

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144723.3 + phase: 0
(127 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
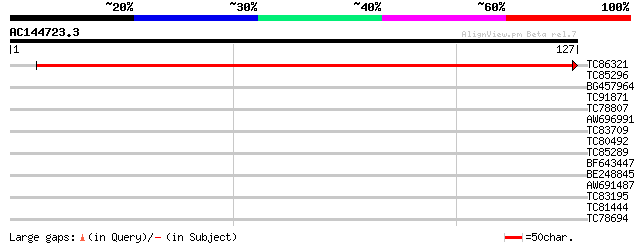
Score E
Sequences producing significant alignments: (bits) Value
TC86321 similar to GP|3868853|dbj|BAA34247.1 GPI-anchored protei... 253 2e-68
TC85296 weakly similar to GP|2505870|emb|CAA72908.1 hypothetical... 28 0.73
BG457964 similar to SP|O22224|Y240 Protein At2g41620. [Mouse-ear... 28 0.95
TC91871 similar to GP|5725518|gb|AAD48086.1| replication origin ... 27 2.1
TC78807 similar to GP|21593058|gb|AAM65007.1 unknown {Arabidopsi... 27 2.8
AW696991 weakly similar to PIR|E86300|E863 protein F3O9.30 [impo... 26 3.6
TC83709 homologue to PIR|S55344|S55344 outer envelope membrane p... 26 3.6
TC80492 similar to GP|21594204|gb|AAM65980.1 unknown {Arabidopsi... 25 6.2
TC85289 similar to SP|Q9SQI2|GIGA_ARATH GIGANTEA protein. [Mouse... 25 6.2
BF643447 similar to GP|16323334|gb AT3g15355/MJK13_1 {Arabidopsi... 25 8.0
BE248845 weakly similar to PIR|T48246|T48 ribonuclease II-like p... 25 8.0
AW691487 similar to SP|Q9THX6|TL29_ Putative L-ascorbate peroxid... 25 8.0
TC83195 homologue to GP|6648199|gb|AAF21197.1| unknown protein {... 25 8.0
TC81444 PIR|C89872|C89872 hypothetical protein [imported] - Stap... 25 8.0
TC78694 similar to GP|10177333|dbj|BAB10682. 2-oxoglutarate dehy... 25 8.0
>TC86321 similar to GP|3868853|dbj|BAA34247.1 GPI-anchored protein {Vigna
radiata}, partial (74%)
Length = 889
Score = 253 bits (645), Expect = 2e-68
Identities = 119/121 (98%), Positives = 119/121 (98%)
Frame = +2
Query: 7 APAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 66
A GCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY
Sbjct: 299 AKKGCSVNFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTNDCASTMFSY 478
Query: 67 INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILL 126
INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILL
Sbjct: 479 INLYGRYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHTPSLVLVLTACIFLILL 658
Query: 127 F 127
F
Sbjct: 659 F 661
>TC85296 weakly similar to GP|2505870|emb|CAA72908.1 hypothetical protein
{Arabidopsis thaliana}, partial (3%)
Length = 774
Score = 28.5 bits (62), Expect = 0.73
Identities = 19/74 (25%), Positives = 32/74 (42%)
Frame = +1
Query: 49 ADVINDLTNDCASTMFSYINLYGRYPPGLFASECREGKEGLACDALPPSVSADDTANQIV 108
A+V ++L NDCA + + + PGLF C + C+ ++ IV
Sbjct: 349 ANVHHELANDCAFNITDMYSCKDKM*PGLFYYPC-----PILCNTYKKKKKKKFLSSFIV 513
Query: 109 HTPSLVLVLTACIF 122
H+ +L L +F
Sbjct: 514 HSHTLQLYFNPSLF 555
>BG457964 similar to SP|O22224|Y240 Protein At2g41620. [Mouse-ear cress]
{Arabidopsis thaliana}, partial (19%)
Length = 670
Score = 28.1 bits (61), Expect = 0.95
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 11/40 (27%)
Frame = -2
Query: 27 CKGPKYPPKECCG------SFKEFACPY-----ADVINDL 55
CK P PP+E C S F+CP+ A +IND+
Sbjct: 306 CKFPTPPPREACAG*LCIVSMMYFSCPHSRWRLAPIINDI 187
>TC91871 similar to GP|5725518|gb|AAD48086.1| replication origin activator 2
{Zea mays}, partial (18%)
Length = 554
Score = 26.9 bits (58), Expect = 2.1
Identities = 15/48 (31%), Positives = 21/48 (43%), Gaps = 1/48 (2%)
Frame = -2
Query: 13 VNFEFLNYTIITSKCKGPKYPPKECCGSFKEF-ACPYADVINDLTNDC 59
+N E + + + KGP P C SFK F + P + NDC
Sbjct: 361 INSERRSLGVTRRETKGPSNPTSTCSPSFKYFGSLPRTVSVTASQNDC 218
>TC78807 similar to GP|21593058|gb|AAM65007.1 unknown {Arabidopsis
thaliana}, partial (74%)
Length = 904
Score = 26.6 bits (57), Expect = 2.8
Identities = 22/95 (23%), Positives = 40/95 (41%), Gaps = 19/95 (20%)
Frame = -3
Query: 32 YPPKECCGSFKEFACPYADVIN---DLTNDC-----ASTMFSYINLYGRYPPGL------ 77
Y K+CC + A PY+ N DL C ST +++++ + P
Sbjct: 548 YYQKDCCSLSHKAAKPYSTTANKSLDLMKQCFASVPKSTS*AFMSVNSTFEPAFTFSASA 369
Query: 78 --FASECREGKEGLACD---ALPPSVSADDTANQI 107
F + + L D +PP+V +D+++ +I
Sbjct: 368 PTFRTSPTRYRSNLTADLGNKIPPAVFSDNSSRRI 264
>AW696991 weakly similar to PIR|E86300|E863 protein F3O9.30 [imported] -
Arabidopsis thaliana, partial (8%)
Length = 779
Score = 26.2 bits (56), Expect = 3.6
Identities = 29/123 (23%), Positives = 49/123 (39%), Gaps = 15/123 (12%)
Frame = -1
Query: 6 IAPAGCSVNFEFLNYTIITSKCKGPKYPPKECCGSF----KEFACPYADVINDLTNDCAS 61
+AP C +F++ ++ + C + ++ C SF C YA ++ + ND AS
Sbjct: 695 VAP-DCGALAKFMSRMLLLTTCCRFAFCHRDVCTSFCISAHRLLCTYACIVI-VHNDTAS 522
Query: 62 TM--------FSYINLYG---RYPPGLFASECREGKEGLACDALPPSVSADDTANQIVHT 110
T ++ +Y R PP L + LPP +S + + H
Sbjct: 521 TSQYTSVARHLPHLRVYALCSRVPPFLTRTSHPHTPR-----PLPPPLSVCSPPSSLTHL 357
Query: 111 PSL 113
PSL
Sbjct: 356 PSL 348
>TC83709 homologue to PIR|S55344|S55344 outer envelope membrane protein
OEP75 precursor - garden pea, partial (36%)
Length = 991
Score = 26.2 bits (56), Expect = 3.6
Identities = 18/60 (30%), Positives = 26/60 (43%), Gaps = 4/60 (6%)
Frame = +2
Query: 2 TLFHIAPAGCSV----NFEFLNYTIITSKCKGPKYPPKECCGSFKEFACPYADVINDLTN 57
TL + P G N + LN ++ S P++ E+A PY D +NDL N
Sbjct: 326 TLASLQPGGTITFEHRNLQGLNRSLTGSVTTSNFLNPQDDLAFKMEYAHPYLDGVNDLRN 505
>TC80492 similar to GP|21594204|gb|AAM65980.1 unknown {Arabidopsis
thaliana}, partial (96%)
Length = 1038
Score = 25.4 bits (54), Expect = 6.2
Identities = 9/23 (39%), Positives = 15/23 (65%)
Frame = +2
Query: 1 MTLFHIAPAGCSVNFEFLNYTII 23
+T+ ++AP GC+V L +T I
Sbjct: 650 LTMVNLAPGGCAVGIPLLYFTAI 718
>TC85289 similar to SP|Q9SQI2|GIGA_ARATH GIGANTEA protein. [Mouse-ear cress]
{Arabidopsis thaliana}, partial (74%)
Length = 3492
Score = 25.4 bits (54), Expect = 6.2
Identities = 12/34 (35%), Positives = 18/34 (52%)
Frame = +3
Query: 49 ADVINDLTNDCASTMFSYINLYGRYPPGLFASEC 82
A+V ++L NDCA + + + PGLF C
Sbjct: 3327 ANVHHELANDCAFNITDMYSCKDKM*PGLFYYPC 3428
>BF643447 similar to GP|16323334|gb AT3g15355/MJK13_1 {Arabidopsis thaliana},
partial (21%)
Length = 671
Score = 25.0 bits (53), Expect = 8.0
Identities = 9/30 (30%), Positives = 19/30 (63%)
Frame = +3
Query: 35 KECCGSFKEFACPYADVINDLTNDCASTMF 64
+ CCG+F + + Y+ V+ ++ C+S +F
Sbjct: 357 RPCCGAFLQPST*YSGVV*NIHGRCSSWLF 446
>BE248845 weakly similar to PIR|T48246|T48 ribonuclease II-like protein -
Arabidopsis thaliana, partial (26%)
Length = 653
Score = 25.0 bits (53), Expect = 8.0
Identities = 8/18 (44%), Positives = 13/18 (71%)
Frame = -1
Query: 77 LFASECREGKEGLACDAL 94
+F+S C E EGL+C+ +
Sbjct: 461 IFSSSCHEDLEGLSCEGI 408
>AW691487 similar to SP|Q9THX6|TL29_ Putative L-ascorbate peroxidase
chloroplast precursor (EC 1.11.1.11), partial (21%)
Length = 562
Score = 25.0 bits (53), Expect = 8.0
Identities = 9/23 (39%), Positives = 14/23 (60%)
Frame = -1
Query: 5 HIAPAGCSVNFEFLNYTIITSKC 27
H P G + NF+F ++ +T KC
Sbjct: 298 HFLPPGQTFNFKF*LFSFLTKKC 230
>TC83195 homologue to GP|6648199|gb|AAF21197.1| unknown protein {Arabidopsis
thaliana}, partial (2%)
Length = 1085
Score = 25.0 bits (53), Expect = 8.0
Identities = 13/43 (30%), Positives = 19/43 (43%), Gaps = 4/43 (9%)
Frame = -2
Query: 10 GCS--VNFEFLNYTIITSKCKGPKY--PPKECCGSFKEFACPY 48
GC + F + T + CK +Y CC +K+F PY
Sbjct: 637 GCRSLLGFRNMGQTNKVNHCKNSQY*TYQNHCCKQYKDFVYPY 509
>TC81444 PIR|C89872|C89872 hypothetical protein [imported] - Staphylococcus
aureus (strain N315), partial (16%)
Length = 1434
Score = 25.0 bits (53), Expect = 8.0
Identities = 10/24 (41%), Positives = 13/24 (53%)
Frame = -3
Query: 5 HIAPAGCSVNFEFLNYTIITSKCK 28
H+A G + + LN I T KCK
Sbjct: 244 HLAAEGARIIVKLLNLVICTQKCK 173
>TC78694 similar to GP|10177333|dbj|BAB10682. 2-oxoglutarate dehydrogenase
E1 component {Arabidopsis thaliana}, partial (29%)
Length = 1224
Score = 25.0 bits (53), Expect = 8.0
Identities = 7/10 (70%), Positives = 9/10 (90%)
Frame = -1
Query: 32 YPPKECCGSF 41
YPP++CCG F
Sbjct: 531 YPPQQCCGHF 502
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.325 0.141 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,204,344
Number of Sequences: 36976
Number of extensions: 79344
Number of successful extensions: 448
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 447
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 448
length of query: 127
length of database: 9,014,727
effective HSP length: 85
effective length of query: 42
effective length of database: 5,871,767
effective search space: 246614214
effective search space used: 246614214
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC144723.3


