

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.8 - phase: 0 /pseudo
(82 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
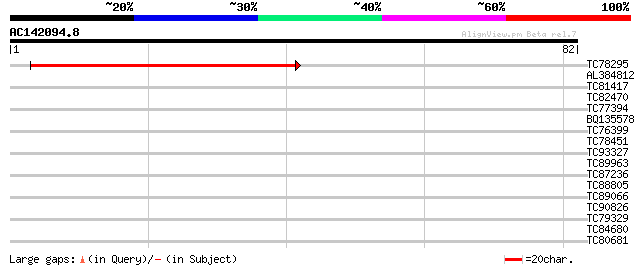
Score E
Sequences producing significant alignments: (bits) Value
TC78295 similar to PIR|T48496|T48496 membrane protein - Arabidop... 44 1e-05
AL384812 27 1.1
TC81417 weakly similar to GP|10998142|dbj|BAB03113. gene_id:MEC1... 27 1.4
TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical prot... 27 1.8
TC77394 similar to GP|4874305|gb|AAD31367.1| expressed protein {... 27 1.8
BQ135578 27 1.8
TC76399 similar to GP|14165163|gb|AAK55409.1 2-on-2 hemoglobin {... 26 3.1
TC78451 similar to GP|9279625|dbj|BAB01083.1 gb|AAF23830.1~gene_... 25 5.4
TC93327 25 5.4
TC89963 weakly similar to GP|6681331|gb|AAF23248.1| B' regulator... 25 7.0
TC87236 homologue to GP|7381203|gb|AAF61434.1| pathogenesis-rela... 25 7.0
TC88805 25 7.0
TC89066 similar to PIR|B84597|B84597 probable disease resistance... 24 9.2
TC90826 homologue to GP|6446577|gb|AAD39534.2| cellulose synthas... 24 9.2
TC79329 similar to PIR|T00456|T00456 protein kinase homolog T14N... 24 9.2
TC84680 similar to GP|21536636|gb|AAM60968.1 unknown {Arabidopsi... 24 9.2
TC80681 weakly similar to PIR|H86265|H86265 protein F3F19.18 [im... 24 9.2
>TC78295 similar to PIR|T48496|T48496 membrane protein - Arabidopsis
thaliana, partial (82%)
Length = 1223
Score = 43.5 bits (101), Expect = 1e-05
Identities = 19/39 (48%), Positives = 28/39 (71%)
Frame = +1
Query: 4 LSIGMVPVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
+++ + V NL IG++P V+NF IGG + GFLLGF+ L
Sbjct: 661 MTLLFIIVINLVIGMLPHVDNFAHIGGFLTGFLLGFIFL 777
>AL384812
Length = 483
Score = 27.3 bits (59), Expect = 1.1
Identities = 10/14 (71%), Positives = 12/14 (85%)
Frame = +3
Query: 39 FVLLCKKDPFVLPD 52
FV LC +DPF+LPD
Sbjct: 135 FVFLCFRDPFLLPD 176
>TC81417 weakly similar to GP|10998142|dbj|BAB03113.
gene_id:MEC18.14~ref|NP_037458.1~similar to unknown
protein {Arabidopsis thaliana}, partial (23%)
Length = 841
Score = 26.9 bits (58), Expect = 1.4
Identities = 12/36 (33%), Positives = 19/36 (52%)
Frame = +1
Query: 42 LCKKDPFVLPDQKLHKRCLPIICFILLSTGWTGVLT 77
LCK++P +L Q RC + +L+ W +LT
Sbjct: 226 LCKRNPNLLQQQSHPMRCTTLSFRLLMKRWWETLLT 333
>TC82470 homologue to GP|23497460|gb|AAN37003.1 hypothetical protein
{Plasmodium falciparum 3D7}, partial (8%)
Length = 1064
Score = 26.6 bits (57), Expect = 1.8
Identities = 14/35 (40%), Positives = 19/35 (54%)
Frame = -2
Query: 36 LLGFVLLCKKDPFVLPDQKLHKRCLPIICFILLST 70
L F L C VL +Q LH+ LP +C+ +L T
Sbjct: 421 LTKFQLFCL*FFLVLSNQVLHQTELPFLCYKILDT 317
>TC77394 similar to GP|4874305|gb|AAD31367.1| expressed protein {Arabidopsis
thaliana}, partial (61%)
Length = 1396
Score = 26.6 bits (57), Expect = 1.8
Identities = 21/62 (33%), Positives = 32/62 (50%), Gaps = 7/62 (11%)
Frame = -1
Query: 14 LTIGIVPIVNNFGLIGGLIP--GFL-----LGFVLLCKKDPFVLPDQKLHKRCLPIICFI 66
L +GI+ + ++F L IP FL LGF+L F+L D K+ + L + F+
Sbjct: 1003 LLLGILGLSSDFFLFAIYIPINTFLILRCTLGFILALTIIVFILCDSKIS*KSLKLFKFL 824
Query: 67 LL 68
LL
Sbjct: 823 LL 818
>BQ135578
Length = 789
Score = 26.6 bits (57), Expect = 1.8
Identities = 9/17 (52%), Positives = 14/17 (81%)
Frame = -3
Query: 6 IGMVPVFNLTIGIVPIV 22
IG+ P+FNLTI +P++
Sbjct: 268 IGVCPIFNLTINTLPLI 218
>TC76399 similar to GP|14165163|gb|AAK55409.1 2-on-2 hemoglobin {Arabidopsis
thaliana}, partial (88%)
Length = 908
Score = 25.8 bits (55), Expect = 3.1
Identities = 16/43 (37%), Positives = 23/43 (53%)
Frame = +1
Query: 33 PGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFILLSTGWTGV 75
P F +GF+++ KK+ FV Q L K+ L I L W G+
Sbjct: 232 PIFTIGFMMMKKKNGFVPYLQILRKKKLFRISMNSLCKEWEGL 360
>TC78451 similar to GP|9279625|dbj|BAB01083.1
gb|AAF23830.1~gene_id:MOJ10.9~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (88%)
Length = 2452
Score = 25.0 bits (53), Expect = 5.4
Identities = 20/67 (29%), Positives = 34/67 (49%), Gaps = 5/67 (7%)
Frame = +1
Query: 13 NLTIGIVPIVN-NFGLIGGLI----PGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFIL 67
NLT+GI+P +N GL+G G L+ ++ K PF + + + C+ + C+ L
Sbjct: 358 NLTVGIIPSLNVAAGLLGFFFVKTWTGLLMKLGIVTK--PFTRQENTVIQTCV-VACYGL 528
Query: 68 LSTGWTG 74
+G G
Sbjct: 529 SFSGGFG 549
>TC93327
Length = 756
Score = 25.0 bits (53), Expect = 5.4
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = +2
Query: 18 IVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLP 51
++P++N G ++ FLL +V LC+ F LP
Sbjct: 95 LLPLINLVGFCV-VLAQFLLSYVALCQISSFFLP 193
>TC89963 weakly similar to GP|6681331|gb|AAF23248.1| B' regulatory subunit
of PP2A (AtB'beta) {Arabidopsis thaliana}, partial (15%)
Length = 981
Score = 24.6 bits (52), Expect = 7.0
Identities = 21/70 (30%), Positives = 28/70 (40%)
Frame = +1
Query: 13 NLTIGIVPIVNNFGLIGGLIPGFLLGFVLLCKKDPFVLPDQKLHKRCLPIICFILLSTGW 72
N+ I +V N F ++ L+ +G VL C +DP L R L W
Sbjct: 448 NIIICLVQTTNTFKIVLFLLRWC*VGDVLFCLEDP-------LGNR---------LQWWW 579
Query: 73 TGVLTKGSEH 82
GVL G H
Sbjct: 580 GGVLCVGGSH 609
>TC87236 homologue to GP|7381203|gb|AAF61434.1| pathogenesis-related protein
4A {Pisum sativum}, partial (93%)
Length = 701
Score = 24.6 bits (52), Expect = 7.0
Identities = 15/52 (28%), Positives = 27/52 (51%), Gaps = 3/52 (5%)
Frame = -1
Query: 24 NFGLIGGLIPGFLLGFVL---LCKKDPFVLPDQKLHKRCLPIICFILLSTGW 72
+ G + L+P L ++ +C+K+ +L Q +HKR IC+ + GW
Sbjct: 323 SIGPLFSLLPELQLQYLSQSSICRKNLCLLA-QLVHKRQSNHICYAMTRAGW 171
>TC88805
Length = 491
Score = 24.6 bits (52), Expect = 7.0
Identities = 9/28 (32%), Positives = 16/28 (57%)
Frame = +3
Query: 27 LIGGLIPGFLLGFVLLCKKDPFVLPDQK 54
L+ G++ + +L K DPF PD++
Sbjct: 90 LVAGVVATVAVANFVLVKNDPFTKPDER 173
>TC89066 similar to PIR|B84597|B84597 probable disease resistance response
protein [imported] - Arabidopsis thaliana, partial (47%)
Length = 639
Score = 24.3 bits (51), Expect = 9.2
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = +3
Query: 8 MVPVFNLTIGIVPIVNNFGLIGGLIPGFLL 37
+VP F++ G++P++ + GLI +P LL
Sbjct: 480 VVPAFSVWQGVMPLLRHIGLIS--LPVMLL 563
>TC90826 homologue to GP|6446577|gb|AAD39534.2| cellulose synthase catalytic
subunit {Gossypium hirsutum}, partial (20%)
Length = 702
Score = 24.3 bits (51), Expect = 9.2
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +3
Query: 22 VNNFGLIGGLIPGFLLG 38
+NNFG +GG+ FLLG
Sbjct: 651 MNNFGXLGGVXLIFLLG 701
>TC79329 similar to PIR|T00456|T00456 protein kinase homolog T14N5.13 -
Arabidopsis thaliana, partial (32%)
Length = 1497
Score = 24.3 bits (51), Expect = 9.2
Identities = 12/32 (37%), Positives = 20/32 (62%)
Frame = -1
Query: 34 GFLLGFVLLCKKDPFVLPDQKLHKRCLPIICF 65
GFLLGF+L+ P ++ L+K + I+C+
Sbjct: 1431 GFLLGFLLIKNHIP-IINGFPLNKNQIEILCY 1339
>TC84680 similar to GP|21536636|gb|AAM60968.1 unknown {Arabidopsis
thaliana}, partial (26%)
Length = 431
Score = 24.3 bits (51), Expect = 9.2
Identities = 12/31 (38%), Positives = 20/31 (63%)
Frame = -1
Query: 13 NLTIGIVPIVNNFGLIGGLIPGFLLGFVLLC 43
+L + +PIV+ F L+ L+ +L F+LLC
Sbjct: 95 HLMLSAIPIVSFFCLLLLLVSVLILLFLLLC 3
>TC80681 weakly similar to PIR|H86265|H86265 protein F3F19.18 [imported] -
Arabidopsis thaliana, partial (6%)
Length = 750
Score = 24.3 bits (51), Expect = 9.2
Identities = 11/33 (33%), Positives = 16/33 (48%)
Frame = +3
Query: 10 PVFNLTIGIVPIVNNFGLIGGLIPGFLLGFVLL 42
P F L + + N+ L+ FLLGFV +
Sbjct: 282 PPFTLNTSLNSLTNSLSCFVALLEHFLLGFVTI 380
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.331 0.154 0.501
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,330,671
Number of Sequences: 36976
Number of extensions: 53273
Number of successful extensions: 309
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 307
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 309
length of query: 82
length of database: 9,014,727
effective HSP length: 58
effective length of query: 24
effective length of database: 6,870,119
effective search space: 164882856
effective search space used: 164882856
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC142094.8


