

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144514.8 - phase: 0 /pseudo
(555 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
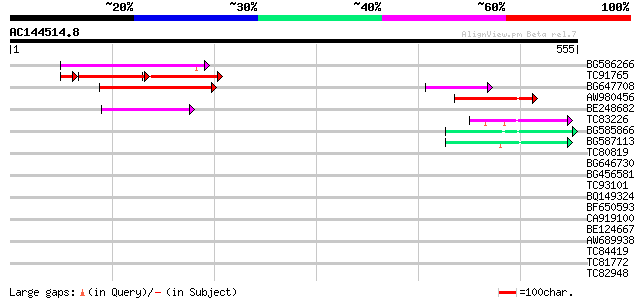
Score E
Sequences producing significant alignments: (bits) Value
BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-li... 117 2e-26
TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative ret... 91 1e-21
BG647708 weakly similar to GP|13786450|gb| putative reverse tran... 95 6e-20
AW980456 80 2e-15
BE248682 similar to GP|18568269|gb putative gag-pol polyprotein ... 66 3e-11
TC83226 weakly similar to PIR|G86419|G86419 probable reverse tra... 55 5e-08
BG585866 48 9e-06
BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At... 44 2e-04
TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non... 40 0.003
BG646730 39 0.007
BG456581 37 0.026
TC93101 36 0.044
BQ149324 29 0.33
BF650593 32 0.83
CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [i... 32 0.83
BE124667 31 1.4
AW689938 weakly similar to GP|22296343|db similar to leucine-ric... 30 1.9
TC84419 similar to SP|Q42920|PME_MEDSA Pectinesterase precursor ... 30 1.9
TC81772 30 3.2
TC82948 29 5.4
>BG586266 similar to GP|7267666|em RNA-directed DNA polymerase-like protein
{Arabidopsis thaliana}, partial (18%)
Length = 789
Score = 117 bits (292), Expect = 2e-26
Identities = 61/150 (40%), Positives = 87/150 (57%), Gaps = 4/150 (2%)
Frame = -3
Query: 50 RAVMVKTGFNNRWIY*MSMCVETVDFSVLVNNDAVGPFIPGRGLRQGDPLSPYMFILCVE 109
R V+ + GF+ WI + CV TV +S L+N G +P RGLRQGDPLSPY+FILC E
Sbjct: 511 REVLTRLGFHGIWISWIMECVSTVSYSFLINGGPQGRVLPSRGLRQGDPLSPYLFILCTE 332
Query: 110 GLSSLIRDAEEHRIISGISICRGAPSISHLLFADDCFLFFKADEGQTQVMKNILITYKAA 169
LS L + A + G+ + R P I+HLLFADD F K++ ++ +I+ Y+AA
Sbjct: 331 VLSGLCQQALRKGTLPGVKVARNCPPINHLLFADDTMFFGKSNASSCAILLSIMDKYRAA 152
Query: 170 SGQAISLPISEI----YCNQNVFDALKASI 195
SG+ I+ S I +Q + D +K +
Sbjct: 151 SGRCIN*TKSAITFSSKTSQAIIDRVKGEL 62
>TC91765 weakly similar to GP|19881779|gb|AAM01180.1 Putative retroelement
{Oryza sativa (japonica cultivar-group)}, partial (1%)
Length = 625
Score = 90.9 bits (224), Expect(2) = 1e-21
Identities = 46/70 (65%), Positives = 53/70 (75%)
Frame = +2
Query: 68 MCVETVDFSVLVNNDAVGPFIPGRGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGI 127
MCVE+ D+ VLVNNDAV P IP RGL+QGD LSPY+FI+CVEGLS LI A+E G
Sbjct: 101 MCVESNDYYVLVNNDAVDPIIPSRGLQQGDHLSPYIFIICVEGLSFLIPHAKERGDTHGT 280
Query: 128 SICRGAPSIS 137
SI RGAP +S
Sbjct: 281 SI*RGAPPVS 310
Score = 69.7 bits (169), Expect = 3e-12
Identities = 41/78 (52%), Positives = 52/78 (66%)
Frame = +3
Query: 131 RGAPSISHLLFADDCFLFFKADEGQTQVMKNILITYKAASGQAISLPISEIYCNQNVFDA 190
RGA ++ LLF + +E Q+MKNILI Y+ SG+AISL S IYC++NV D
Sbjct: 258 RGAIPMA-LLFEGAPLRSLRVEEHHAQIMKNILILYEEDSGKAISLRKS*IYCSRNVPDI 434
Query: 191 LKASITNIIGVQDVLGTC 208
LK SIT I+GVQ +LGTC
Sbjct: 435 LKTSITYILGVQFMLGTC 488
Score = 30.8 bits (68), Expect(2) = 1e-21
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = +1
Query: 50 RAVMVKTGFNNRWIY*M 66
+ +M+K GFNNRWIY M
Sbjct: 46 KEIMIKMGFNNRWIYWM 96
>BG647708 weakly similar to GP|13786450|gb| putative reverse transcriptase
{Oryza sativa}, partial (9%)
Length = 708
Score = 95.1 bits (235), Expect = 6e-20
Identities = 48/114 (42%), Positives = 71/114 (62%)
Frame = +1
Query: 89 PGRGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPSISHLLFADDCFLF 148
P +GLRQGDPLSPY+FILC LS L++ + + GI + R P I+HLLFADD LF
Sbjct: 4 PEKGLRQGDPLSPYLFILCANVLSGLLKREGNKQNLHGIQVARSDPKITHLLFADDSLLF 183
Query: 149 FKADEGQTQVMKNILITYKAASGQAISLPISEIYCNQNVFDALKASITNIIGVQ 202
+A+ + + +L +Y++ASGQ ++ SE+ +QNV + K I I ++
Sbjct: 184 ARANLTEAATIMQVLHSYQSASGQLVNFEKSEVSYSQNVPNQEKEMICQQIAIK 345
Score = 47.8 bits (112), Expect = 1e-05
Identities = 27/66 (40%), Positives = 38/66 (56%), Gaps = 1/66 (1%)
Frame = +3
Query: 408 VDSLIDDETKCWDVPFIQ*VFSKDIAYDVLNTSLVNQVTNDKLIWKAEKNGIYSVKSTFR 467
VD LID +TK W+ I F+ A+ ++ L + DK+IW EK+GIYSV+S
Sbjct: 402 VDELIDYDTKQWNRDLIFHSFNNYAAHQIIKIPLSMRQPEDKIIWHWEKDGIYSVRSAHH 581
Query: 468 -LCVED 472
LC E+
Sbjct: 582 ALCDEN 599
>AW980456
Length = 779
Score = 80.5 bits (197), Expect = 2e-15
Identities = 39/81 (48%), Positives = 57/81 (70%)
Frame = -2
Query: 436 VLNTSLVNQVTNDKLIWKAEKNGIYSVKSTFRLCVEDLMDTTHLRCPEFWMGIWRLKVPP 495
+++T L++ D+LIWK EK+G Y VKS +R CVE+L D+++L P W GIW+LKVPP
Sbjct: 244 IMSTPLISHA*LDRLIWKDEKHGKYYVKSAYRFCVEELFDSSYLHRPGNWSGIWKLKVPP 65
Query: 496 KIKKSHVDDMLDEILPTRMQL 516
K+ ++ V M LPTR++L
Sbjct: 64 KV-QNLVWRMCRGCLPTRIRL 5
>BE248682 similar to GP|18568269|gb putative gag-pol polyprotein {Zea mays},
partial (1%)
Length = 441
Score = 66.2 bits (160), Expect = 3e-11
Identities = 31/91 (34%), Positives = 51/91 (55%)
Frame = +3
Query: 91 RGLRQGDPLSPYMFILCVEGLSSLIRDAEEHRIISGISICRGAPSISHLLFADDCFLFFK 150
RGL+QGDPL+P++F+L EG+S L+++A + G + RG +SHL +ADD
Sbjct: 24 RGLKQGDPLAPFLFLLVAEGISGLMKNAVNRNLFQGFDVKRGGTRVSHLQYADDTLCIGM 203
Query: 151 ADEGQTQVMKNILITYKAASGQAISLPISEI 181
+K +L ++ ASG ++ S +
Sbjct: 204 PTVDNLWTLKALLQGFEMASGLKVNFHKSSL 296
>TC83226 weakly similar to PIR|G86419|G86419 probable reverse transcriptase
100033-105622 [imported] - Arabidopsis thaliana, partial
(2%)
Length = 885
Score = 55.5 bits (132), Expect = 5e-08
Identities = 34/106 (32%), Positives = 49/106 (46%), Gaps = 5/106 (4%)
Frame = +3
Query: 451 IWKAEKNGIYSVKS---TFRLCVEDLMDTTHLRCPE--FWMGIWRLKVPPKIKKSHVDDM 505
+W GIYSVKS T R ++ T E W IW L P+ K + +
Sbjct: 3 MWMHNPTGIYSVKSGYNTLRTWQTQQINNTSTSSDETLIWKKIWSLHTIPR-HKVLLWRI 179
Query: 506 LDEILPTRMQLKDKGV*CPTYCVSYEHQEEDLEHIFFQCPFSVQVW 551
L++ LP R L+ +G+ C C + E + H+F CP S +VW
Sbjct: 180 LNDSLPVRSSLRKRGIQCYPLCPRCHSKTETITHLFMSCPLSKRVW 317
>BG585866
Length = 828
Score = 48.1 bits (113), Expect = 9e-06
Identities = 36/130 (27%), Positives = 49/130 (37%), Gaps = 1/130 (0%)
Frame = +3
Query: 427 VFSKDIAYDVLNTSL-VNQVTNDKLIWKAEKNGIYSVKSTFRLCVEDLMDTTHLRCPEFW 485
+ DIA + NT L N D IW NG+Y+ KS + + + W
Sbjct: 282 ILPTDIAEVINNTHLNFNASIGDAYIWPHNSNGVYTAKSGYSWILSQTETVNYNNSS--W 455
Query: 486 MGIWRLKVPPKIKKSHVDDMLDEILPTRMQLKDKGV*CPTYCVSYEHQEEDLEHIFFQCP 545
IWRLK+P K K + +PT L + + C EE H C
Sbjct: 456 SWIWRLKIPEKY-KFFLWLACHNAVPTLSLLNHRNMVNSAICSRCGEHEESFFHCVRDCR 632
Query: 546 FSVQVWQFAG 555
FS +W G
Sbjct: 633 FSKIIWHKIG 662
>BG587113 weakly similar to PIR|A84888|A8 hypothetical protein At2g45230
[imported] - Arabidopsis thaliana, partial (10%)
Length = 767
Score = 43.9 bits (102), Expect = 2e-04
Identities = 26/131 (19%), Positives = 53/131 (39%), Gaps = 6/131 (4%)
Frame = -3
Query: 427 VFSKDIAYDVLNTSLVNQVTNDKLIWKAEKNGIYSVKSTFRLCVEDLMDTTHL------R 480
+F + +L+ + D W+ K+G YSVKS + + +
Sbjct: 489 IFPEGTRRKILSIHPQGPIGEDSYSWEYSKSGHYSVKSGYYVQTNIIAAANQRGTVDQPS 310
Query: 481 CPEFWMGIWRLKVPPKIKKSHVDDMLDEILPTRMQLKDKGV*CPTYCVSYEHQEEDLEHI 540
+ + +W+ PK++ + + LPT ++ + + C + E + HI
Sbjct: 309 LDDLYQRVWKYNTSPKVRH-FLWRCISNSLPTAANMRSRHISKDGSCSRCGMESETVNHI 133
Query: 541 FFQCPFSVQVW 551
FQCP++ +W
Sbjct: 132 LFQCPYARLIW 100
>TC80819 weakly similar to GP|10140689|gb|AAG13524.1 putative non-LTR
retroelement reverse transcriptase {Oryza sativa
(japonica cultivar-group)}, partial (2%)
Length = 1262
Score = 39.7 bits (91), Expect = 0.003
Identities = 32/118 (27%), Positives = 47/118 (39%), Gaps = 2/118 (1%)
Frame = +2
Query: 436 VLNTSLVNQVTNDKLIWKAEKNGIYSVKSTFRLCVEDLMDTTHLRCPEFWMGIWRLKVPP 495
+LN ++ NDK W + YSVK +R + T H+ +W +P
Sbjct: 482 LLNNFVLQDNVNDKWRWLLDPVNGYSVKVFYRY----ITSTGHISDRSLVDDVWHKHIPS 649
Query: 496 KIKKSHVDDMLDEILPTRMQLKDKGV*CPT--YCVSYEHQEEDLEHIFFQCPFSVQVW 551
K+ V +L LPT+ L +GV T CV E H+F C +W
Sbjct: 650 KV-SLFVWRLLRNRLPTKDNLVHRGVLLATNAACVCGCVDSESTTHLFLHCNVFCSLW 820
>BG646730
Length = 799
Score = 38.5 bits (88), Expect = 0.007
Identities = 26/90 (28%), Positives = 40/90 (43%)
Frame = +1
Query: 379 ISILDEPWIKDGRSINDNLPGADFVRNFCVDSLIDDETKCWDVPFIQ*VFSKDIAYDVLN 438
+ I + W+ + P DF C+ LID TK ++ VFS A +LN
Sbjct: 481 VRIWKDDWLPNQTGFKVCSPMVDFEEATCISELIDTNTKSSKRDLVRNVFSAFEAKQILN 660
Query: 439 TSLVNQVTNDKLIWKAEKNGIYSVKSTFRL 468
L ++ D+ I E++ YSV+S L
Sbjct: 661 IPLSWRLWPDRRILYWERDRNYSVRSAHHL 750
>BG456581
Length = 683
Score = 36.6 bits (83), Expect = 0.026
Identities = 23/81 (28%), Positives = 35/81 (42%), Gaps = 3/81 (3%)
Frame = +2
Query: 476 TTHLRCPEFWMGIWRLKVPPKIKKSHVD---DMLDEILPTRMQLKDKGV*CPTYCVSYEH 532
T+H C + IW +PP SH + + LPT L +G + C
Sbjct: 44 TSHTSCAPWASTIWNSCIPP----SHSFICWRLAHDRLPTDDNLSSRGCALVSMCSFCLE 211
Query: 533 QEEDLEHIFFQCPFSVQVWQF 553
Q E +H+F +C F V +W +
Sbjct: 212 QVETSDHLFLRCKFVVTLWSW 274
>TC93101
Length = 675
Score = 35.8 bits (81), Expect = 0.044
Identities = 16/42 (38%), Positives = 20/42 (47%)
Frame = -1
Query: 510 LPTRMQLKDKGV*CPTYCVSYEHQEEDLEHIFFQCPFSVQVW 551
LP R +L +GV CP C E HIF C + +VW
Sbjct: 663 LPVRXELNKRGVNCPPLCPRCYFNLETTNHIFMSCERTQRVW 538
>BQ149324
Length = 374
Score = 28.9 bits (63), Expect(2) = 0.33
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = +3
Query: 510 LPTRMQLKDKGV*CPTYCVSYEHQEED 536
LPTR++LKDK V CP C ED
Sbjct: 3 LPTRVRLKDKRVTCPMDCTLCTVGSED 83
Score = 22.7 bits (47), Expect(2) = 0.33
Identities = 7/13 (53%), Positives = 9/13 (68%)
Frame = +1
Query: 539 HIFFQCPFSVQVW 551
H+ FQC S+ VW
Sbjct: 91 HLIFQCSSSLNVW 129
>BF650593
Length = 486
Score = 31.6 bits (70), Expect = 0.83
Identities = 19/49 (38%), Positives = 24/49 (48%), Gaps = 2/49 (4%)
Frame = +3
Query: 505 MLDEILPTRMQLKDKGV*CPT--YCVSYEHQEEDLEHIFFQCPFSVQVW 551
+ L TR L +GV CV +EE + H FF+CPFS VW
Sbjct: 339 LFQNXLATRDNLSKRGVLDQNSIXCVXDCGREESVSHFFFECPFS-XVW 482
>CA919100 homologue to PIR|G90291|G902 endoglucanase precursor [imported] -
Sulfolobus solfataricus, partial (3%)
Length = 789
Score = 31.6 bits (70), Expect = 0.83
Identities = 22/67 (32%), Positives = 32/67 (46%), Gaps = 3/67 (4%)
Frame = -2
Query: 488 IWRLKVPPKIKKSHVDDMLDEILPTRMQLKDKGV*CPT---YCVSYEHQEEDLEHIFFQC 544
IW +VP K+ +L + LPT+ L +GV PT CVS E +H+F C
Sbjct: 584 IWHRQVPLKVSV-FAWRLLRDRLPTKSNLIYRGV-IPTEAGLCVSGCGALESAQHLFLSC 411
Query: 545 PFSVQVW 551
+ +W
Sbjct: 410 SYFASLW 390
>BE124667
Length = 540
Score = 30.8 bits (68), Expect = 1.4
Identities = 10/30 (33%), Positives = 18/30 (59%)
Frame = -1
Query: 524 PTYCVSYEHQEEDLEHIFFQCPFSVQVWQF 553
P+ C S + E H+FF+C F+ ++W +
Sbjct: 393 PSMCSSCQASSETTFHLFFECNFATKMWSW 304
>AW689938 weakly similar to GP|22296343|db similar to leucine-rich repeat
transmembrane protein kinase 1, partial (16%)
Length = 655
Score = 30.4 bits (67), Expect = 1.9
Identities = 12/40 (30%), Positives = 21/40 (52%)
Frame = +3
Query: 162 ILITYKAASGQAISLPISEIYCNQNVFDALKASITNIIGV 201
+ +TY S A+SLPI ++ N + D L + +G+
Sbjct: 501 VSLTYLNVSNNALSLPIGXVFANHSHLDTLDYHLITFLGI 620
>TC84419 similar to SP|Q42920|PME_MEDSA Pectinesterase precursor (EC
3.1.1.11) (Pectin methylesterase) (PE) (P65). [Alfalfa],
partial (53%)
Length = 732
Score = 30.4 bits (67), Expect = 1.9
Identities = 22/56 (39%), Positives = 30/56 (53%)
Frame = -3
Query: 428 FSKDIAYDVLNTSLVNQVTNDKLIWKAEKNGIYSVKSTFRLCVEDLMDTTHLRCPE 483
FS D + VL S N +T DK+ NG +V+ +F LCVE ++ T HL E
Sbjct: 328 FSND-EFAVLENS--NCITKDKV--DCSGNGAVAVELSFGLCVECILVTVHLTVVE 176
>TC81772
Length = 982
Score = 29.6 bits (65), Expect = 3.2
Identities = 18/66 (27%), Positives = 34/66 (51%), Gaps = 2/66 (3%)
Frame = -3
Query: 436 VLNTSLVNQVTNDKLIWKAEKNGIYSVKSTFRLCVEDLMDTTHLRCPE--FWMGIWRLKV 493
+L +L+ D+ IW + G +VK+ + +C E L++ L+ F++ IW +
Sbjct: 452 ILKITLLRLNKPDERIWIFDGRGTNAVKNAYIVCAEHLVNIKELKIQGEIFYVEIWE-*L 276
Query: 494 PPKIKK 499
PK+ K
Sbjct: 275 SPKLSK 258
>TC82948
Length = 705
Score = 28.9 bits (63), Expect = 5.4
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = +3
Query: 535 EDLEHIFFQCPFSVQVWQF 553
E H+FF+C SVQ+W +
Sbjct: 468 ESTHHLFFECSVSVQIWDW 524
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.343 0.152 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,174,726
Number of Sequences: 36976
Number of extensions: 313452
Number of successful extensions: 2462
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 1768
Number of HSP's successfully gapped in prelim test: 61
Number of HSP's that attempted gapping in prelim test: 649
Number of HSP's gapped (non-prelim): 1884
length of query: 555
length of database: 9,014,727
effective HSP length: 101
effective length of query: 454
effective length of database: 5,280,151
effective search space: 2397188554
effective search space used: 2397188554
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144514.8


