C + +GQ + K+ + S C ACC
Sbjct: 445 TSFISGIDPGTTCLICEGLFFTWWMCGIHTGQ-VRQSLQKKYHLKNSPCNACC 600
>AJ498097 weakly similar to PIR|T14441|T144 glycine-rich protein - wild
cabbage (fragment), partial (25%)
Length = 592
Score = 29.6 bits (65), Expect = 6.3
Identities = 15/54 (27%), Positives = 24/54 (43%), Gaps = 11/54 (20%)
Frame = +1
Query: 431 YIEKCNRENQTWPYH---RYHGPVRGWQDNISFSS--------SWKSTWMLGYW 473
Y +K R +TW + +++G R W + +SS W S W G+W
Sbjct: 250 YYKKGRRRRRTWRFRWPWKWNGRKRNW*RKVRWSSCRNYRRCSGWWSCWWSGWW 411
>TC93035
Length = 766
Score = 29.6 bits (65), Expect = 6.3
Identities = 15/47 (31%), Positives = 20/47 (41%), Gaps = 7/47 (14%)
Frame = -1
Query: 63 LTSWVSGCEPGWACSVPSGQKI-------DLKDSKEMPARTSNCRAC 102
+ SW+S GWA S P +K +L S P +CR C
Sbjct: 616 INSWLSSSSTGWAKSTPRNRKSLTQPPANNLSRSTRAPLGRPSCRGC 476
>TC77758 similar to GP|1255951|emb|CAA65634.1 PS60 {Nicotiana tabacum},
partial (96%)
Length = 2197
Score = 29.3 bits (64), Expect = 8.2
Identities = 9/23 (39%), Positives = 15/23 (65%)
Frame = +3
Query: 199 SNPKHACIWSYAHCCFEYFATHY 221
++ + +C WS A CF +F +HY
Sbjct: 171 NDSQFSCCWSLASMCFYFFVSHY 239
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.354 0.153 0.599
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,703,462
Number of Sequences: 36976
Number of extensions: 829229
Number of successful extensions: 11099
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 3378
Number of HSP's successfully gapped in prelim test: 482
Number of HSP's that attempted gapping in prelim test: 7346
Number of HSP's gapped (non-prelim): 4589
length of query: 1036
length of database: 9,014,727
effective HSP length: 106
effective length of query: 930
effective length of database: 5,095,271
effective search space: 4738602030
effective search space used: 4738602030
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144928.9


