

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144405.9 - phase: 0
(166 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
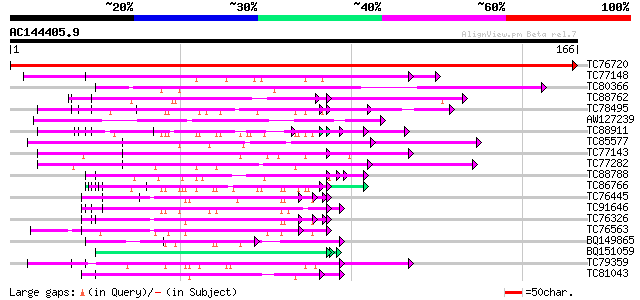
Score E
Sequences producing significant alignments: (bits) Value
TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {... 331 7e-92
TC77148 ENOD20 75 2e-14
TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like... 71 2e-13
TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Ar... 66 7e-12
TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 ... 63 6e-11
AW127239 weakly similar to SP|Q9FPQ6|GP1_C Vegetative cell wall ... 62 1e-10
TC88911 weakly similar to GP|13926280|gb|AAK49610.1 AT5g14920/F2... 61 2e-10
TC85577 homologue to GP|6715512|gb|AAF26445.1| vacuolar H+-ATPas... 60 3e-10
TC77143 similar to GP|559557|gb|AAA51649.1|| arabinogalactan-pro... 60 4e-10
TC77282 homologue to GP|17065300|gb|AAL32804.1 Unknown protein {... 60 5e-10
TC88788 weakly similar to SP|P05790|FBOH_BOMMO Fibroin heavy cha... 60 5e-10
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 59 9e-10
TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - so... 59 9e-10
TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 58 2e-09
TC76326 homologue to PIR|C29356|C29356 hydroxyproline-rich glyco... 58 2e-09
TC76563 homologue to PIR|C29356|C29356 hydroxyproline-rich glyco... 58 2e-09
BQ149865 weakly similar to GP|15128223|db hypothetical protein~s... 58 2e-09
BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melano... 58 2e-09
TC79359 weakly similar to PIR|S49915|S49915 extensin-like protei... 57 3e-09
TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproli... 57 3e-09
>TC76720 similar to GP|11121502|emb|CAC14888. putative extensin {Nicotiana
sylvestris}, partial (51%)
Length = 1290
Score = 331 bits (849), Expect = 7e-92
Identities = 166/166 (100%), Positives = 166/166 (100%)
Frame = +1
Query: 1 MAIPRFSLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTP 60
MAIPRFSLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTP
Sbjct: 487 MAIPRFSLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTP 666
Query: 61 TPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMS 120
TPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMS
Sbjct: 667 TPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMS 846
Query: 121 SGKKAGIAVGVIAAVGVVALGAMVVKKRRQNIQRSEYGYTARRELL 166
SGKKAGIAVGVIAAVGVVALGAMVVKKRRQNIQRSEYGYTARRELL
Sbjct: 847 SGKKAGIAVGVIAAVGVVALGAMVVKKRRQNIQRSEYGYTARRELL 984
Score = 33.9 bits (76), Expect = 0.031
Identities = 43/142 (30%), Positives = 54/142 (37%), Gaps = 29/142 (20%)
Frame = -2
Query: 46 TPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSP---SPSSSPAPSPDEAADNNA 102
T +P + P TPT P P PD+ P SSP P S A S D A D+
Sbjct: 917 TIAPNATTPTAAITPTAIPAFLPELIPPPDDFP-SSPIPVWLIALLSAASSGDGAGDDEG 741
Query: 103 ISH-------TGIGEDGKS----------SGGGMSSGKKAGIAVG-------VIAAVGV- 137
+G G G+S GG +G+ GI VG +A VG
Sbjct: 740 EGEGEEEGELSGEGAGGESE*GDGVGVGVGAGGEFAGEGVGIGVGEGEFSDEFVAGVGAG 561
Query: 138 -VALGAMVVKKRRQNIQRSEYG 158
A AM+ KK R+ R G
Sbjct: 560 ESAEEAMLTKKERRRKTRENLG 495
Score = 32.3 bits (72), Expect = 0.090
Identities = 25/79 (31%), Positives = 28/79 (34%), Gaps = 9/79 (11%)
Frame = +2
Query: 25 SPAPTPATNSSLNSPSP------TPIPTPSPANSPPAPT---PTPTPSPHSDSPPAPSPD 75
SP P P SP P +P P SPP P +P P + P P P
Sbjct: 32 SPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPPPP- 208
Query: 76 NSPSSSPSPSPSSSPAPSP 94
P PSP P SP
Sbjct: 209 KKPYKYPSPPPPVYKYKSP 265
Score = 30.4 bits (67), Expect = 0.34
Identities = 24/80 (30%), Positives = 26/80 (32%), Gaps = 14/80 (17%)
Frame = +2
Query: 27 APTPATNSSLNSPSPTP----IPTPSPA----NSPPAPT------PTPTPSPHSDSPPAP 72
+P P PSP P +P P SPP P P P S PP
Sbjct: 32 SPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPPPPK 211
Query: 73 SPDNSPSSSPSPSPSSSPAP 92
P PS P SP P
Sbjct: 212 KPYKYPSPPPPVYKYKSPPP 271
Score = 28.1 bits (61), Expect = 1.7
Identities = 21/80 (26%), Positives = 28/80 (34%), Gaps = 12/80 (15%)
Frame = +2
Query: 27 APTPATNSSLNSPSPTPIPTPSPANSPPA---PTPTPTPSPHSDSPP------APSPDNS 77
+P P + P P P P+ PP +P P + PP +P P
Sbjct: 2 SPPPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPPPVY 181
Query: 78 PSSSPSPSPSSS---PAPSP 94
SP P P P+P P
Sbjct: 182KYKSPPPPPKKPYKYPSPPP 241
Score = 28.1 bits (61), Expect = 1.7
Identities = 17/57 (29%), Positives = 20/57 (34%)
Frame = +2
Query: 38 SPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
SP P SP P P P+P P +P P SP P +P P
Sbjct: 2 SPPPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPP 172
Score = 28.1 bits (61), Expect = 1.7
Identities = 17/57 (29%), Positives = 20/57 (34%)
Frame = +2
Query: 38 SPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
SP P SP P P P+P P +P P SP P +P P
Sbjct: 161 SPPPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPP 331
Score = 27.3 bits (59), Expect = 2.9
Identities = 21/69 (30%), Positives = 25/69 (35%), Gaps = 11/69 (15%)
Frame = +2
Query: 27 APTPATNSSLNSPSPTP----IPTPSPA----NSPPAPT---PTPTPSPHSDSPPAPSPD 75
+P P PSP P +P P SPP P +P P + P P P
Sbjct: 191 SPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSPPPPVYKYKSP-PPPP 367
Query: 76 NSPSSSPSP 84
P PSP
Sbjct: 368 KKPYKYPSP 394
Score = 26.2 bits (56), Expect = 6.4
Identities = 23/79 (29%), Positives = 28/79 (35%), Gaps = 11/79 (13%)
Frame = +2
Query: 27 APTPATNSSLNSPSPT-PIPTPSPANSPPAPTPTPTPSPHS-DSPPAP-----SPDNSPS 79
+P P + P P +P P P P+P P + SPP P SP
Sbjct: 131 SPPPPVYKYKSPPPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPVY 310
Query: 80 SSPSPSPS----SSPAPSP 94
SP P SP P P
Sbjct: 311 KYKSPPPPVYKYKSPPPPP 367
>TC77148 ENOD20
Length = 1108
Score = 74.7 bits (182), Expect = 2e-14
Identities = 48/132 (36%), Positives = 62/132 (46%), Gaps = 10/132 (7%)
Frame = +3
Query: 5 RFSLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSP 64
R L ++ + + SS P P SS P P PSP + P+P+P+P+PSP
Sbjct: 372 RLGLKLAVVVMVAPVLSSPPPPPSPPTPRSSTPIPHPPRRSLPSPPSPSPSPSPSPSPSP 551
Query: 65 HSDSPPAP--------SPDNSPSSSPSPSPSSSP--APSPDEAADNNAISHTGIGEDGKS 114
S P P SP SPS S SPSPS SP APSP ++ + A S + E
Sbjct: 552 SPRSTPIPHPRKRSPASPSPSPSLSKSPSPSESPSLAPSPSDSVASLAPSSSPSDESPSP 731
Query: 115 SGGGMSSGKKAG 126
+ SSG K G
Sbjct: 732 APSPSSSGSKGG 767
Score = 69.3 bits (168), Expect = 7e-13
Identities = 44/108 (40%), Positives = 59/108 (53%), Gaps = 12/108 (11%)
Frame = +3
Query: 23 ADSPAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPTPTP---SPHSDSPP-----APS 73
+ +P P P S + PSP+P P+PSP+ SP P TP P P SP S SP +PS
Sbjct: 459 SSTPIPHPPRRSLPSPPSPSPSPSPSPSPSPSPRSTPIPHPRKRSPASPSPSPSLSKSPS 638
Query: 74 PDNSPSSSPSPS---PSSSPAPSPDEAADNNAISHTGIGEDGKSSGGG 118
P SPS +PSPS S +P+ SP + + + A S + G G +G G
Sbjct: 639 PSESPSLAPSPSDSVASLAPSSSPSDESPSPAPSPSSSGSKGGGAGHG 782
>TC80366 similar to GP|13877617|gb|AAK43886.1 protein kinase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 814
Score = 70.9 bits (172), Expect = 2e-13
Identities = 50/136 (36%), Positives = 67/136 (48%), Gaps = 4/136 (2%)
Frame = +3
Query: 26 PAPTPATNSSLNSPSPTP-IPTPS-PANSPPAPTPTPTPSPHSDSPPAPSPDN--SPSSS 81
P TPAT SS PS TP P PS P+NSPP+P TP SP S PSP + PSS
Sbjct: 84 PPSTPATPSS-PPPSTTPSAPPPSTPSNSPPSPPTTPAISPPSGGGTTPSPPSRTPPSSD 260
Query: 82 PSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAAVGVVALG 141
SPSP SS P P +++I S+G GIAVG + + ++
Sbjct: 261 DSPSPPSSKTPPPPSPPSSSSI----------------STGTVIGIAVGAVVVLVFFSIC 392
Query: 142 AMVVKKRRQNIQRSEY 157
+ +K+++ + EY
Sbjct: 393 CICFRKKKRRRRDEEY 440
Score = 34.3 bits (77), Expect = 0.024
Identities = 22/66 (33%), Positives = 27/66 (40%), Gaps = 4/66 (6%)
Frame = +3
Query: 45 PTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSS----PAPSPDEAADN 100
P+ + PP P P P P +PP P P SS S S S P PSP A +
Sbjct: 540 PSDHVVSKPPPPAPIPPRPPSHVAPPPPPPAFISSSGGSGSNYSGGELLPPPSPGIAFSS 719
Query: 101 NAISHT 106
+ T
Sbjct: 720 GKSTFT 737
Score = 29.6 bits (65), Expect = 0.58
Identities = 24/86 (27%), Positives = 32/86 (36%), Gaps = 9/86 (10%)
Frame = +3
Query: 27 APTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSP---------PAPSPDNS 77
A P+ + P P PIP P++ P P P S S P PSP +
Sbjct: 531 AHPPSDHVVSKPPPPAPIPPRPPSHVAPPPPPPAFISSSGGSGSNYSGGELLPPPSPGIA 710
Query: 78 PSSSPSPSPSSSPAPSPDEAADNNAI 103
SS S A + D +D N +
Sbjct: 711 FSSGKSTFTYEELARATDGFSDANLL 788
Score = 26.2 bits (56), Expect = 6.4
Identities = 26/86 (30%), Positives = 31/86 (35%), Gaps = 11/86 (12%)
Frame = +3
Query: 54 PAPTPTPTPSPHSDSPPA------PSPDNSPSSSPSPS-----PSSSPAPSPDEAADNNA 102
P P P P+ P P D+ S P P+ P S AP P A
Sbjct: 465 PPPAQRPKVEPYGGPPQQWQNNAHPPSDHVVSKPPPPAPIPPRPPSHVAPPPPPPA---F 635
Query: 103 ISHTGIGEDGKSSGGGMSSGKKAGIA 128
IS +G G SGG + GIA
Sbjct: 636 ISSSG-GSGSNYSGGELLPPPSPGIA 710
>TC88762 similar to GP|22136928|gb|AAM91808.1 unknown protein {Arabidopsis
thaliana}, partial (3%)
Length = 805
Score = 65.9 bits (159), Expect = 7e-12
Identities = 43/120 (35%), Positives = 65/120 (53%), Gaps = 3/120 (2%)
Frame = +3
Query: 18 NIASSADSPAPTPATNSSLNSPSPTPIPTP-SPANSPPAPTPTPTPSPHSDSPPAPSPDN 76
N S + SP+ + +++ + P+ TP +P S SPPA TP +P P + SPPAP+P +
Sbjct: 81 NPTSPSSSPSASSPVSTTTSPPASTPASSPVSTTTSPPASTPASSPVPTTTSPPAPTPAS 260
Query: 77 SPSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKA--GIAVGVIAA 134
SP S+ SP+ +SPA S AA + S T S+ G S + G ++GV +A
Sbjct: 261 SPVSTNSPT--ASPAGSLPAAATPSPSSTTAGSPSPSSTPPGTRSSNETTPGRSIGVASA 434
Score = 53.1 bits (126), Expect = 5e-08
Identities = 28/70 (40%), Positives = 40/70 (57%)
Frame = +3
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSP 84
+P+P N + S SP+ S SPPA TP +P + SPPA ++P+SSP P
Sbjct: 57 APSPYQFNNPTSPSSSPSASSPVSTTTSPPASTPASSPVSTTTSPPA----STPASSPVP 224
Query: 85 SPSSSPAPSP 94
+ +S PAP+P
Sbjct: 225 TTTSPPAPTP 254
Score = 40.0 bits (92), Expect = 4e-04
Identities = 24/75 (32%), Positives = 39/75 (52%), Gaps = 2/75 (2%)
Frame = +3
Query: 19 IASSADSPAPTPATNS-SLNSPSPTPIPT-PSPANSPPAPTPTPTPSPHSDSPPAPSPDN 76
+ ++ PAPTPA++ S NSP+ +P + P+ A P+ T +PSP S P S +
Sbjct: 219 VPTTTSPPAPTPASSPVSTNSPTASPAGSLPAAATPSPSSTTAGSPSPSSTPPGTRSSNE 398
Query: 77 SPSSSPSPSPSSSPA 91
+ S+SP+
Sbjct: 399 TTPGRSIGVASASPS 443
Score = 37.4 bits (85), Expect = 0.003
Identities = 20/54 (37%), Positives = 29/54 (53%)
Frame = +3
Query: 37 NSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPSSSP 90
+S P TPS P+P P+ S SP A SP ++ +S P+ +P+SSP
Sbjct: 9 SSLPPGVAQTPSGNGIAPSPYQFNNPTSPSSSPSASSPVSTTTSPPASTPASSP 170
Score = 32.0 bits (71), Expect = 0.12
Identities = 22/56 (39%), Positives = 30/56 (53%), Gaps = 10/56 (17%)
Frame = +3
Query: 49 PANS-PPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS---------PSSSPAPSP 94
P++S PP TP+ + + SP + SPSSSPS S P+S+PA SP
Sbjct: 3 PSSSLPPGVAQTPSGNGIAPSPYQFNNPTSPSSSPSASSPVSTTTSPPASTPASSP 170
>TC78495 homologue to PIR|S53504|S53504 extensin-like protein S3 - alfalfa,
partial (97%)
Length = 1049
Score = 62.8 bits (151), Expect = 6e-11
Identities = 41/115 (35%), Positives = 61/115 (52%), Gaps = 3/115 (2%)
Frame = +3
Query: 19 IASSADSPAPTPATNSSLNSPSPTPIP-TPSPANSPPAPTPTPTPSPHSDSPPAPSPDNS 77
+ S+ +P PA+ + P TP P TP PA PPA TPTP SP + + PAP+P
Sbjct: 477 VQSTPPPASPPPASPPPFSPPPATPPPATPPPATPPPALTPTPLSSPPA-TTPAPAPAKL 653
Query: 78 PSSSPSPSPSSSP--APSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVG 130
S +P+ +P SP AP+P ++ + +IS +G + G + S K G+ G
Sbjct: 654 KSKAPALAPVLSPSDAPAPGLSSLSPSISPSGTDDSGAEK---LWSHKMVGLVFG 809
Score = 50.4 bits (119), Expect = 3e-07
Identities = 25/67 (37%), Positives = 35/67 (51%), Gaps = 10/67 (14%)
Frame = +3
Query: 38 SPSPTPI-PTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS---------PS 87
SP PTP+ P P + PPA P +P P S P P P P ++P P+ P+
Sbjct: 444 SPPPTPLTPPPVQSTPPPASPPPASPPPFSPPPATPPPATPPPATPPPALTPTPLSSPPA 623
Query: 88 SSPAPSP 94
++PAP+P
Sbjct: 624 TTPAPAP 644
Score = 48.9 bits (115), Expect = 9e-07
Identities = 33/90 (36%), Positives = 43/90 (47%), Gaps = 8/90 (8%)
Frame = +3
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPP-----APTPTP-TPSPHSDSPPAPSPDNSP 78
SP P ++ L S P TP P S P +P PTP TP P +PP SP P
Sbjct: 339 SPPPVQSSPPPLQSSPPPAQSTPPPVQSSPPPVQQSPPPTPLTPPPVQSTPPPASP--PP 512
Query: 79 SSSP--SPSPSSSPAPSPDEAADNNAISHT 106
+S P SP P++ P +P A A++ T
Sbjct: 513 ASPPPFSPPPATPPPATPPPATPPPALTPT 602
Score = 47.0 bits (110), Expect = 4e-06
Identities = 27/74 (36%), Positives = 36/74 (48%)
Frame = +3
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
S++ + +P P T + N+P TP +P P S P P + P S PPA S S
Sbjct: 246 STSPNSSPAPPTPPA-NTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQSTPPPVQS 422
Query: 81 SPSPSPSSSPAPSP 94
SP P SP P+P
Sbjct: 423 SP-PPVQQSPPPTP 461
Score = 46.2 bits (108), Expect = 6e-06
Identities = 33/90 (36%), Positives = 41/90 (44%), Gaps = 16/90 (17%)
Frame = +3
Query: 21 SSADSPAP-TPATNSSLNSPSPTPIP---TPSPANSPPAP---TPTP---TPSPHSDSPP 70
S SPAP TP N+ +P +P P +P P S P P +P P TP P SPP
Sbjct: 252 SPNSSPAPPTPPANTPPTTPQASPPPVQSSPPPVQSSPPPLQSSPPPAQSTPPPVQSSPP 431
Query: 71 A------PSPDNSPSSSPSPSPSSSPAPSP 94
P+P P +P P+S P SP
Sbjct: 432 PVQQSPPPTPLTPPPVQSTPPPASPPPASP 521
Score = 46.2 bits (108), Expect = 6e-06
Identities = 27/84 (32%), Positives = 39/84 (46%)
Frame = +3
Query: 9 VFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDS 68
+ L++ L+ + S+ P+T SP+ +P P PAN+PP TP +P P S
Sbjct: 180 IVLVVIGLICVVFSSVGAQQAPST-----SPNSSPAPPTPPANTPPT-TPQASPPPVQSS 341
Query: 69 PPAPSPDNSPSSSPSPSPSSSPAP 92
PP P S P S+P P
Sbjct: 342 PPPVQSSPPPLQSSPPPAQSTPPP 413
>AW127239 weakly similar to SP|Q9FPQ6|GP1_C Vegetative cell wall protein gp1
precursor (Hydroxyproline-rich glycoprotein 1)., partial
(5%)
Length = 362
Score = 62.0 bits (149), Expect = 1e-10
Identities = 39/103 (37%), Positives = 56/103 (53%)
Frame = +2
Query: 8 LVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSD 67
L F+LL+ +++ +ADSP SPSP P +PSP ++ P+P P SP
Sbjct: 113 LSFILLTLSLSLHVTADSPP----------SPSPAPSLSPSPTDT-PSPYYPPASSPPVS 259
Query: 68 SPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNAISHTGIGE 110
SPPAPSP N PSP +P PSP+ D+ +++H + E
Sbjct: 260 SPPAPSPLN-------PSPIPAPVPSPE---DSTSLNHIDVDE 358
>TC88911 weakly similar to GP|13926280|gb|AAK49610.1 AT5g14920/F2G14_40
{Arabidopsis thaliana}, partial (11%)
Length = 1320
Score = 60.8 bits (146), Expect = 2e-10
Identities = 42/115 (36%), Positives = 58/115 (49%), Gaps = 25/115 (21%)
Frame = +3
Query: 9 VFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIP-TPSPANSPPAPTPT-----PTP 62
+FL +L ++ S P P PA+ SP+P +P P+PA PAP PT P P
Sbjct: 486 LFLYHHYLYHLVSLPPLPLP-PASKPPKVSPAPAKVPKAPAPAKEAPAPAPTHKKKAPAP 662
Query: 63 SPHSDSP-PAPS--PDNSPSSSPSPS----------------PSSSPAPSPDEAA 98
+P ++P PAP+ +P SSP PS PSS+PAPSP++ A
Sbjct: 663 APDKETPAPAPTHKKKKAPKSSPVPSPAILPPSPAPTPAIDTPSSAPAPSPEDDA 827
Score = 50.1 bits (118), Expect = 4e-07
Identities = 33/83 (39%), Positives = 39/83 (46%), Gaps = 18/83 (21%)
Frame = +3
Query: 20 ASSADSPAPT-----PATNSSLNSPSPTPI----------PTPSPANSPPAPTPTP---T 61
A A +PAPT PA +P+P P P PSPA PP+P PTP T
Sbjct: 609 AKEAPAPAPTHKKKAPAPAPDKETPAPAPTHKKKKAPKSSPVPSPAILPPSPAPTPAIDT 788
Query: 62 PSPHSDSPPAPSPDNSPSSSPSP 84
PS S PAPSP++ P P
Sbjct: 789 PS----SAPAPSPEDDAPEPPPP 845
Score = 47.4 bits (111), Expect = 3e-06
Identities = 27/82 (32%), Positives = 40/82 (47%), Gaps = 4/82 (4%)
Frame = +2
Query: 21 SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPP--APTPTPTPSPHSDSP-PAPSPDNS 77
++ + AP +P P+P+ P P + PP AP +P P P +P P P +
Sbjct: 239 TTTPTSAPLKPPVPKAPAPKPSPVKPPVPKSKPPTTAPVKSPVPDPPKLAPVKLPVPKVT 418
Query: 78 PSSSP-SPSPSSSPAPSPDEAA 98
P+ SP +PSP P P P + A
Sbjct: 419 PALSPKTPSPKIQPPPHPPKKA 484
Score = 47.0 bits (110), Expect = 4e-06
Identities = 36/114 (31%), Positives = 52/114 (45%), Gaps = 21/114 (18%)
Frame = +3
Query: 25 SPAPTPATNSSLNSPSPTP---IPTPSPANSPP---------------APTPTPTPSPHS 66
SPAPTPA ++ ++P+P+P P P P + A P PT S +
Sbjct: 759 SPAPTPAIDTPSSAPAPSPEDDAPEPPPPHKHKRRKHKHSKHKKHHALALAPEPTSSSST 938
Query: 67 ---DSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGG 117
SPPAP D++ + S PS +P+PS N A S+ G ++GG
Sbjct: 939 IIRRSPPAPLADDNTTMSSDEGPSPAPSPSA-----NGAQSYQGQWRKMLATGG 1085
Score = 41.2 bits (95), Expect = 2e-04
Identities = 25/78 (32%), Positives = 30/78 (38%), Gaps = 8/78 (10%)
Frame = +2
Query: 23 ADSPAPTPATNSSLNSPSPTPIPTPSPANSP--------PAPTPTPTPSPHSDSPPAPSP 74
A +P P+P S PT P SP P P P TP SP + SP P
Sbjct: 284 APAPKPSPVKPPVPKSKPPTTAPVKSPVPDPPKLAPVKLPVPKVTPALSPKTPSPKIQPP 463
Query: 75 DNSPSSSPSPSPSSSPAP 92
+ P +P P S P
Sbjct: 464 PHPPKKAPVSLPPLSIPP 517
Score = 39.7 bits (91), Expect = 6e-04
Identities = 25/78 (32%), Positives = 31/78 (39%), Gaps = 8/78 (10%)
Frame = +2
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDS-PPAPSPDNSPSSSPS 83
+P P + +PT P P PAP P+P P S PP +P SP P
Sbjct: 200 TPTAAPVKPIVPKTTTPTSAPLKPPVPKAPAPKPSPVKPPVPKSKPPTTAPVKSPVPDPP 379
Query: 84 -------PSPSSSPAPSP 94
P P +PA SP
Sbjct: 380 KLAPVKLPVPKVTPALSP 433
Score = 39.7 bits (91), Expect = 6e-04
Identities = 23/66 (34%), Positives = 37/66 (55%), Gaps = 3/66 (4%)
Frame = +3
Query: 43 PIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPSPS---SSPAPSPDEAAD 99
P+P P PA+ PP +P P P + PAP+ + +P+P+P+ +PAP+PD+
Sbjct: 531 PLPLP-PASKPPKVSPAPAKVPKA---PAPAKE-----APAPAPTHKKKAPAPAPDKETP 683
Query: 100 NNAISH 105
A +H
Sbjct: 684 APAPTH 701
Score = 38.9 bits (89), Expect = 0.001
Identities = 22/75 (29%), Positives = 32/75 (42%), Gaps = 3/75 (4%)
Frame = +2
Query: 27 APTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSP---SSSPS 83
AP + +P+ P+ P + P P P P + +P PSP P S P+
Sbjct: 167 APAKPLVPKVTTPTAAPVKPIVPKTTTPTSAPLKPPVPKAPAPK-PSPVKPPVPKSKPPT 343
Query: 84 PSPSSSPAPSPDEAA 98
+P SP P P + A
Sbjct: 344 TAPVKSPVPDPPKLA 388
Score = 34.7 bits (78), Expect = 0.018
Identities = 24/74 (32%), Positives = 32/74 (42%), Gaps = 4/74 (5%)
Frame = +2
Query: 25 SPAPTPATNSSLNSPSPTP---IPTPSPANSPPAPTPTPTPSPHSDS-PPAPSPDNSPSS 80
SP P T +S P P PT +PA TPT +P P +P ++P
Sbjct: 89 SPVPKATTPTSAPVKPPVPKVATPTVAPAKPLVPKVTTPTAAPVKPIVPKTTTPTSAPLK 268
Query: 81 SPSPSPSSSPAPSP 94
P P + +P PSP
Sbjct: 269 PPVPK-APAPKPSP 307
Score = 34.7 bits (78), Expect = 0.018
Identities = 19/71 (26%), Positives = 29/71 (40%), Gaps = 3/71 (4%)
Frame = +2
Query: 27 APTPATNSSLNSPSPTPIP---TPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPS 83
+P P + ++P P+P TP+ A + P TP+ P P S+
Sbjct: 89 SPVPKATTPTSAPVKPPVPKVATPTVAPAKPLVPKVTTPTAAPVKPIVPKTTTPTSAPLK 268
Query: 84 PSPSSSPAPSP 94
P +PAP P
Sbjct: 269 PPVPKAPAPKP 301
Score = 29.3 bits (64), Expect = 0.76
Identities = 22/67 (32%), Positives = 29/67 (42%), Gaps = 2/67 (2%)
Frame = +2
Query: 25 SPAPTPATNSSLNSPSP--TPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSP 82
SP P P + + P P TP SP P+P P PH PP +P + P S
Sbjct: 359 SPVPDPPKLAPVKLPVPKVTPA------LSPKTPSPKIQPPPH---PPKKAPVSLPPLSI 511
Query: 83 SPSPSSS 89
P +S+
Sbjct: 512 PPGFTST 532
>TC85577 homologue to GP|6715512|gb|AAF26445.1| vacuolar H+-ATPase B subunit
{Nicotiana tabacum}, partial (14%)
Length = 841
Score = 60.5 bits (145), Expect = 3e-10
Identities = 39/108 (36%), Positives = 61/108 (56%), Gaps = 6/108 (5%)
Frame = -1
Query: 6 FSLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPS---PANSPPAPT---PT 59
+++V +L++ L+ ++ A SP+ +P + P + P+PS P SPP P P+
Sbjct: 538 YTVVLMLVATLLVSSTIAQSPSSSPTISPVATPPKSSQAPSPSAVSPVASPPVPVKNAPS 359
Query: 60 PTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAADNNAISHTG 107
P+P SDSP A SP +PSS S +PS +P+PS + AA N + G
Sbjct: 358 PSPPASSDSP-AVSPAVTPSSI-STTPSEAPSPSDNSAAAFNRFTVAG 221
Score = 47.8 bits (112), Expect = 2e-06
Identities = 35/107 (32%), Positives = 52/107 (47%), Gaps = 2/107 (1%)
Frame = -1
Query: 34 SSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSD--SPPAPSPDNSPSSSPSPSPSSSPA 91
SS + SP+ PT SP +PP + P+PS S SPP P N+PS SP P+ S SPA
Sbjct: 499 SSTIAQSPSSSPTISPVATPPKSSQAPSPSAVSPVASPPVPVK-NAPSPSP-PASSDSPA 326
Query: 92 PSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAAVGVV 138
SP + + + + +S + AG A ++ A ++
Sbjct: 325 VSPAVTPSSISTTPSEAPSPSDNSAAAFNRFTVAGSAAVMVFAAALM 185
>TC77143 similar to GP|559557|gb|AAA51649.1|| arabinogalactan-protein {Pyrus
communis}, partial (30%)
Length = 956
Score = 60.1 bits (144), Expect = 4e-10
Identities = 38/104 (36%), Positives = 50/104 (47%), Gaps = 18/104 (17%)
Frame = +3
Query: 9 VFLLLSFLVNIA-SSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHS- 66
VFL+L L + A APT +++ P+P P+ TP A P A PT TP P +
Sbjct: 132 VFLILGLLATSCFAQAPGAAPTQPPSATPTPPTPAPVATPPTATPPTATPPTATPPPAAA 311
Query: 67 ---------DSPPAPSP-------DNSPSSSPSPSPSSSPAPSP 94
SPPAP+P D++ SS P+P P PAP P
Sbjct: 312 PTPATPAPATSPPAPTPTSDAPTPDSTSSSPPAPGP-GGPAPGP 440
Score = 48.9 bits (115), Expect = 9e-07
Identities = 29/95 (30%), Positives = 49/95 (51%), Gaps = 10/95 (10%)
Frame = +3
Query: 34 SSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNS---PSSSPSPS----P 86
+S + +P PT P+ +P PTP P +P + +PP +P + P+++P+P+
Sbjct: 159 TSCFAQAPGAAPTQPPSATPTPPTPAPVATPPTATPPTATPPTATPPPAAAPTPATPAPA 338
Query: 87 SSSPAPSPDEAA---DNNAISHTGIGEDGKSSGGG 118
+S PAP+P A D+ + S G G + G G
Sbjct: 339 TSPPAPTPTSDAPTPDSTSSSPPAPGPGGPAPGPG 443
Score = 34.7 bits (78), Expect = 0.018
Identities = 23/68 (33%), Positives = 32/68 (46%), Gaps = 2/68 (2%)
Frame = -2
Query: 80 SSPSPSPSSSPAPSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGI--AVGVIAAVGV 137
+S P P + P P P + S G E G +GG ++ AG+ A G AVG
Sbjct: 454 ASVLPGPGAGP-PGPGAGGEEEVESGVGASEVGVGAGGEVAGAGVAGVGAAAGGGVAVGG 278
Query: 138 VALGAMVV 145
VA+G + V
Sbjct: 277 VAVGGVAV 254
>TC77282 homologue to GP|17065300|gb|AAL32804.1 Unknown protein {Arabidopsis
thaliana}, partial (19%)
Length = 787
Score = 59.7 bits (143), Expect = 5e-10
Identities = 40/104 (38%), Positives = 60/104 (57%), Gaps = 6/104 (5%)
Frame = +2
Query: 9 VFLLLSFLV-NIASSADSPAPTPATNSSLNSPSP-TPIPTPSPANSP-PAPTPTPTPSPH 65
+F++L L + + A +PAPT + +S +PSP T PTPSPA SP + +P +PSP+
Sbjct: 149 LFVILGLLATSCLAQAPAPAPTQLSPNSAPAPSPSTKSPTPSPAVSPSTSSSPGSSPSPN 328
Query: 66 SDSPPAPSPDNSPSSSP---SPSPSSSPAPSPDEAADNNAISHT 106
+ SPPAP N P+S P +P + APS + N ++ T
Sbjct: 329 TFSPPAPG-SNVPASGPGGATPGQPGADAPSAAFSITNTFVAVT 457
Score = 49.3 bits (116), Expect = 7e-07
Identities = 34/107 (31%), Positives = 47/107 (43%), Gaps = 3/107 (2%)
Frame = +2
Query: 34 SSLNSPSPTPIPTPSPANSPPAPTPT---PTPSPHSDSPPAPSPDNSPSSSPSPSPSSSP 90
+S + +P P PT NS PAP+P+ PTPSP A SP S S SPSP++
Sbjct: 176 TSCLAQAPAPAPTQLSPNSAPAPSPSTKSPTPSP------AVSPSTSSSPGSSPSPNTFS 337
Query: 91 APSPDEAADNNAISHTGIGEDGKSSGGGMSSGKKAGIAVGVIAAVGV 137
P+P + G+ G + S +AV A + V
Sbjct: 338 PPAPGSNVPASGPGGATPGQPGADAPSAAFSITNTFVAVTAFAGIFV 478
>TC88788 weakly similar to SP|P05790|FBOH_BOMMO Fibroin heavy chain
precursor (Fib-H) (H-fibroin). [Silk moth] {Bombyx
mori}, partial (2%)
Length = 782
Score = 59.7 bits (143), Expect = 5e-10
Identities = 28/72 (38%), Positives = 35/72 (47%)
Frame = -2
Query: 23 ADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSP 82
A PAP PA P P P PA+ P P P P P ++ P P PD +P+ P
Sbjct: 382 ASEPAPDPA-------PDPARDPAWEPASEPAPEDPDPDPDPEVEAEPDPEPDPAPAPEP 224
Query: 83 SPSPSSSPAPSP 94
+P P SP+P P
Sbjct: 223 APPPPPSPSPKP 188
Score = 52.0 bits (123), Expect = 1e-07
Identities = 30/81 (37%), Positives = 36/81 (44%), Gaps = 6/81 (7%)
Frame = -2
Query: 23 ADSPAPTPATNSSLNSPSPTPIPTPSPANSP---PAPTPTPTPSPHSDSPPAPS---PDN 76
A PA PA+ + P P P PA+ P PAP P P+ S PAP PD
Sbjct: 463 AYEPASEPASEPAPEDPDPA*EPAYEPASEPAPDPAPDPARDPAWEPASEPAPEDPDPDP 284
Query: 77 SPSSSPSPSPSSSPAPSPDEA 97
P P P PAP+P+ A
Sbjct: 283 DPEVEAEPDPEPDPAPAPEPA 221
Score = 51.2 bits (121), Expect = 2e-07
Identities = 32/88 (36%), Positives = 43/88 (48%), Gaps = 5/88 (5%)
Frame = -2
Query: 23 ADSPAPTPATNSS---LNSPSPT-PIPTPSP-ANSPPAPTPTPTPSPHSDSPPAPSPDNS 77
A PAP PA + + + P+P P P P P + P P P P P+P PAP P S
Sbjct: 370 APDPAPDPARDPAWEPASEPAPEDPDPDPDPEVEAEPDPEPDPAPAPE----PAPPPPPS 203
Query: 78 PSSSPSPSPSSSPAPSPDEAADNNAISH 105
PS P+P+P +P S + +SH
Sbjct: 202 PSPKPAPNPIPTPPKSNPRSFLAALLSH 119
Score = 48.5 bits (114), Expect = 1e-06
Identities = 29/81 (35%), Positives = 37/81 (44%), Gaps = 7/81 (8%)
Frame = -2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPA------PTPTPTPSPHSDSPPAPSPDNSPS 79
PA PA+ +L P P P PA+ P + P P P+ S PAP P P+
Sbjct: 523 PAYEPASEPALEDPDPA*DPAYEPASEPASEPAPEDPDPA*EPAYEPASEPAPDPAPDPA 344
Query: 80 SSPSPSPSSSPAP-SPDEAAD 99
P+ P+S PAP PD D
Sbjct: 343 RDPAWEPASEPAPEDPDPDPD 281
Score = 38.5 bits (88), Expect = 0.001
Identities = 25/68 (36%), Positives = 32/68 (46%), Gaps = 7/68 (10%)
Frame = -2
Query: 39 PSPTPIPTPSPANSP----PAPTPTPTPSPHSDSPPAPSP-DNSPSSSPSPSPSSSPA-- 91
P P P PA+ P P P P P S+ P+P D P+ P+ P+S PA
Sbjct: 541 PDPA*DPAYEPASEPALEDPDPA*DPAYEPASEPASEPAPEDPDPA*EPAYEPASEPAPD 362
Query: 92 PSPDEAAD 99
P+PD A D
Sbjct: 361 PAPDPARD 338
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 58.9 bits (141), Expect = 9e-10
Identities = 32/79 (40%), Positives = 37/79 (46%), Gaps = 8/79 (10%)
Frame = +2
Query: 24 DSPAPTPATNSSLNSPSPTPI----PTPSPANSPPAPTPTPTPSPHSDSPPA----PSPD 75
+SP P P + P P P P P PA+SPP P +P P PHS +PP P
Sbjct: 47 NSPPPPPPPAPVFSPPPPVPYYYSSPPPPPAHSPPPPPXSPPPPPHSPTPPVYPYLSPPP 226
Query: 76 NSPSSSPSPSPSSSPAPSP 94
P SP P S P PSP
Sbjct: 227 PPPVHSPPPPVYSPPPPSP 283
Score = 54.3 bits (129), Expect = 2e-08
Identities = 31/76 (40%), Positives = 38/76 (49%), Gaps = 8/76 (10%)
Frame = +2
Query: 27 APTPATNSSLNSPSPTPIPTPSP-ANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
+PTP L+ P P P+ +P P SPP P+P P P PP P + P SSP+P
Sbjct: 185 SPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEP-PPPPPPPCVEPPPPSSPAPH 361
Query: 86 -------PSSSPAPSP 94
PS SP PSP
Sbjct: 362 QTPYHPPPSPSPPPSP 409
Score = 50.4 bits (119), Expect = 3e-07
Identities = 31/85 (36%), Positives = 34/85 (39%), Gaps = 2/85 (2%)
Frame = +2
Query: 23 ADSPAPTPATNSSLNSPSPTPIPTPS--PANSPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
A SP P P + P P PTP P SPP P P +P P SPP PSP
Sbjct: 137 AHSPPPPPXS-----PPPPPHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEP 301
Query: 81 SPSPSPSSSPAPSPDEAADNNAISH 105
P P P P P A + H
Sbjct: 302 PPPPPPPCVEPPPPSSPAPHQTPYH 376
Score = 48.1 bits (113), Expect = 2e-06
Identities = 29/75 (38%), Positives = 35/75 (46%), Gaps = 6/75 (8%)
Frame = +2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAP---SPDNSP---S 79
P P P P +P P +P + PP+P+P P+P SPP P SP SP
Sbjct: 299 PPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPPVYTSPPPSPVYAY 478
Query: 80 SSPSPSPSSSPAPSP 94
SP P SSP P P
Sbjct: 479 PSPPPPVYSSPPPPP 523
Score = 47.4 bits (111), Expect = 3e-06
Identities = 29/70 (41%), Positives = 31/70 (43%)
Frame = +2
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSP 84
SP P P SP P P P+P SPP P P SP PPA SP P S P P
Sbjct: 26 SPPPPPPN-------SPPPPPPPAPVFSPPPPVPYYYSSP--PPPPAHSPPPPPXSPPPP 178
Query: 85 SPSSSPAPSP 94
S +P P
Sbjct: 179PHSPTPPVYP 208
Score = 46.2 bits (108), Expect = 6e-06
Identities = 32/85 (37%), Positives = 38/85 (44%), Gaps = 15/85 (17%)
Frame = +2
Query: 25 SPAPTPATNS------SLNSPSPTPIPTPSP------ANSPPAPTPTPTPSPHSDSPPAP 72
SP P P +S S PSP P P P PP +P P +P+ PP+P
Sbjct: 215 SPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPPPPPCVEPPPPSSPAPHQTPY-HPPPSP 391
Query: 73 SPDNSP---SSSPSPSPSSSPAPSP 94
SP SP SP P +SP PSP
Sbjct: 392 SPPPSPVYAYPSPPPPVYTSPPPSP 466
Score = 45.1 bits (105), Expect = 1e-05
Identities = 27/74 (36%), Positives = 31/74 (41%), Gaps = 9/74 (12%)
Frame = +2
Query: 28 PTPATNSSLNSPSPTPIPTPSPA----NSPPAPTPTPTPSPHSDSPPAPSPDNSPS---- 79
P P NS P P P+ +P P S P P P +P P SPP P +P
Sbjct: 32 PPPPPNSPPPPPPPAPVFSPPPPVPYYYSSPPPPPAHSPPPPPXSPPPPPHSPTPPVYPY 211
Query: 80 -SSPSPSPSSSPAP 92
S P P P SP P
Sbjct: 212 LSPPPPPPVHSPPP 253
Score = 44.7 bits (104), Expect = 2e-05
Identities = 27/65 (41%), Positives = 29/65 (44%), Gaps = 13/65 (20%)
Frame = +2
Query: 41 PTP-----IPTPSPANSPPAPTPT------PTPSPHSDSPPAPSPDNSPSSSPS--PSPS 87
PTP P P P NSPP P P P P P+ S P P P +SP P P P
Sbjct: 2 PTPPYCVRSPPPPPPNSPPPPPPPAPVFSPPPPVPYYYSSPPPPPAHSPPPPPXSPPPPP 181
Query: 88 SSPAP 92
SP P
Sbjct: 182HSPTP 196
Score = 41.6 bits (96), Expect = 1e-04
Identities = 27/73 (36%), Positives = 34/73 (45%), Gaps = 6/73 (8%)
Frame = +2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTP----TPTPSP--HSDSPPAPSPDNSPS 79
P +PA + + P P+P P PSP + P+P P +P PSP SPP P S
Sbjct: 338 PPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPPVYTSPPPSPVYAYPSPPPPV----YS 505
Query: 80 SSPSPSPSSSPAP 92
S P P P P
Sbjct: 506 SPPPPPVYEGPIP 544
Score = 40.8 bits (94), Expect = 3e-04
Identities = 29/75 (38%), Positives = 34/75 (44%), Gaps = 6/75 (8%)
Frame = +2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSP-PAPTPT-PTPSPHSDSPPA---PSPDNSPSS 80
P P+P P P P P P +SP P TP P PSP P PSP +
Sbjct: 269 PPPSPPPCVEPPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPPVYT 448
Query: 81 SPSPSP-SSSPAPSP 94
SP PSP + P+P P
Sbjct: 449 SPPPSPVYAYPSPPP 493
Score = 36.2 bits (82), Expect = 0.006
Identities = 23/67 (34%), Positives = 31/67 (45%), Gaps = 8/67 (11%)
Frame = +2
Query: 26 PAPTPATNSSLNSPSPTP----IPTPSPANSPPAPTP----TPTPSPHSDSPPAPSPDNS 77
P+P+P + PSP P P PSP + P+P P +P P P + P P S
Sbjct: 383 PSPSPPPSPVYAYPSPPPPVYTSPPPSPVYAYPSPPPPVYSSPPPPPVYEGPIPPVFGIS 562
Query: 78 PSSSPSP 84
+S P P
Sbjct: 563 YASPPPP 583
>TC76445 similar to PIR|T07623|T07623 extensin homolog HRGP2 - soybean
(fragment), partial (76%)
Length = 821
Score = 58.9 bits (141), Expect = 9e-10
Identities = 31/76 (40%), Positives = 39/76 (50%), Gaps = 3/76 (3%)
Frame = +2
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
S P+P+P SP P P P+P P SPP P+P+P P H SPP PSP + P
Sbjct: 80 SPPPPSPSPPPPYYYKSPPP-PSPSPPPPYYYKSPPPPSPSPPPPYHYQSPPPPSPISHP 256
Query: 79 SSSPSPSPSSSPAPSP 94
+ P SP+P P
Sbjct: 257 PNYYKSPPPPSPSPPP 304
Score = 55.8 bits (133), Expect = 8e-09
Identities = 32/76 (42%), Positives = 39/76 (51%), Gaps = 7/76 (9%)
Frame = +2
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
S P+P+P + SP P P P+P P SPP P+P+P P + SPP PSP P
Sbjct: 32 SPPPPSPSPPSPYYYKSPPP-PSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPP 208
Query: 79 ----SSSPSPSPSSSP 90
S P PSP S P
Sbjct: 209 PYHYQSPPPPSPISHP 256
Score = 55.1 bits (131), Expect = 1e-08
Identities = 34/79 (43%), Positives = 39/79 (49%), Gaps = 12/79 (15%)
Frame = +2
Query: 28 PTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSP----SSSPS 83
P P S PSP+P P+P SPP P+P+P P + SPP PSP P S P
Sbjct: 11 PPPYYYKSPPPPSPSP-PSPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPP 187
Query: 84 PSPS--------SSPAPSP 94
PSPS S P PSP
Sbjct: 188PSPSPPPPYHYQSPPPPSP 244
Score = 48.9 bits (115), Expect = 9e-07
Identities = 30/78 (38%), Positives = 36/78 (45%), Gaps = 6/78 (7%)
Frame = +2
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
S P+P+P SP P P P+P P SPP P+P P + SPP PSP P
Sbjct: 128 SPPPPSPSPPPPYYYKSPPP-PSPSPPPPYHYQSPPPPSPISHPPNYYKSPPPPSPSPPP 304
Query: 79 S---SSPSPSPSSSPAPS 93
SP P S P P+
Sbjct: 305 PYHYVSPPPPVKSPPPPA 358
Score = 43.5 bits (101), Expect = 4e-05
Identities = 26/68 (38%), Positives = 31/68 (45%), Gaps = 3/68 (4%)
Frame = +2
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPAN---SPPAPTPTPTPSPHSDSPPAPSPDNSP 78
S P+P+P SP P P P P N SPP P+P+P P H SPP P P
Sbjct: 176 SPPPPSPSPPPPYHYQSPPP-PSPISHPPNYYKSPPPPSPSPPPPYHYVSPPPPVKSPPP 352
Query: 79 SSSPSPSP 86
+ SP
Sbjct: 353 PAYIYASP 376
Score = 41.2 bits (95), Expect = 2e-04
Identities = 27/59 (45%), Positives = 31/59 (51%), Gaps = 3/59 (5%)
Frame = +2
Query: 39 PSPTPIPTPSPANSPPAPTPTPTPSPH---SDSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
PSP P P SPP P+P+P PSP+ S PP+PSP P SP P S P P
Sbjct: 2 PSPPP---PYYYKSPPPPSPSP-PSPYYYKSPPPPSPSPP-PPYYYKSPPPPSPSPPPP 163
Score = 36.2 bits (82), Expect = 0.006
Identities = 22/64 (34%), Positives = 31/64 (48%), Gaps = 5/64 (7%)
Frame = +2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPAN-----SPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
P P+P ++ SP P P+PSP SPP P +P P + + P P N P +
Sbjct: 230 PPPSPISHPPNYYKSPPP-PSPSPPPPYHYVSPPPPVKSPPPPAYIYASPPPPIYN*PHN 406
Query: 81 SPSP 84
+ SP
Sbjct: 407 TKSP 418
>TC91646 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (15%)
Length = 984
Score = 58.2 bits (139), Expect = 2e-09
Identities = 34/93 (36%), Positives = 42/93 (44%), Gaps = 21/93 (22%)
Frame = +1
Query: 23 ADSPAPTPATNSSLNSPSPTPIPTPS------------PANSPPAPTPT---------PT 61
A SP P+P+ S L PS P+ TP P SPP P P P
Sbjct: 247 APSPRPSPSPLSPLRFPSRLPLTTPPSLTTSPSPTASPPPRSPPRPPPRSPPRQLMSPPR 426
Query: 62 PSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSP 94
P P S SPP +P ++P+P P SP+PSP
Sbjct: 427 PLPRSPSPPPRNPPELSRTTPTPRPPMSPSPSP 525
Score = 48.9 bits (115), Expect = 9e-07
Identities = 33/96 (34%), Positives = 39/96 (40%), Gaps = 22/96 (22%)
Frame = +1
Query: 25 SPAPTPATNSSLNSPSPTP--------IPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDN 76
SP+P P L+ +PTP PT PA +PP P P P+ PP P P
Sbjct: 439 SPSPPPRNPPELSRTTPTPRPPMSPSPSPTSPPAATPPVSRPPPRPALRRLLPPPPLPPT 618
Query: 77 SPSS--------------SPSPSPSSSPAPSPDEAA 98
PSS SP + SS P P P AA
Sbjct: 619 LPSSMASLFLAASVLPPASPPTT*SSPPTP*PSSAA 726
Score = 48.1 bits (113), Expect = 2e-06
Identities = 30/77 (38%), Positives = 40/77 (50%), Gaps = 10/77 (12%)
Frame = +1
Query: 28 PTPATNSSLNSPSPTPI-----PTPSPANSPPAPT--PTPTPSPHSDSPPAPSPDNSP-- 78
P+P + PSP+P+ P+ P +PP+ T P+PT SP SPP P P + P
Sbjct: 229 PSPLMEAPSPRPSPSPLSPLRFPSRLPLTTPPSLTTSPSPTASPPPRSPPRPPPRSPPRQ 408
Query: 79 -SSSPSPSPSSSPAPSP 94
S P P P SP+P P
Sbjct: 409 LMSPPRPLP-RSPSPPP 456
Score = 47.4 bits (111), Expect = 3e-06
Identities = 27/74 (36%), Positives = 35/74 (46%), Gaps = 1/74 (1%)
Frame = +1
Query: 22 SADSPAPTPATNSSLNSPSPTP-IPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
S P P ++ P P P P+P P N P TPTP P P+PSP + P++
Sbjct: 370 SPPRPPPRSPPRQLMSPPRPLPRSPSPPPRNPPELSRTTPTPRPPMS--PSPSPTSPPAA 543
Query: 81 SPSPSPSSSPAPSP 94
+P P S P P P
Sbjct: 544 TP---PVSRPPPRP 576
Score = 31.2 bits (69), Expect = 0.20
Identities = 28/93 (30%), Positives = 41/93 (43%), Gaps = 2/93 (2%)
Frame = +1
Query: 4 PRFSLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPS 63
P + + L L + S + +P + +S PS T S + P+ P
Sbjct: 712 PSSAALLLCLPASATLPCSEQTASPVTSWTTSTRCPSARTSATLSAPVTLPSAAVVVLPL 891
Query: 64 PHSDSPPAPSPDNSPSSSPSPSPS--SSPAPSP 94
S S +P P SPS+S S +PS +SP PSP
Sbjct: 892 LSSVSS-SPRPSASPSTSASTAPSVPTSP-PSP 984
Score = 30.8 bits (68), Expect = 0.26
Identities = 20/58 (34%), Positives = 29/58 (49%), Gaps = 4/58 (6%)
Frame = +3
Query: 21 SSADSPAPT----PATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSP 74
+S + P T P T +S S S T +PT S + S P+ T T T + + AP+P
Sbjct: 447 TSTEEPTGTVTDDPNTTASDESFSFTDVPTGSDSTSVPSATTTGTETATPSATVAPNP 620
>TC76326 homologue to PIR|C29356|C29356 hydroxyproline-rich glycoprotein
(clone Hyp2.13) - kidney bean (fragment), partial (32%)
Length = 777
Score = 58.2 bits (139), Expect = 2e-09
Identities = 33/76 (43%), Positives = 39/76 (50%), Gaps = 7/76 (9%)
Frame = +1
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
S P+P+P SP P P P+P P SPP P+P+P P H SPP PSP + P
Sbjct: 55 SPPPPSPSPPPPYYYKSPPP-PSPSPPPPYYYKSPPPPSPSPPPPYHYQSPPPPSPISHP 231
Query: 79 ----SSSPSPSPSSSP 90
S P PSPS P
Sbjct: 232 PYYYKSPPPPSPSPPP 279
Score = 54.7 bits (130), Expect = 2e-08
Identities = 31/72 (43%), Positives = 37/72 (51%), Gaps = 7/72 (9%)
Frame = +1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAPSPDNSP---- 78
P+P+P SP P P P+P P SPP P+P+P P + SPP PSP P
Sbjct: 19 PSPSPPPPYYYKSPPP-PSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYHY 195
Query: 79 SSSPSPSPSSSP 90
S P PSP S P
Sbjct: 196QSPPPPSPISHP 231
Score = 53.1 bits (126), Expect = 5e-08
Identities = 34/82 (41%), Positives = 39/82 (47%), Gaps = 12/82 (14%)
Frame = +1
Query: 25 SPAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSP----SS 80
+P P P PSP+P P P SPP P+P+P P + SPP PSP P S
Sbjct: 1 APGPPP--------PSPSP-PPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKS 153
Query: 81 SPSPSPS--------SSPAPSP 94
P PSPS S P PSP
Sbjct: 154PPPPSPSPPPPYHYQSPPPPSP 219
Score = 48.5 bits (114), Expect = 1e-06
Identities = 30/78 (38%), Positives = 36/78 (45%), Gaps = 6/78 (7%)
Frame = +1
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
S P+P+P SP P P P+P P SPP P+P P + SPP PSP P
Sbjct: 103 SPPPPSPSPPPPYYYKSPPP-PSPSPPPPYHYQSPPPPSPISHPPYYYKSPPPPSPSPPP 279
Query: 79 S---SSPSPSPSSSPAPS 93
SP P S P P+
Sbjct: 280 PYHYVSPPPPVKSPPPPT 333
Score = 41.2 bits (95), Expect = 2e-04
Identities = 25/68 (36%), Positives = 30/68 (43%), Gaps = 3/68 (4%)
Frame = +1
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
S P+P+P SP P P P P SPP P+P+P P H SPP P P
Sbjct: 151 SPPPPSPSPPPPYHYQSPPP-PSPISHPPYYYKSPPPPSPSPPPPYHYVSPPPPVKSPPP 327
Query: 79 SSSPSPSP 86
+ SP
Sbjct: 328 PTYIYASP 351
Score = 33.9 bits (76), Expect = 0.031
Identities = 22/65 (33%), Positives = 29/65 (43%), Gaps = 3/65 (4%)
Frame = +1
Query: 25 SPAPTPATNSSLNSPSPTPIPTPS---PANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSS 81
SP+P P + + P P+PI P + PP+P+P P S PP SP
Sbjct: 166 SPSPPPPYHYQ-SPPPPSPISHPPYYYKSPPPPSPSPPPPYHYVSPPPPVKSPPPPTYIY 342
Query: 82 PSPSP 86
SP P
Sbjct: 343 ASPPP 357
Score = 32.7 bits (73), Expect = 0.069
Identities = 20/64 (31%), Positives = 29/64 (45%), Gaps = 5/64 (7%)
Frame = +1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPAN-----SPPAPTPTPTPSPHSDSPPAPSPDNSPSS 80
P P+P ++ SP P P+PSP SPP P +P P + + P P N
Sbjct: 205 PPPSPISHPPYYYKSPPP-PSPSPPPPYHYVSPPPPVKSPPPPTYIYASPPPPHYN*AQR 381
Query: 81 SPSP 84
+ +P
Sbjct: 382 AEAP 393
>TC76563 homologue to PIR|C29356|C29356 hydroxyproline-rich glycoprotein
(clone Hyp2.13) - kidney bean (fragment), partial (34%)
Length = 507
Score = 58.2 bits (139), Expect = 2e-09
Identities = 37/88 (42%), Positives = 42/88 (47%), Gaps = 15/88 (17%)
Frame = +2
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAPSPDNSP 78
S P+P P T SP P P P+P P SPP P+P+P P + SPP PSP P
Sbjct: 203 SPPPPSPIPKTPYYYKSPPP-PSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPP 379
Query: 79 ----SSSPSPSPS--------SSPAPSP 94
S P PSPS S P PSP
Sbjct: 380 PYYYKSPPPPSPSPPPPYYYKSPPPPSP 463
Score = 57.0 bits (136), Expect = 3e-09
Identities = 37/87 (42%), Positives = 43/87 (48%), Gaps = 14/87 (16%)
Frame = +2
Query: 22 SADSPAPTPATNSSLNSPSP-TPIP-TPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSP- 78
S P+P+P SP P +PIP TP SPP P+P+P P + SPP PSP P
Sbjct: 155 SPPPPSPSPPPPYYYKSPPPPSPIPKTPYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPP 334
Query: 79 ---SSSPSPSPS--------SSPAPSP 94
S P PSPS S P PSP
Sbjct: 335 YYYKSPPPPSPSPPPPYYYKSPPPPSP 415
Score = 53.1 bits (126), Expect = 5e-08
Identities = 40/104 (38%), Positives = 50/104 (47%), Gaps = 16/104 (15%)
Frame = +2
Query: 7 SLVFLLLSFLVNIASSADS----PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTP 62
+L +L+S V ++++ADS P P S PSP+P P P SPP P+P P
Sbjct: 62 ALAIVLISTSV-VSAAADSYIYSSPPPPYYYKSPPPPSPSP-PPPYYYKSPPPPSPIPKT 235
Query: 63 SPHSDSPPAPSPDNSP----SSSPSPSPS--------SSPAPSP 94
+ SPP PSP P S P PSPS S P PSP
Sbjct: 236 PYYYKSPPPPSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSP 367
Score = 48.1 bits (113), Expect = 2e-06
Identities = 28/69 (40%), Positives = 36/69 (51%), Gaps = 4/69 (5%)
Frame = +2
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAPSP-DNS 77
S P+P+P SP P P P+P P SPP P+P+P P + SPP PSP ++
Sbjct: 299 SPPPPSPSPPPPYYYKSPPP-PSPSPPPPYYYKSPPPPSPSPPPPYYYKSPPPPSPIPHT 475
Query: 78 PSSSPSPSP 86
P SP P
Sbjct: 476 PYYYKSPPP 502
Score = 37.4 bits (85), Expect = 0.003
Identities = 21/54 (38%), Positives = 26/54 (47%), Gaps = 3/54 (5%)
Frame = +2
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPA---NSPPAPTPTPTPSPHSDSPPAP 72
S P+P+P SP P P P+P P SPP P+P P + SPP P
Sbjct: 347 SPPPPSPSPPPPYYYKSPPP-PSPSPPPPYYYKSPPPPSPIPHTPYYYKSPPPP 505
>BQ149865 weakly similar to GP|15128223|db hypothetical protein~similar to
Oryza sativa chromosome 1 P0489A01.2, partial (3%)
Length = 672
Score = 57.8 bits (138), Expect = 2e-09
Identities = 30/56 (53%), Positives = 33/56 (58%), Gaps = 3/56 (5%)
Frame = +1
Query: 46 TPSPANSPPAPTPT---PTPSPHSDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAA 98
TP+PA SPPAPTPT PTP S SPPAP P P+P P S+ AP AA
Sbjct: 13 TPAPATSPPAPTPTSDAPTPDSTSSSPPAPGP-----GGPAPGPGSTDAPPSPSAA 165
Score = 42.0 bits (97), Expect = 1e-04
Identities = 28/57 (49%), Positives = 34/57 (59%), Gaps = 6/57 (10%)
Frame = +1
Query: 23 ADSPAPTPATNSSLNSPSPTPI---PTP-SPANSPPAPTP-TPTPSPHS-DSPPAPS 73
A P PAT+ P+PTP PTP S ++SPPAP P P P P S D+PP+PS
Sbjct: 1 ARGATPAPATSP----PAPTPTSDAPTPDSTSSSPPAPGPGGPAPGPGSTDAPPSPS 159
>BQ151059 similar to GP|7291525|gb|A CG9888-PA {Drosophila melanogaster},
partial (26%)
Length = 308
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 304 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 125
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 124 PPPPPPPPP 98
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 274 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 95
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 94 PPPPPPPPP 68
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 299 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 120
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 119 PPPPPPPPP 93
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 296 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 117
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 116 PPPPPPPPP 90
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 308 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 129
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 128 PPPPPPPPP 102
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -3
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 288 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 109
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 108 PPPPPPPPP 82
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 287 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 108
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 107 PPPPPPPPP 81
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 283 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 104
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 103 PPPPPPPPP 77
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 281 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 102
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 101 PPPPPPPPP 75
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -3
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 276 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 97
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 96 PPPPPPPPP 70
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 302 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 123
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 122 PPPPPPPPP 96
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 271 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 92
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 91 PPPPPPPPP 65
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 269 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 90
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 89 PPPPPPPPP 63
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 265 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 86
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 85 PPPPPPPPP 59
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 263 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 84
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 83 PPPPPPPPP 57
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 260 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 81
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 80 PPPPPPPPP 54
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -3
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 255 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 76
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 75 PPPPPPPPP 49
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 254 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 75
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 74 PPPPPPPPP 48
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -3
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 249 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 70
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 69 PPPPPPPPP 43
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 248 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 69
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 68 PPPPPPPPP 42
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 244 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 65
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 64 PPPPPPPPP 38
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 241 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 62
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 61 PPPPPPPPP 35
Score = 57.8 bits (138), Expect = 2e-09
Identities = 27/69 (39%), Positives = 27/69 (39%)
Frame = -3
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 291 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 112
Query: 86 PSSSPAPSP 94
P P P P
Sbjct: 111 PPPPPPPPP 85
Score = 56.6 bits (135), Expect = 4e-09
Identities = 27/70 (38%), Positives = 28/70 (39%)
Frame = -3
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 228 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 49
Query: 86 PSSSPAPSPD 95
P P SP+
Sbjct: 48 PPPPPGFSPE 19
Score = 52.4 bits (124), Expect = 8e-08
Identities = 25/72 (34%), Positives = 27/72 (36%)
Frame = -2
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P P P P P P P PP P P P P P PP P P P P P
Sbjct: 223 PPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPPP 44
Query: 86 PSSSPAPSPDEA 97
P P ++
Sbjct: 43 PPPGVFPGKGDS 8
>TC79359 weakly similar to PIR|S49915|S49915 extensin-like protein - maize,
partial (3%)
Length = 1170
Score = 57.4 bits (137), Expect = 3e-09
Identities = 46/118 (38%), Positives = 58/118 (48%), Gaps = 25/118 (21%)
Frame = +3
Query: 6 FSLVFLLLSFLVNIASSADSPAPTPATNSSLNSPSPTPI-PTPSPAN-------SPPAPT 57
FSL L + F V A +A PA +P ++ + PS + PT SPA+ SPP+ T
Sbjct: 174 FSLFVLSMLFTVLFAQNA--PASSPKSSVTAKPPSSVSVSPTNSPASPAKSPTLSPPSQT 347
Query: 58 PTPTPS------PHSDSPPAPSPDNSPSSSPSPS-----------PSSSPAPSPDEAA 98
P +PS P + SPPA SP P SS SP+ P SSPA SP AA
Sbjct: 348 PAVSPSGSASTPPPATSPPAKSPAVQPPSSVSPAISPSNNVSSTPPVSSPASSPPTAA 521
Score = 48.9 bits (115), Expect = 9e-07
Identities = 39/113 (34%), Positives = 52/113 (45%), Gaps = 17/113 (15%)
Frame = +3
Query: 23 ADSPAPTPATN-----SSLNSPSPTPIPTPSPANSPP----APTPTPTPSPHSDSPPAPS 73
A SPA P ++ S N+ S TP P SPA+SPP +P +P +P SPP S
Sbjct: 405 AKSPAVQPPSSVSPAISPSNNVSSTP-PVSSPASSPPTAAVSPVSSPVEAPSVSSPPEAS 581
Query: 74 PDNSPSSSPSPS-------PSS-SPAPSPDEAADNNAISHTGIGEDGKSSGGG 118
PSSS +P+ PSS SP SP ++ + + G S G
Sbjct: 582 SAGIPSSSATPADAPAATLPSSKSPGTSPASSSPETSQGPAAADDSGSRSSFG 740
Score = 45.8 bits (107), Expect = 8e-06
Identities = 30/93 (32%), Positives = 44/93 (47%), Gaps = 13/93 (13%)
Frame = +3
Query: 19 IASSADSPAPTPATNSSLNSPSPTPIPTPSPA--------NSPPAPTPTPTP-----SPH 65
++ S + P PAT+ SP+ P + SPA ++PP +P +P SP
Sbjct: 354 VSPSGSASTPPPATSPPAKSPAVQPPSSVSPAISPSNNVSSTPPVSSPASSPPTAAVSPV 533
Query: 66 SDSPPAPSPDNSPSSSPSPSPSSSPAPSPDEAA 98
S APS + P +S + PSSS P+ AA
Sbjct: 534 SSPVEAPSVSSPPEASSAGIPSSSATPADAPAA 632
>TC81043 weakly similar to GP|6523547|emb|CAB62280.1 hydroxyproline-rich
glycoprotein DZ-HRGP {Volvox carteri f. nagariensis},
partial (19%)
Length = 555
Score = 57.0 bits (136), Expect = 3e-09
Identities = 34/73 (46%), Positives = 38/73 (51%)
Frame = +1
Query: 26 PAPTPATNSSLNSPSPTPIPTPSPANSPPAPTPTPTPSPHSDSPPAPSPDNSPSSSPSPS 85
P P P S SP P P PSP SPPAP P P P P SPPAP SP S P S
Sbjct: 199 PKPPPPAPPSPGSPPKPPAP-PSPG-SPPAPAPAPPPIP--PSPPAPPSPGSPPSPPPAS 366
Query: 86 PSSSPAPSPDEAA 98
P+ P P+P + +
Sbjct: 367 PAP-PKPTPPKGS 402
Score = 56.6 bits (135), Expect = 4e-09
Identities = 31/74 (41%), Positives = 40/74 (53%), Gaps = 3/74 (4%)
Frame = +1
Query: 22 SADSPAPTPATNSSLNSPSPTPIPTPSPAN--SPPAPTPTPTPSPHSDSPPAPSPDNSPS 79
S SP PA S + P+P P P P P + +PP+P P+P P S +PP P+P P
Sbjct: 226 SPGSPPKPPAPPSPGSPPAPAPAPPPIPPSPPAPPSPGSPPSPPPASPAPPKPTP---PK 396
Query: 80 SSP-SPSPSSSPAP 92
SP P P +PAP
Sbjct: 397 GSPLLPMPEKTPAP 438
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.305 0.124 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,128,669
Number of Sequences: 36976
Number of extensions: 244968
Number of successful extensions: 23828
Number of sequences better than 10.0: 1899
Number of HSP's better than 10.0 without gapping: 4655
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 12147
length of query: 166
length of database: 9,014,727
effective HSP length: 89
effective length of query: 77
effective length of database: 5,723,863
effective search space: 440737451
effective search space used: 440737451
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144405.9


