

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144928.8 + phase: 0 /pseudo
(359 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
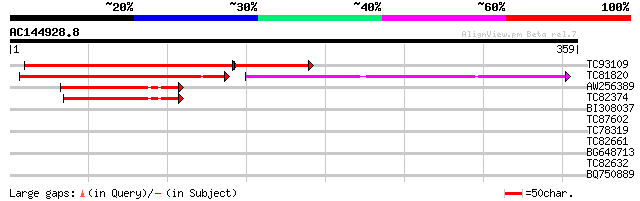
Score E
Sequences producing significant alignments: (bits) Value
TC93109 similar to GP|5824321|emb|CAB54139.1 ATPase {Solanum tub... 263 6e-96
TC81820 similar to GP|6056208|gb|AAF02825.1| putative ATPase {Ar... 174 3e-63
AW256389 similar to GP|21689765|gb putative arsA-like protein hA... 59 4e-09
TC82374 similar to GP|21689765|gb|AAM67526.1 putative arsA-like ... 50 1e-06
BI308037 weakly similar to GP|12583663|dbj chloroplast RelA homo... 33 0.22
TC87602 similar to GP|21741453|emb|CAD41397. OJ000223_09.9 {Oryz... 31 0.65
TC78319 homologue to GP|21592386|gb|AAM64337.1 mrp protein puta... 30 1.9
TC82661 similar to PIR|H86386|H86386 hypothetical protein AAG381... 28 4.2
BG648713 similar to GP|16924108|gb putative nucleotide-binding p... 28 4.2
TC82632 homologue to GP|18086370|gb|AAL57645.1 At2g47390/T8I13.2... 28 5.5
BQ750889 28 5.5
>TC93109 similar to GP|5824321|emb|CAB54139.1 ATPase {Solanum tuberosum},
partial (44%)
Length = 743
Score = 263 bits (671), Expect(2) = 6e-96
Identities = 130/135 (96%), Positives = 133/135 (98%)
Frame = +3
Query: 10 VKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTD 69
VKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTD
Sbjct: 204 VKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTD 383
Query: 70 PAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMD 129
PAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMD
Sbjct: 384 PAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKDFMD 563
Query: 130 GMGLGMIVDQTAKVR 144
GMGLGMIVDQ +++
Sbjct: 564 GMGLGMIVDQLGELK 608
Score = 106 bits (264), Expect(2) = 6e-96
Identities = 50/50 (100%), Positives = 50/50 (100%)
Frame = +2
Query: 143 VRRAKTGRVTGYTSSWTG*SYCDFQGYTIS*ITGI*YVHSNCI*YGTYWS 192
VRRAKTGRVTGYTSSWTG*SYCDFQGYTIS*ITGI*YVHSNCI*YGTYWS
Sbjct: 593 VRRAKTGRVTGYTSSWTG*SYCDFQGYTIS*ITGI*YVHSNCI*YGTYWS 742
>TC81820 similar to GP|6056208|gb|AAF02825.1| putative ATPase {Arabidopsis
thaliana}, partial (69%)
Length = 1320
Score = 174 bits (440), Expect(2) = 3e-63
Identities = 85/133 (63%), Positives = 103/133 (76%)
Frame = +2
Query: 7 LLSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVV 66
L + + ES+ FD+MV ER+YYMLGGKGGVGKTSCAASLA+KFAN+GHPT+VV
Sbjct: 29 LSQARRIGNVAESVCGFDEMVRDKERRYYMLGGKGGVGKTSCAASLAIKFANHGHPTMVV 208
Query: 67 STDPAHSLSDSFAQDLTGGALVQVDGPDYPLFALEINPEKAREDFRDVAKQNGGSTGVKD 126
STDPAHSLSDSFAQDLTGG LV V+G + PL+ALEINP+K+ E+FR ++ GG G K
Sbjct: 209 STDPAHSLSDSFAQDLTGGKLVPVEGVNSPLYALEINPDKSMEEFRTAGQKLGGG-GAKS 385
Query: 127 FMDGMGLGMIVDQ 139
M MGLG++ DQ
Sbjct: 386 LMQSMGLGVVADQ 424
Score = 85.9 bits (211), Expect(2) = 3e-63
Identities = 71/206 (34%), Positives = 112/206 (53%)
Frame = +1
Query: 150 RVTGYTSSWTG*SYCDFQGYTIS*ITGI*YVHSNCI*YGTYWSYITFIVLA*FLGCIHRE 209
R ++S+W *++C+ G IS*+ I*YV S+ *Y T SY + ++A* LG I+R+
Sbjct: 445 RAVAHSSTWNR*NHCNC*GDAIS*V*RI*YVQSDSF*YSTNGSYTSATIIA*LLGWIYRQ 624
Query: 210 DTKAKTKDSFCYLCN*ICVWARKSSSRCY*QIGEA*GEDDKSARAFS*H*FDRICYCYNP 269
+ + + + I V R +S + + Q +A GE K+ +F * RI +
Sbjct: 625 TYEDEDEIRVSFQ---ILVGKRTASKQSFRQT*KAQGESGKNTGSFP*FRHYRIHNSDDS 795
Query: 270 HGDGS**VIQIECLIEERKCSCEETHCQSTSSTICI*LQVLCNEKKGSNTCS*YDSE*SR 329
+G G+**+I++ C+++ERKC+C +T+ + + T LQVL +E KGSN +* +* R
Sbjct: 796 YGYGN**IIKVTCILKERKCTC*KTN-R*SGLTSYDGLQVLFDETKGSNARN*NYPQ*FR 972
Query: 330 TLQPINDPSTTC*C*DKRCSCSQISG 355
T T +K CS S I G
Sbjct: 973 TWWIEIMSGTLGGYGNKGCSSSHIHG 1050
>AW256389 similar to GP|21689765|gb putative arsA-like protein hASNA-I
{Arabidopsis thaliana}, partial (33%)
Length = 601
Score = 58.5 bits (140), Expect = 4e-09
Identities = 32/78 (41%), Positives = 49/78 (62%)
Frame = +1
Query: 33 KYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDG 92
K+ GG GVGKT+C++ L++ A+ L++STDPAH+LSD+F Q T + V+G
Sbjct: 31 KWVFFGGNCGVGKTTCSSILSILLASVRSSVLIISTDPAHNLSDAFQQRFTKTPTL-VNG 207
Query: 93 PDYPLFALEINPEKARED 110
L+A+E++P ED
Sbjct: 208 FS-NLYAMEVDPTVEHED 258
>TC82374 similar to GP|21689765|gb|AAM67526.1 putative arsA-like protein
hASNA-I {Arabidopsis thaliana}, partial (97%)
Length = 1176
Score = 50.1 bits (118), Expect = 1e-06
Identities = 29/76 (38%), Positives = 47/76 (61%)
Frame = +2
Query: 35 YMLGGKGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGPD 94
+ L K +GKT+C++ L++ A+ L++STDPAH+LSD+F Q T + V+G
Sbjct: 44 FSLVAKVVLGKTTCSSILSILLASVRSSVLIISTDPAHNLSDAFQQRFTKTPTL-VNGFS 220
Query: 95 YPLFALEINPEKARED 110
L+A+E++P ED
Sbjct: 221 -NLYAMEVDPTVEHED 265
>BI308037 weakly similar to GP|12583663|dbj chloroplast RelA homologue 2
{Oryza sativa (japonica cultivar-group)}, partial (10%)
Length = 532
Score = 32.7 bits (73), Expect = 0.22
Identities = 20/53 (37%), Positives = 28/53 (52%), Gaps = 6/53 (11%)
Frame = -2
Query: 1 MIITVFLLSVKSVAAPTESISVFDDMVAGTERKYYMLGGK------GGVGKTS 47
++ F+LSVKS+ APT SI++F TE +L + G VGK S
Sbjct: 240 LVDNTFILSVKSLKAPTSSITIFPPPGTSTEHALGVLDHRRVVERVGRVGKNS 82
>TC87602 similar to GP|21741453|emb|CAD41397. OJ000223_09.9 {Oryza sativa},
partial (95%)
Length = 1263
Score = 31.2 bits (69), Expect = 0.65
Identities = 19/39 (48%), Positives = 23/39 (58%), Gaps = 6/39 (15%)
Frame = +2
Query: 25 DMVAGTER------KYYMLGGKGGVGKTSCAASLAVKFA 57
DMVA ER K +L GKGGVGK++ +A LA A
Sbjct: 218 DMVAIAERMATVKHKILVLSGKGGVGKSTFSAQLAFALA 334
>TC78319 homologue to GP|21592386|gb|AAM64337.1 mrp protein putative
{Arabidopsis thaliana}, partial (60%)
Length = 1243
Score = 29.6 bits (65), Expect = 1.9
Identities = 22/67 (32%), Positives = 33/67 (48%)
Frame = +2
Query: 40 KGGVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVDGPDYPLFA 99
KGGVGK++ A +LA A+ G + F D+ G +L + P+ +
Sbjct: 584 KGGVGKSTVAVNLAYTLADMGARVGI------------FDADIYGPSLPTMVSPENRI-- 721
Query: 100 LEINPEK 106
LE+NPEK
Sbjct: 722 LEMNPEK 742
>TC82661 similar to PIR|H86386|H86386 hypothetical protein AAG38125.1
[imported] - Arabidopsis thaliana, partial (27%)
Length = 1044
Score = 28.5 bits (62), Expect = 4.2
Identities = 21/56 (37%), Positives = 28/56 (49%)
Frame = +2
Query: 11 KSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGHPTLVV 66
K + A S S + VA T+ ++ GKGG GKT+ A A FA G T +V
Sbjct: 98 KPLMAVASSSSSSSEDVASTKLLTFL--GKGGSGKTTAAIFAAQHFAMAGLNTCLV 259
>BG648713 similar to GP|16924108|gb putative nucleotide-binding protein
{Oryza sativa}, partial (69%)
Length = 754
Score = 28.5 bits (62), Expect = 4.2
Identities = 13/32 (40%), Positives = 20/32 (61%)
Frame = +1
Query: 27 VAGTERKYYMLGGKGGVGKTSCAASLAVKFAN 58
+ G + + GKGGVGK++ A +LAV A+
Sbjct: 139 IDGVKDTIAIASGKGGVGKSTTAVNLAVALAS 234
>TC82632 homologue to GP|18086370|gb|AAL57645.1 At2g47390/T8I13.23
{Arabidopsis thaliana}, partial (26%)
Length = 848
Score = 28.1 bits (61), Expect = 5.5
Identities = 15/50 (30%), Positives = 24/50 (48%)
Frame = +3
Query: 42 GVGKTSCAASLAVKFANNGHPTLVVSTDPAHSLSDSFAQDLTGGALVQVD 91
G+G TS LA +FA PT+ + + +DS+ + L A V+
Sbjct: 159 GIGSTSALLWLAKRFAILSGPTIPIIGEGEVEANDSYVEQLVASAEAAVE 308
>BQ750889
Length = 789
Score = 28.1 bits (61), Expect = 5.5
Identities = 22/84 (26%), Positives = 38/84 (45%)
Frame = -1
Query: 2 IITVFLLSVKSVAAPTESISVFDDMVAGTERKYYMLGGKGGVGKTSCAASLAVKFANNGH 61
++ V LSVK ++ + + + + R +LGG+ G +T+ S +K G
Sbjct: 522 VVLVEALSVKEISRRKSTNPCGEHRIRSSNR*GMILGGRPGSSRTA-TTSRTMKSGLQGG 346
Query: 62 PTLVVSTDPAHSLSDSFAQDLTGG 85
+S D L DS +D+T G
Sbjct: 345 MLYTLSID----LLDSHVEDVTNG 286
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.347 0.152 0.531
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,022,003
Number of Sequences: 36976
Number of extensions: 192136
Number of successful extensions: 1429
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1417
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1427
length of query: 359
length of database: 9,014,727
effective HSP length: 97
effective length of query: 262
effective length of database: 5,428,055
effective search space: 1422150410
effective search space used: 1422150410
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144928.8


