

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
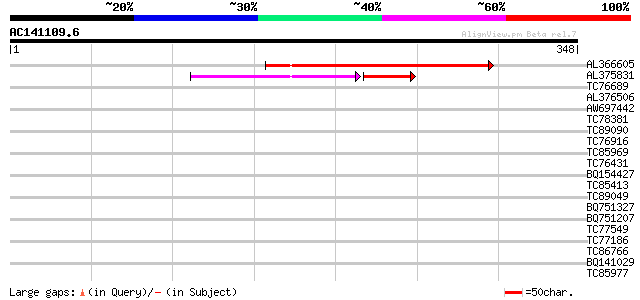
Query= AC141109.6 - phase: 0
(348 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AL366605 132 3e-31
AL375831 72 2e-18
TC76689 MtN12 37 0.015
AL376506 homologue to GP|15021744|gb root nodule extensin {Pisum... 35 0.033
AW697442 homologue to PIR|T00881|T008 probable PCF2-like DNA bin... 34 0.073
TC78381 similar to GP|14335166|gb|AAK59863.1 AT3g25860/MPE11_1 {... 33 0.12
TC89090 33 0.16
TC76916 MtN4 33 0.16
TC85969 similar to GP|21553908|gb|AAM62991.1 unknown {Arabidopsi... 33 0.21
TC76431 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sat... 32 0.28
BQ154427 weakly similar to GP|13277220|emb ZF-HD homeobox protei... 32 0.47
TC85413 similar to PIR|S23737|S23737 proline-rich protein precur... 32 0.47
TC89049 similar to PIR|B96702|B96702 hypothetical protein T23K23... 31 0.62
BQ751327 weakly similar to SP|P43076|PHR1 PH responsive protein ... 31 0.81
BQ751207 weakly similar to GP|604427|gb|AA ACOB protein {Emerice... 31 0.81
TC77549 similar to GP|21592984|gb|AAM64933.1 photolyase/blue-lig... 30 1.1
TC77186 phosphate transporter 30 1.1
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 30 1.4
BQ141029 similar to GP|8132441|gb| extensin {Pisum sativum}, par... 29 2.4
TC85977 similar to GP|10176918|dbj|BAB10162. auxin response fact... 29 2.4
>AL366605
Length = 422
Score = 132 bits (331), Expect = 3e-31
Identities = 60/140 (42%), Positives = 91/140 (64%)
Frame = -3
Query: 158 DMRALRGKEKFGKTAYDLCLVLNVQIPHKFKVPYFEKYKGNSCPEEHLKMYVRRMSTYAR 217
+++A RG KT DLCLV V +P KFK+P F++Y G +CP+ H+ YVR+M Y
Sbjct: 417 EIKANRGNADSFKTQ-DLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNYKD 241
Query: 218 NDQVFIYYFQESLASPASKWYTNLDKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMT 277
ND + I+ FQ+SL A++WYT+L K I TF +L+ F Y +N + P+R L++++
Sbjct: 240 NDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTFDELAAAFKSHYGFNTRLKPNREFLRSLS 61
Query: 278 QENKETFKEYAHRWRDTATQ 297
Q+ +E+F+EYA RWR A +
Sbjct: 60 QKKEESFREYAQRWRGAAAR 1
>AL375831
Length = 467
Score = 71.6 bits (174), Expect(2) = 2e-18
Identities = 37/105 (35%), Positives = 59/105 (55%), Gaps = 1/105 (0%)
Frame = +2
Query: 112 PLVVMTYSAPMIHTVPQIEEPIFHSGNMEVCDRVNDLQEKY-DELQRDMRALRGKEKFGK 170
P + P++HT+PQ ++ V + ++ E+ EL+ +++A RG K
Sbjct: 47 PQTTAAVTEPLVHTLPQGININTQHRSIPVTKTMEEMMEELAKELRHEIQANRGNADSVK 226
Query: 171 TAYDLCLVLNVQIPHKFKVPYFEKYKGNSCPEEHLKMYVRRMSTY 215
T DLCLV V +P KFK+P F++Y G +CP+ H+ YVR+M Y
Sbjct: 227 TQ-DLCLVSKVDVPKKFKIPDFDRYNGLTCPQNHIIKYVRKMGNY 358
Score = 38.5 bits (88), Expect(2) = 2e-18
Identities = 16/32 (50%), Positives = 22/32 (68%)
Frame = +3
Query: 218 NDQVFIYYFQESLASPASKWYTNLDKTRIQTF 249
ND + I+ FQ+SL A++WYT+L K I TF
Sbjct: 366 NDSLMIHCFQDSLMEDAAEWYTSLSKNDIHTF 461
>TC76689 MtN12
Length = 759
Score = 36.6 bits (83), Expect = 0.015
Identities = 30/104 (28%), Positives = 38/104 (35%), Gaps = 6/104 (5%)
Frame = +1
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTH 98
PPPPPV T + P + +P PP P+ P H
Sbjct: 31 PPPPPVHTYPK---------------PVYHSP-------PPPVHHTYPKPVYHSPPPPVH 144
Query: 99 QY-----VAHVPPPPVR-APLVVMTYSAPMIHTVPQIEEPIFHS 136
Y V H PPPPV P V P +HT +P++HS
Sbjct: 145 TYPHPKPVYHSPPPPVHTYPKPVYHSPPPPVHTYVPHPKPVYHS 276
Score = 30.0 bits (66), Expect = 1.4
Identities = 27/83 (32%), Positives = 31/83 (36%), Gaps = 10/83 (12%)
Frame = +1
Query: 40 PPPPVRTQVEASSSTISEWTICADTPTHSAPQ---HSMPWFPPFTTGEILCPIACEAQMP 96
PPPPV T S P H+ P+ HS P P T P+ P
Sbjct: 127 PPPPVHTYPHPKPVYHSP-----PPPVHTYPKPVYHSPPP-PVHTYVPHPKPVYHSPPPP 288
Query: 97 THQYVAHV-------PPPPVRAP 112
H Y HV PPPPV +P
Sbjct: 289 VHTYPPHVPHPVYHSPPPPVHSP 357
>AL376506 homologue to GP|15021744|gb root nodule extensin {Pisum sativum},
partial (68%)
Length = 470
Score = 35.4 bits (80), Expect = 0.033
Identities = 29/106 (27%), Positives = 39/106 (36%), Gaps = 8/106 (7%)
Frame = +2
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTH 98
PPPPPV T + P + +P + +P P+ P H
Sbjct: 134 PPPPPVHTYPK---------------PVYHSPPPPVHTYPK--------PVYHSPPPPVH 244
Query: 99 QY-----VAHVPPPPVRA---PLVVMTYSAPMIHTVPQIEEPIFHS 136
+Y V H PPPPV P V P +H P P++HS
Sbjct: 245 KYPHPKPVYHSPPPPVHKYPHPHPVYHSPPPPVHKYPH-PHPVYHS 379
Score = 28.1 bits (61), Expect = 5.2
Identities = 15/42 (35%), Positives = 21/42 (49%), Gaps = 1/42 (2%)
Frame = +2
Query: 96 PTHQYVAHVPPPPVRA-PLVVMTYSAPMIHTVPQIEEPIFHS 136
P + + PPPPV P V P +HT P +P++HS
Sbjct: 110 PPPPHYSSPPPPPVHTYPKPVYHSPPPPVHTYP---KPVYHS 226
>AW697442 homologue to PIR|T00881|T008 probable PCF2-like DNA binding protein
[imported] - Arabidopsis thaliana, partial (26%)
Length = 652
Score = 34.3 bits (77), Expect = 0.073
Identities = 22/56 (39%), Positives = 28/56 (49%), Gaps = 3/56 (5%)
Frame = +1
Query: 10 EVIKTMAEAQAQAQVLAQAQAQAHAQAQVPPPPPVR---TQVEASSSTISEWTICA 62
+V K + +AQAQAQ QAQAQA AQ + + T+VE I CA
Sbjct: 169 QVNKALVQAQAQAQAQTQAQAQAQAQQTIQKRSSTKDRHTKVEGRGRRIRMPATCA 336
>TC78381 similar to GP|14335166|gb|AAK59863.1 AT3g25860/MPE11_1 {Arabidopsis
thaliana}, partial (80%)
Length = 1830
Score = 33.5 bits (75), Expect = 0.12
Identities = 36/123 (29%), Positives = 52/123 (42%), Gaps = 9/123 (7%)
Frame = +2
Query: 8 MAEVIKTMAEAQAQAQVLAQAQAQAHA----QAQVPPPPPVRTQVEASSSTISEWTICAD 63
+AE + +AEAQAQA+ + A + + + +Q PPPPP V++ S + T
Sbjct: 470 LAETAEDIAEAQAQAKSVKSASSSSSSPPQETSQSPPPPPPPAAVKSVSDGPKKIT-ATP 646
Query: 64 TPTHSAPQHSMPWFPPFTTGEI--LCPIACEAQ---MPTHQYVAHVPPPPVRAPLVVMTY 118
A QH + TG + P EA P VA V P AP +
Sbjct: 647 QAKKLAKQHKVDIASVNGTGPFGRITPADVEAAAGITPVKSNVAPVATPTPVAPKGGSSA 826
Query: 119 SAP 121
+AP
Sbjct: 827 AAP 835
>TC89090
Length = 548
Score = 33.1 bits (74), Expect = 0.16
Identities = 16/71 (22%), Positives = 35/71 (48%)
Frame = +1
Query: 7 EMAEVIKTMAEAQAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPT 66
E E+++T++E+ + ++Q + H + PP PP+++ + SS+ +D
Sbjct: 55 EQGELLRTISESTESNEYISQMR---HMTVRQPPSPPIKSICSSGSSSFQFSDESSDYEA 225
Query: 67 HSAPQHSMPWF 77
H+ MP +
Sbjct: 226 HNRSNTEMPTY 258
>TC76916 MtN4
Length = 1850
Score = 33.1 bits (74), Expect = 0.16
Identities = 22/72 (30%), Positives = 29/72 (39%)
Frame = +1
Query: 64 TPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAPMI 123
+PT P H P +PP + CP P H H PP V P V + P +
Sbjct: 583 SPTPKPPVHKPPRYPPKPSP---CPPPSSTPKPPH----HPKPPAVHPPHVPKPH-PPYV 738
Query: 124 HTVPQIEEPIFH 135
P ++ PI H
Sbjct: 739 PKPPIVKPPIVH 774
>TC85969 similar to GP|21553908|gb|AAM62991.1 unknown {Arabidopsis thaliana},
partial (31%)
Length = 2756
Score = 32.7 bits (73), Expect = 0.21
Identities = 14/30 (46%), Positives = 21/30 (69%)
Frame = +2
Query: 10 EVIKTMAEAQAQAQVLAQAQAQAHAQAQVP 39
+V++ +AQ QAQ +QAQ+QA Q+Q P
Sbjct: 1292 QVVQVQPQAQNQAQTQSQAQSQAQTQSQAP 1381
>TC76431 similar to GP|8132441|gb|AAF73291.1| extensin {Pisum sativum},
partial (30%)
Length = 890
Score = 32.3 bits (72), Expect = 0.28
Identities = 22/75 (29%), Positives = 30/75 (39%)
Frame = +1
Query: 41 PPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTHQY 100
PPP + + SS P + HS P PP+T P+ P H Y
Sbjct: 274 PPPPKKPYKYSSPPPPVHPPYTPKPVY----HSPPVHPPYTPH----PVYHSPPPPVHSY 429
Query: 101 VAHVPPPPVRAPLVV 115
+ PPPP + +VV
Sbjct: 430 IYASPPPPYHS*IVV 474
>BQ154427 weakly similar to GP|13277220|emb ZF-HD homeobox protein
{Flaveria bidentis}, partial (28%)
Length = 645
Score = 31.6 bits (70), Expect = 0.47
Identities = 12/26 (46%), Positives = 19/26 (72%)
Frame = +3
Query: 8 MAEVIKTMAEAQAQAQVLAQAQAQAH 33
+ +++K A+ + QAQV+AQ Q QAH
Sbjct: 60 VVQLVKAQAQVKVQAQVIAQVQXQAH 137
>TC85413 similar to PIR|S23737|S23737 proline-rich protein precursor -
kidney bean, partial (56%)
Length = 1139
Score = 31.6 bits (70), Expect = 0.47
Identities = 22/73 (30%), Positives = 31/73 (42%), Gaps = 1/73 (1%)
Frame = +1
Query: 62 ADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMPTHQYVAHVPPPPVRAPLVVMTYSA- 120
A TPT S P PP+ + + A+ PTH + H P PP P+ A
Sbjct: 226 APTPTPSPSPIHTPLHPPYHSAPV------PAKPPTHGHHHHHPHPPAPTPVHTPVAPAH 387
Query: 121 PMIHTVPQIEEPI 133
P +H P + P+
Sbjct: 388 PPLH--PPVHTPV 420
Score = 29.3 bits (64), Expect = 2.4
Identities = 19/66 (28%), Positives = 27/66 (40%), Gaps = 5/66 (7%)
Frame = +1
Query: 75 PWFPPFTTGEILCPIACEAQMPTHQYVAHV-----PPPPVRAPLVVMTYSAPMIHTVPQI 129
P +PP TT P+ A P H + H+ P P +P + T P H+ P
Sbjct: 130 PTYPPHTTP---APLHPPANAPHHHHHHHIHSPTPAPTPTPSPSPIHTPLHPPYHSAPVP 300
Query: 130 EEPIFH 135
+P H
Sbjct: 301 AKPPTH 318
>TC89049 similar to PIR|B96702|B96702 hypothetical protein T23K23.22
[imported] - Arabidopsis thaliana, partial (34%)
Length = 880
Score = 31.2 bits (69), Expect = 0.62
Identities = 38/173 (21%), Positives = 64/173 (36%), Gaps = 20/173 (11%)
Frame = +2
Query: 66 THSAPQHSMPWFPPFTTGEILCPIACEAQMPTHQYVAHVPPPP---------------VR 110
+HS P P+ PP++ P P H A +PPPP +
Sbjct: 2 SHSIPSAPTPFSPPYSH----LPSTPPRSPPPHSPPAPLPPPPKNSTTQLASSRISSVPK 169
Query: 111 APLVVMTYS--APM---IHTVPQIEEPIFHSGNMEVCDRVNDLQEKYDELQRDMRALRGK 165
+ LV M+ S +P+ + T+ + +P+ H N V+DL Q + L
Sbjct: 170 SSLVTMSSSRNSPLSITLITLSPLSDPLSHLFNPPFA--VSDLNYPIHTAQLPPKPLNSP 343
Query: 166 EKFGKTAYDLCLVLNVQIPHKFKVPYFEKYKGNSCPEEHLKMYVRRMSTYARN 218
++ ++ + F +P+ EK K + L + R S Y N
Sbjct: 344 TSTAPPSFFSIPFVHSESQRSFAIPWLEK*K--KLI*QRLLSFTLRSSVYVMN 496
>BQ751327 weakly similar to SP|P43076|PHR1 PH responsive protein 1 precursor
(PH-regulated protein 1). [Yeast] {Candida albicans},
partial (24%)
Length = 574
Score = 30.8 bits (68), Expect = 0.81
Identities = 24/87 (27%), Positives = 34/87 (38%), Gaps = 10/87 (11%)
Frame = +3
Query: 36 AQVPPPPPVRTQVEAS----------SSTISEWTICADTPTHSAPQHSMPWFPPFTTGEI 85
++ PPPP V ++E+S S+T W A TP+ P P TT
Sbjct: 174 SRTPPPPAVSPRLESSRTPSPTRPPASATFPSWPRLAPTPSEPTPS-----TPSRTTPPA 338
Query: 86 LCPIACEAQMPTHQYVAHVPPPPVRAP 112
A A + +V+ PP V P
Sbjct: 339 *SSSATPASTSSPTWVSPRPPSTVTTP 419
>BQ751207 weakly similar to GP|604427|gb|AA ACOB protein {Emericella
nidulans}, partial (12%)
Length = 758
Score = 30.8 bits (68), Expect = 0.81
Identities = 31/115 (26%), Positives = 41/115 (34%), Gaps = 7/115 (6%)
Frame = +1
Query: 14 TMAEAQAQAQVLAQAQAQAHAQAQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHS 73
T + A+ Q +Q + P PPP T SSS +++ A TPT S P
Sbjct: 118 TSSPARHPTQTPMSSQNSSKPLRSRPSPPPTFTPTSPSSS----FSLTAHTPTISPPPPP 285
Query: 74 MPWF------PPFTTGEILCPI-ACEAQMPTHQYVAHVPPPPVRAPLVVMTYSAP 121
P PP + P PT + PP R P T S+P
Sbjct: 286 PPAITSPSSTPPRPSSSANSPS*LSLTPAPTSPTPPYSPPSTYRRPAFSRTSSSP 450
>TC77549 similar to GP|21592984|gb|AAM64933.1 photolyase/blue-light receptor
PHR2 {Arabidopsis thaliana}, partial (72%)
Length = 1653
Score = 30.4 bits (67), Expect = 1.1
Identities = 14/57 (24%), Positives = 30/57 (52%)
Frame = +1
Query: 40 PPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQMP 96
PPPP++ +V +S+++ ++ T THS+P + F++ + P++ P
Sbjct: 187 PPPPIKLKVPTQASSLTHLSL--STKTHSSPSNKTFKKSSFSSNPLKSPLSLIPHRP 351
>TC77186 phosphate transporter
Length = 2314
Score = 30.4 bits (67), Expect = 1.1
Identities = 28/121 (23%), Positives = 51/121 (42%), Gaps = 5/121 (4%)
Frame = +1
Query: 175 LCLVLNVQIPHKFKVPYFEKYKGNSCPEEHLKMYVRRMSTYARNDQVFIYYFQESLASPA 234
+ L++ HK+KVP FE+ S R + + YY++ + P
Sbjct: 937 VALIVASIFDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWR--MKMPE 1110
Query: 235 SKWYTNL-DKTRIQTFCDLSEVFVEQYNYNVDMAPDRSDLQAMTQENKETF----KEYAH 289
+ YT L K Q D+S+V V++ + +Q MT + + ++ K++A
Sbjct: 1111 TARYTALVAKNAKQAAADMSKVL------QVELEVEEEKVQKMTSDKRNSYGLFSKQFAA 1272
Query: 290 R 290
R
Sbjct: 1273 R 1275
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 30.0 bits (66), Expect = 1.4
Identities = 25/87 (28%), Positives = 31/87 (34%), Gaps = 4/87 (4%)
Frame = +2
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQ-HSMPWFPPFTTGEILCPIACEAQMPT 97
PPPPP C + P S+P H P+ PP P P
Sbjct: 308 PPPPP-----------------CVEPPPPSSPAPHQTPYHPP--------PSPSPPPSPV 412
Query: 98 HQYVAHVPPPPVRA---PLVVMTYSAP 121
+ Y + PPPPV P V Y +P
Sbjct: 413 YAYPS--PPPPVYTSPPPSPVYAYPSP 487
Score = 29.3 bits (64), Expect = 2.4
Identities = 21/79 (26%), Positives = 28/79 (34%), Gaps = 4/79 (5%)
Frame = +2
Query: 39 PPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFP----PFTTGEILCPIACEAQ 94
PPPPP + SS T P+ S P + +P P T P+
Sbjct: 305 PPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPPVYTSPPPSPVYAYPS 484
Query: 95 MPTHQYVAHVPPPPVRAPL 113
P Y + PPP P+
Sbjct: 485 PPPPVYSSPPPPPVYEGPI 541
Score = 29.3 bits (64), Expect = 2.4
Identities = 28/102 (27%), Positives = 32/102 (30%), Gaps = 16/102 (15%)
Frame = +2
Query: 36 AQVPPPPPVRTQVEASSSTISEWTICADTPTHSAPQHSMPWFPPFTTGEILCPIACEAQM 95
A PPPPP S T + + P P HS P PP + P C
Sbjct: 137 AHSPPPPPXSPPPPPHSPTPPVYPYLSPPPP--PPVHSPP--PPVYSPPPPSPPPCVEPP 304
Query: 96 PT----------------HQYVAHVPPPPVRAPLVVMTYSAP 121
P HQ H PP P P V Y +P
Sbjct: 305 PPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSP 430
>BQ141029 similar to GP|8132441|gb| extensin {Pisum sativum}, partial (54%)
Length = 613
Score = 29.3 bits (64), Expect = 2.4
Identities = 14/37 (37%), Positives = 18/37 (47%)
Frame = +2
Query: 96 PTHQYVAHVPPPPVRAPLVVMTYSAPMIHTVPQIEEP 132
P Y H PPPPV +P YS+P V + + P
Sbjct: 125 PPKPYYYHSPPPPVHSPPPPYHYSSPPPPPVYKYKSP 235
>TC85977 similar to GP|10176918|dbj|BAB10162. auxin response factor-like
protein {Arabidopsis thaliana}, partial (65%)
Length = 3162
Score = 29.3 bits (64), Expect = 2.4
Identities = 28/112 (25%), Positives = 46/112 (41%), Gaps = 12/112 (10%)
Frame = +3
Query: 3 IKMREMAEVIKTMAEAQAQA-QVLAQA-------QAQAHAQAQVPPPPPVRTQVEASSST 54
++ + + VI M +A+ +V AQ Q + + + PP PP R V + T
Sbjct: 447 LRPKILCRVINVMLKAEPDTDEVFAQVTLVPEPNQDENAVEKEAPPAPPPRFHVHSFCKT 626
Query: 55 ISEWTICADTPTHSA----PQHSMPWFPPFTTGEILCPIACEAQMPTHQYVA 102
++ +DT TH +H+ E L P+ Q PT + VA
Sbjct: 627 LT----ASDTSTHGGFSVLKRHA---------DECLPPLDMSKQPPTQELVA 743
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.319 0.130 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,818,101
Number of Sequences: 36976
Number of extensions: 187580
Number of successful extensions: 1637
Number of sequences better than 10.0: 68
Number of HSP's better than 10.0 without gapping: 1510
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1595
length of query: 348
length of database: 9,014,727
effective HSP length: 97
effective length of query: 251
effective length of database: 5,428,055
effective search space: 1362441805
effective search space used: 1362441805
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC141109.6


