

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142094.3 + phase: 0
(85 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
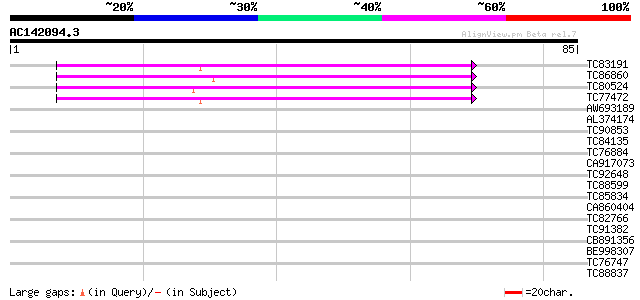
Score E
Sequences producing significant alignments: (bits) Value
TC83191 similar to GP|6714272|gb|AAF25968.1| F6N18.8 {Arabidopsi... 56 3e-09
TC86860 similar to PIR|T05076|T05076 hypothetical protein T6K21.... 55 4e-09
TC80524 similar to GP|6714272|gb|AAF25968.1| F6N18.8 {Arabidopsi... 55 6e-09
TC77472 similar to PIR|T05076|T05076 hypothetical protein T6K21.... 52 4e-08
AW693189 similar to PIR|T05076|T050 hypothetical protein T6K21.8... 30 0.13
AL374174 weakly similar to GP|15810036|gb| At1g10180/F14N23_6 {A... 30 0.16
TC90853 similar to GP|2342427|dbj|BAA21857.1 NPK1-related protei... 29 0.28
TC84135 similar to SP|Q9M5Q1|FUT1_PEA Galactoside 2-L-fucosyltra... 26 2.4
TC76884 similar to SP|P26690|6DCS_SOYBN NAD(P)H dependent 6'-deo... 26 2.4
CA917073 similar to GP|10177918|db kinesin-like protein {Arabido... 26 3.1
TC92648 similar to GP|12039342|gb|AAG46129.1 hypothetical protei... 25 4.0
TC88599 25 4.0
TC85834 similar to GP|16974583|gb|AAL31187.1 AT4g39080/F19H22_18... 25 4.0
CA860404 25 5.3
TC82766 similar to GP|3128183|gb|AAC16087.1| unknown protein {Ar... 25 5.3
TC91382 GP|21327675|gb|AAC37438.2 CPH1 {Chlamydomonas reinhardti... 25 5.3
CB891356 similar to GP|20269067|em protease inhibitor {Sesbania ... 25 5.3
BE998307 25 5.3
TC76747 similar to GP|20269067|emb|CAD29731. protease inhibitor ... 25 5.3
TC88837 similar to GP|23617012|dbj|BAC20708. putative translatio... 25 6.9
>TC83191 similar to GP|6714272|gb|AAF25968.1| F6N18.8 {Arabidopsis
thaliana}, partial (58%)
Length = 1053
Score = 55.8 bits (133), Expect = 3e-09
Identities = 27/64 (42%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Frame = +3
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C +H HR QI R SY DV R SE+QK D
Sbjct: 198 CKVHADSNKSECNMYCLDCMNGALCSSCLASHKDHRAIQIRRSSYHDVIRVSEIQKFLDI 377
Query: 67 SNIQ 70
+ +Q
Sbjct: 378 TGVQ 389
>TC86860 similar to PIR|T05076|T05076 hypothetical protein T6K21.80 -
Arabidopsis thaliana, partial (68%)
Length = 1198
Score = 55.5 bits (132), Expect = 4e-09
Identities = 27/64 (42%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Frame = +3
Query: 8 CDDHRDLRSNEKNTFCVDCAVR-FCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H HR QI R SY DV R +E+QKH D
Sbjct: 210 CKIHADSHKSECNMYCLDCMNGPLCSLCLAYHKDHRYIQIRRSSYHDVIRVNEIQKHLDI 389
Query: 67 SNIQ 70
+ +Q
Sbjct: 390 AGVQ 401
>TC80524 similar to GP|6714272|gb|AAF25968.1| F6N18.8 {Arabidopsis
thaliana}, partial (62%)
Length = 1080
Score = 54.7 bits (130), Expect = 6e-09
Identities = 27/64 (42%), Positives = 34/64 (52%), Gaps = 1/64 (1%)
Frame = +2
Query: 8 CDDHRDLRSNEKNTFCVDC-AVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C +H HR QI R SY DV R SE+QK D
Sbjct: 161 CKVHSDSHKSECNMYCLDCNNGALCSVCLASHKQHRTIQIRRSSYHDVIRVSEIQKFLDI 340
Query: 67 SNIQ 70
+ +Q
Sbjct: 341 AEVQ 352
>TC77472 similar to PIR|T05076|T05076 hypothetical protein T6K21.80 -
Arabidopsis thaliana, partial (68%)
Length = 1426
Score = 52.0 bits (123), Expect = 4e-08
Identities = 26/64 (40%), Positives = 33/64 (50%), Gaps = 1/64 (1%)
Frame = +3
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDC 66
C H D +E N +C+DC C C H HR QI R SY DV R +E+QK D
Sbjct: 267 CKLHADSHKSECNMYCLDCMNGALCSLCLNYHKDHRAIQIRRSSYHDVIRVNEIQKVLDI 446
Query: 67 SNIQ 70
+ +Q
Sbjct: 447 TGVQ 458
>AW693189 similar to PIR|T05076|T050 hypothetical protein T6K21.80 -
Arabidopsis thaliana, partial (30%)
Length = 615
Score = 30.4 bits (67), Expect = 0.13
Identities = 14/40 (35%), Positives = 18/40 (45%), Gaps = 1/40 (2%)
Frame = +3
Query: 8 CDDHRDLRSNEKNTFCVDCA-VRFCRHCKEAHSIHRRFQI 46
C H D +E N +C+DC C C H HR Q+
Sbjct: 207 CKIHADSHKSECNMYCLDCMNGPLCSLCLAYHKDHRYIQV 326
>AL374174 weakly similar to GP|15810036|gb| At1g10180/F14N23_6 {Arabidopsis
thaliana}, partial (8%)
Length = 199
Score = 30.0 bits (66), Expect = 0.16
Identities = 15/35 (42%), Positives = 17/35 (47%), Gaps = 1/35 (2%)
Frame = +1
Query: 2 NSSSGYCDDHRDLRS-NEKNTFCVDCAVRFCRHCK 35
NS YC H DL +K T C C CR+CK
Sbjct: 61 NSIRYYCSCHPDLLC*GDKPTKCTSCG*MVCRNCK 165
>TC90853 similar to GP|2342427|dbj|BAA21857.1 NPK1-related protein kinase 3
{Arabidopsis thaliana}, partial (22%)
Length = 746
Score = 29.3 bits (64), Expect = 0.28
Identities = 15/45 (33%), Positives = 22/45 (48%), Gaps = 5/45 (11%)
Frame = +2
Query: 22 FCVDCAVRFCR-----HCKEAHSIHRRFQIYRYSYQDVFRHSELQ 61
F + C R R HC ++ RR + R+ QD RHS++Q
Sbjct: 83 FLLTCNARLLRLNSPIHCLLNPALRRRCTLLRFRQQDRLRHSQIQ 217
>TC84135 similar to SP|Q9M5Q1|FUT1_PEA Galactoside 2-L-fucosyltransferase
(EC 2.4.1.69) (Xyloglucan alpha- (1
2)-fucosyltransferase), partial (41%)
Length = 814
Score = 26.2 bits (56), Expect = 2.4
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = -3
Query: 10 DHRDLRSNEKNTFCVDCAVRFCR 32
DH D R N+KNT C + +++ CR
Sbjct: 428 DHIDSRVNQKNTICEE-SIKKCR 363
>TC76884 similar to SP|P26690|6DCS_SOYBN NAD(P)H dependent 6'-deoxychalcone
synthase (EC 1.1.-.-). [Soybean] {Glycine max}, partial
(95%)
Length = 1256
Score = 26.2 bits (56), Expect = 2.4
Identities = 15/38 (39%), Positives = 18/38 (46%), Gaps = 1/38 (2%)
Frame = -2
Query: 31 CRHCK-EAHSIHRRFQIYRYSYQDVFRHSELQKHFDCS 67
C C+ EAH HR F + +D FR HF CS
Sbjct: 166 CFGCRAEAH--HRHFPLSGGGGEDYFRDCRCSSHFVCS 59
>CA917073 similar to GP|10177918|db kinesin-like protein {Arabidopsis
thaliana}, partial (4%)
Length = 788
Score = 25.8 bits (55), Expect = 3.1
Identities = 11/41 (26%), Positives = 22/41 (52%)
Frame = -2
Query: 25 DCAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFD 65
+C FC+ + S H+ +++ +Q ++ SEL +H D
Sbjct: 787 NCRTSFCQVMRWKRSHHQWLLQFKWKWQKPWKLSELIRHSD 665
>TC92648 similar to GP|12039342|gb|AAG46129.1 hypothetical protein {Oryza
sativa}, partial (70%)
Length = 900
Score = 25.4 bits (54), Expect = 4.0
Identities = 12/30 (40%), Positives = 15/30 (50%), Gaps = 1/30 (3%)
Frame = +3
Query: 20 NTFCVDCAVRFC-RHCKEAHSIHRRFQIYR 48
N CV C VRFC C+ H+ R + R
Sbjct: 492 NYTCVRCGVRFCSNRCQNVHNDTRCLKFCR 581
>TC88599
Length = 1291
Score = 25.4 bits (54), Expect = 4.0
Identities = 15/60 (25%), Positives = 26/60 (43%)
Frame = +1
Query: 26 CAVRFCRHCKEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCSNIQVTLNALLQVIDSFSL 85
C +R C H + + S + R I SY L + + C+N +N + ++ S L
Sbjct: 1018 CIIRRCVHHRVSDSENSRLGIRSRSYLLYMNKLNLLRSWGCNNRLYLINNVYMILVSMYL 1197
>TC85834 similar to GP|16974583|gb|AAL31187.1 AT4g39080/F19H22_180
{Arabidopsis thaliana}, partial (97%)
Length = 2959
Score = 25.4 bits (54), Expect = 4.0
Identities = 13/31 (41%), Positives = 18/31 (57%), Gaps = 1/31 (3%)
Frame = -3
Query: 41 HRRFQIYRYSYQDVF-RHSELQKHFDCSNIQ 70
H+R I+ S+ +F RH QKHF S +Q
Sbjct: 944 HQRRHIFSESWLSLFHRHKRRQKHFSQSFLQ 852
>CA860404
Length = 496
Score = 25.0 bits (53), Expect = 5.3
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = -2
Query: 47 YRYSYQDVFRHSELQKHF 64
Y Y Y D+ ++SEL +HF
Sbjct: 258 YLYPYVDLEKYSELMRHF 205
>TC82766 similar to GP|3128183|gb|AAC16087.1| unknown protein {Arabidopsis
thaliana}, partial (10%)
Length = 690
Score = 25.0 bits (53), Expect = 5.3
Identities = 11/38 (28%), Positives = 21/38 (54%), Gaps = 2/38 (5%)
Frame = -3
Query: 38 HSIHRRFQIYR--YSYQDVFRHSELQKHFDCSNIQVTL 73
+S HR F + Y++ +F+ +F C N+Q+T+
Sbjct: 286 YSFHRLFTQIKLHYNFSSLFKPMPSTTYF*CFNVQITV 173
>TC91382 GP|21327675|gb|AAC37438.2 CPH1 {Chlamydomonas reinhardtii}, partial
(1%)
Length = 702
Score = 25.0 bits (53), Expect = 5.3
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = +1
Query: 46 IYRYSYQDVFRHSELQKHFDCSNI 69
++ Y DV ++ LQ+HFD +NI
Sbjct: 631 LFMDDYMDVDNYTLLQEHFDNANI 702
>CB891356 similar to GP|20269067|em protease inhibitor {Sesbania rostrata},
partial (51%)
Length = 829
Score = 25.0 bits (53), Expect = 5.3
Identities = 9/35 (25%), Positives = 17/35 (47%)
Frame = +3
Query: 1 MNSSSGYCDDHRDLRSNEKNTFCVDCAVRFCRHCK 35
M ++ C H+ +++ FC C + C HC+
Sbjct: 216 MQRNTPVCRLHQGYGLHQEGMFCFLCVLM*CLHCR 320
>BE998307
Length = 624
Score = 25.0 bits (53), Expect = 5.3
Identities = 13/51 (25%), Positives = 22/51 (42%), Gaps = 4/51 (7%)
Frame = -1
Query: 23 CVDCAVRFCRHC----KEAHSIHRRFQIYRYSYQDVFRHSELQKHFDCSNI 69
C+ C + F HC + HR+ + + +F H E +H + S I
Sbjct: 363 CLICILNFSIHCHPLCRRRLRFHRQGSVRYFHCGSLFHHYESLRHLNHSPI 211
>TC76747 similar to GP|20269067|emb|CAD29731. protease inhibitor {Sesbania
rostrata}, partial (86%)
Length = 1189
Score = 25.0 bits (53), Expect = 5.3
Identities = 9/35 (25%), Positives = 17/35 (47%)
Frame = -2
Query: 1 MNSSSGYCDDHRDLRSNEKNTFCVDCAVRFCRHCK 35
M ++ C H+ +++ FC C + C HC+
Sbjct: 354 MQRNTPVCRLHQGYGLHQEGMFCFLCVLM*CLHCR 250
>TC88837 similar to GP|23617012|dbj|BAC20708. putative translational
inhibitor protein {Oryza sativa (japonica
cultivar-group)}, partial (68%)
Length = 897
Score = 24.6 bits (52), Expect = 6.9
Identities = 11/25 (44%), Positives = 14/25 (56%)
Frame = -1
Query: 19 KNTFCVDCAVRFCRHCKEAHSIHRR 43
+N+F VD + H KE HS RR
Sbjct: 270 RNSFLVDPGISGKTHAKEGHSALRR 196
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.329 0.137 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,549,174
Number of Sequences: 36976
Number of extensions: 53878
Number of successful extensions: 436
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 435
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 435
length of query: 85
length of database: 9,014,727
effective HSP length: 61
effective length of query: 24
effective length of database: 6,759,191
effective search space: 162220584
effective search space used: 162220584
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC142094.3


