

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144483.8 + phase: 0
(253 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
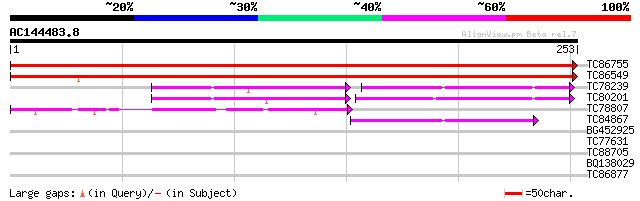
Score E
Sequences producing significant alignments: (bits) Value
TC86755 similar to GP|7331143|gb|AAF60293.1| chaperonin 21 precu... 486 e-138
TC86549 similar to GP|7331143|gb|AAF60293.1| chaperonin 21 precu... 379 e-106
TC78239 similar to SP|P34893|CH10_ARATH 10 kDa chaperonin (Prote... 60 1e-09
TC80201 similar to SP|Q96539|CH10_BRANA 10 kDa chaperonin (Prote... 56 2e-08
TC78807 similar to GP|21593058|gb|AAM65007.1 unknown {Arabidopsi... 53 1e-07
TC84867 similar to GP|21554204|gb|AAM63283.1 putative 10kd chape... 49 1e-06
BG452925 similar to GP|11967921|dbj contains EST AU078264(S21150... 30 1.2
TC77631 similar to PIR|T51579|T51579 cellulose synthase catalyti... 28 2.6
TC88705 similar to GP|10177834|dbj|BAB11263. gene_id:MCD7.9~unkn... 28 3.4
BQ138029 weakly similar to GP|1764098|gb|A putative permease {Ur... 27 7.5
TC86877 similar to GP|8953376|emb|CAB96649.1 putative protein {A... 27 7.5
>TC86755 similar to GP|7331143|gb|AAF60293.1| chaperonin 21 precursor
{Lycopersicon esculentum}, partial (90%)
Length = 985
Score = 486 bits (1250), Expect = e-138
Identities = 252/253 (99%), Positives = 253/253 (99%)
Frame = +3
Query: 1 MATTQLTASSISTRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPKHT 60
MATTQLTASSISTRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPKHT
Sbjct: 51 MATTQLTASSISTRNLSSFERLRTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPKHT 230
Query: 61 TVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAG 120
TVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAG
Sbjct: 231 TVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAG 410
Query: 121 AQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTAG 180
AQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTAG
Sbjct: 411 AQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTAG 590
Query: 181 GLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDGSDY 240
GLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDGSDY
Sbjct: 591 GLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDGSDY 770
Query: 241 IALRASDVMAILS 253
IALRAS+VMAILS
Sbjct: 771 IALRASNVMAILS 809
>TC86549 similar to GP|7331143|gb|AAF60293.1| chaperonin 21 precursor
{Lycopersicon esculentum}, partial (92%)
Length = 1134
Score = 379 bits (973), Expect = e-106
Identities = 190/254 (74%), Positives = 228/254 (88%), Gaps = 1/254 (0%)
Frame = +1
Query: 1 MATTQLTASSISTRNLSSFERLRTSSIQF-PRNNARIAILTQWSLPSMVVKAATVVAPKH 59
MA++QLTASSISTR+++SFE LR SS+QF + RI + S +++VKAATVVAPK+
Sbjct: 76 MASSQLTASSISTRSVASFEGLRPSSVQFHSARHVRIGNPARRSFKALIVKAATVVAPKY 255
Query: 60 TTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKA 119
T +KPLGDRVLVKIK+AEEKTQ GILLPS+AQ+KPQGGEVVA+G+GKT+GKN+V+ISVK
Sbjct: 256 TAIKPLGDRVLVKIKEAEEKTQSGILLPSSAQTKPQGGEVVAVGEGKTLGKNQVEISVKP 435
Query: 120 GAQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTA 179
GAQVVYSKYAGTEV+F+G KHLILKD+DIVGILET+++KDLKPLNDRVLIK+ +AEEKT+
Sbjct: 436 GAQVVYSKYAGTEVDFNGEKHLILKDDDIVGILETDDIKDLKPLNDRVLIKIEKAEEKTS 615
Query: 180 GGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSILPGNTVLYSKYAGNDFKGKDGSD 239
GGL LTEATK+KPS GTV+AVGPG VD+EGNRKPL++ GNTVLYSKYAGNDFKGKDG +
Sbjct: 616 GGLYLTEATKEKPSFGTVVAVGPGLVDEEGNRKPLTVASGNTVLYSKYAGNDFKGKDGFE 795
Query: 240 YIALRASDVMAILS 253
YI LR+SDVMA+LS
Sbjct: 796 YITLRSSDVMAVLS 837
>TC78239 similar to SP|P34893|CH10_ARATH 10 kDa chaperonin (Protein CPN10)
(Protein groES). [Mouse-ear cress] {Arabidopsis
thaliana}, partial (96%)
Length = 546
Score = 59.7 bits (143), Expect = 1e-09
Identities = 36/95 (37%), Positives = 52/95 (53%)
Frame = +2
Query: 158 KDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPLSIL 217
K L P +R+L++ KT+ G+LL E T S G V+AVGPG D GN P+S+
Sbjct: 26 KRLIPTFNRILVEKIIPPSKTSAGILLPEKTSQLNS-GKVVAVGPGSRDKSGNLIPVSVK 202
Query: 218 PGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
G+ VL +Y G+ K D ++ R D++ IL
Sbjct: 203 EGDHVLLPEYGGSQIK-LDDKEFHLFRDEDILGIL 304
Score = 59.7 bits (143), Expect = 1e-09
Identities = 34/90 (37%), Positives = 53/90 (58%), Gaps = 1/90 (1%)
Frame = +2
Query: 64 PLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDG-KTVGKNKVDISVKAGAQ 122
P +R+LV+ KT GILLP S+ G+VVA+G G + N + +SVK G
Sbjct: 38 PTFNRILVEKIIPPSKTSAGILLPEKT-SQLNSGKVVAVGPGSRDKSGNLIPVSVKEGDH 214
Query: 123 VVYSKYAGTEVEFDGSKHLILKDEDIVGIL 152
V+ +Y G++++ D + + +DEDI+GIL
Sbjct: 215 VLLPEYGGSQIKLDDKEFHLFRDEDILGIL 304
>TC80201 similar to SP|Q96539|CH10_BRANA 10 kDa chaperonin (Protein CPN10)
(Protein GROES). [Rape] {Brassica napus}, partial (97%)
Length = 755
Score = 55.8 bits (133), Expect = 2e-08
Identities = 36/90 (40%), Positives = 51/90 (56%), Gaps = 1/90 (1%)
Frame = +1
Query: 64 PLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKV-DISVKAGAQ 122
PL +RVLV+ KT GILLP SK G+VVA+G G K+ ++VK G
Sbjct: 55 PLFNRVLVEKIVPPSKTTAGILLPEKI-SKLNSGKVVAVGPGVHGKDGKLLPVAVKEGDT 231
Query: 123 VVYSKYAGTEVEFDGSKHLILKDEDIVGIL 152
V+ +Y G EV+ D ++ + D+DI+G L
Sbjct: 232 VLLPEYGGVEVKLDHKEYYLYGDDDILGTL 321
Score = 54.3 bits (129), Expect = 4e-08
Identities = 35/98 (35%), Positives = 51/98 (51%)
Frame = +1
Query: 155 EEVKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRKPL 214
E K L PL +RVL++ KT G+LL E K + G V+AVGPG +G P+
Sbjct: 34 EMAKRLIPLFNRVLVEKIVPPSKTTAGILLPEKIS-KLNSGKVVAVGPGVHGKDGKLLPV 210
Query: 215 SILPGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
++ G+TVL +Y G + K D +Y D++ L
Sbjct: 211 AVKEGDTVLLPEYGGVEVK-LDHKEYYLYGDDDILGTL 321
>TC78807 similar to GP|21593058|gb|AAM65007.1 unknown {Arabidopsis
thaliana}, partial (74%)
Length = 904
Score = 53.1 bits (126), Expect = 1e-07
Identities = 48/158 (30%), Positives = 79/158 (49%), Gaps = 5/158 (3%)
Frame = +1
Query: 1 MATTQLTASS---ISTRNLSSFERLRTSSIQFPRNNARI-AILTQWSLPSMVVKAATVVA 56
MA+T LT + + +SF R ++ RN+ ++ AI +W PS VV
Sbjct: 94 MASTFLTIPTTPFLHKTPTASFSTQRLPILK--RNSLKVNAIAKKWE-PSKVV------- 243
Query: 57 PKHTTVKPLGDRVLVKIKDAEEKTQGGILLPSTAQSKPQGGEVVAIGDGKTVGKNKVDIS 116
P DRVL+++++ EKT GGILLP +A E +G+ VG +
Sbjct: 244 -------PQADRVLIRLEELSEKTAGGILLPKSAVK----FERYLVGEVLNVGAEAE--N 384
Query: 117 VKAGAQVVYSKYAGTEVEF-DGSKHLILKDEDIVGILE 153
VKAG++V+++ EV+ +KH +K D++ ++E
Sbjct: 385 VKAGSKVLFTDINAYEVDLGTDAKHCFIKSSDLLAVVE 498
Score = 38.1 bits (87), Expect = 0.003
Identities = 26/99 (26%), Positives = 52/99 (52%), Gaps = 2/99 (2%)
Frame = +1
Query: 156 EVKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATK--DKPSIGTVIAVGPGPVDDEGNRKP 213
E + P DRVLI++ E EKTAGG+LL ++ ++ +G V+ VG +
Sbjct: 226 EPSKVVPQADRVLIRLEELSEKTAGGILLPKSAVKFERYLVGEVLNVG---------AEA 378
Query: 214 LSILPGNTVLYSKYAGNDFKGKDGSDYIALRASDVMAIL 252
++ G+ VL++ + + + +++SD++A++
Sbjct: 379 ENVKAGSKVLFTDINAYEVDLGTDAKHCFIKSSDLLAVV 495
>TC84867 similar to GP|21554204|gb|AAM63283.1 putative 10kd chaperonin
{Arabidopsis thaliana}, partial (80%)
Length = 587
Score = 49.3 bits (116), Expect = 1e-06
Identities = 31/84 (36%), Positives = 44/84 (51%)
Frame = +1
Query: 153 ETEEVKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAVGPGPVDDEGNRK 212
E + K L P + +L + KT+ G+LL E T S G V+AV PG D GN
Sbjct: 97 EKQMAKRLIPTFNCILAEKIVPPSKTSAGVLLPEKTSQLNS-GNVVAVCPGSRDKSGNLI 273
Query: 213 PLSILPGNTVLYSKYAGNDFKGKD 236
PLS+ G+ V+ ++Y G+ K D
Sbjct: 274 PLSVKEGDHVVLAEYGGSQIKLDD 345
Score = 39.3 bits (90), Expect = 0.001
Identities = 22/59 (37%), Positives = 34/59 (57%), Gaps = 1/59 (1%)
Frame = +1
Query: 79 KTQGGILLPSTAQSKPQGGEVVAIGDG-KTVGKNKVDISVKAGAQVVYSKYAGTEVEFD 136
KT G+LLP S+ G VVA+ G + N + +SVK G VV ++Y G++++ D
Sbjct: 169 KTSAGVLLPEKT-SQLNSGNVVAVCPGSRDKSGNLIPLSVKEGDHVVLAEYGGSQIKLD 342
>BG452925 similar to GP|11967921|dbj contains EST AU078264(S21150)~unknown
protein {Oryza sativa (japonica cultivar-group)},
partial (14%)
Length = 685
Score = 29.6 bits (65), Expect = 1.2
Identities = 14/42 (33%), Positives = 21/42 (49%)
Frame = +1
Query: 23 RTSSIQFPRNNARIAILTQWSLPSMVVKAATVVAPKHTTVKP 64
R+ +Q PR R+ QWSLPS + + AP + +P
Sbjct: 157 RSRQLQIPRRRPRL----QWSLPSSLPRRPATTAPHRSAPQP 270
>TC77631 similar to PIR|T51579|T51579 cellulose synthase catalytic subunit
(IRX3) - Arabidopsis thaliana, partial (77%)
Length = 2694
Score = 28.5 bits (62), Expect = 2.6
Identities = 26/91 (28%), Positives = 39/91 (42%), Gaps = 7/91 (7%)
Frame = +1
Query: 166 RVLIKVAEAEEKTAGGLLLTEAT-------KDKPSIGTVIAVGPGPVDDEGNRKPLSILP 218
R+ VA+A++ AGG ++ + T KD P + V G D EGN+ P +
Sbjct: 595 RINALVAKAQKVPAGGWIMQDGTPWPGNNTKDHPGMIQVFLGHSGGHDSEGNQLPRLVY- 771
Query: 219 GNTVLYSKYAGNDFKGKDGSDYIALRASDVM 249
V K G K G+ +R S V+
Sbjct: 772 ---VSREKRPGFQHHKKAGAMNALVRVSAVL 855
>TC88705 similar to GP|10177834|dbj|BAB11263. gene_id:MCD7.9~unknown protein
{Arabidopsis thaliana}, partial (32%)
Length = 1685
Score = 28.1 bits (61), Expect = 3.4
Identities = 22/73 (30%), Positives = 32/73 (43%)
Frame = +1
Query: 110 KNKVDISVKAGAQVVYSKYAGTEVEFDGSKHLILKDEDIVGILETEEVKDLKPLNDRVLI 169
K K + KA + V + T +E + KH ++D I E EVK+ P++
Sbjct: 91 KQKKEAEEKANEKQVKTNEEDTGIENEAEKHSDIEDNFAASIHEKIEVKEDSPVDQ---- 258
Query: 170 KVAEAEEKTAGGL 182
EA EK A L
Sbjct: 259 --DEAGEKLADTL 291
>BQ138029 weakly similar to GP|1764098|gb|A putative permease {Uromyces
fabae}, partial (10%)
Length = 656
Score = 26.9 bits (58), Expect = 7.5
Identities = 11/34 (32%), Positives = 18/34 (52%)
Frame = -2
Query: 158 KDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDK 191
+D +DR+L+ V + K GG+ +T A K
Sbjct: 529 RDFGTQSDRILVPVVPTDSKAKGGIRITSAVLGK 428
>TC86877 similar to GP|8953376|emb|CAB96649.1 putative protein {Arabidopsis
thaliana}, partial (12%)
Length = 1558
Score = 26.9 bits (58), Expect = 7.5
Identities = 33/116 (28%), Positives = 49/116 (41%)
Frame = +2
Query: 85 LLPSTAQSKPQGGEVVAIGDGKTVGKNKVDISVKAGAQVVYSKYAGTEVEFDGSKHLILK 144
++P T K + G + +V +K+ A A SKY +D K L K
Sbjct: 431 VIPLTFDRKKKLGSRPELPVLDSVPASKIGAIYLADAASPISKY------YD-KKKLEPK 589
Query: 145 DEDIVGILETEEVKDLKPLNDRVLIKVAEAEEKTAGGLLLTEATKDKPSIGTVIAV 200
+IV D+KP+ND+V K + E AG + +A DK S V+ V
Sbjct: 590 SNEIVPASGELHTLDVKPVNDKVQPK--DGSENLAGQIQTNQA-GDKNSSNKVVPV 748
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.311 0.131 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,426,240
Number of Sequences: 36976
Number of extensions: 58875
Number of successful extensions: 244
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 228
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 234
length of query: 253
length of database: 9,014,727
effective HSP length: 94
effective length of query: 159
effective length of database: 5,538,983
effective search space: 880698297
effective search space used: 880698297
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC144483.8


