

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
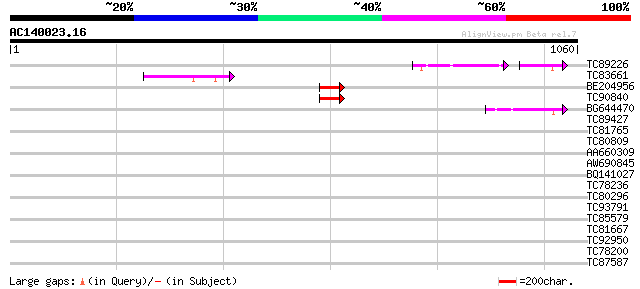
Query= AC140023.16 - phase: 0
(1060 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89226 weakly similar to GP|9759519|dbj|BAB10985.1 gb|AAF16763.... 65 1e-15
TC83661 similar to PIR|E86410|E86410 protein F3M18.14 [imported]... 77 3e-14
BE204956 similar to PIR|E86410|E86 protein F3M18.14 [imported] -... 62 1e-09
TC90840 similar to PIR|E86410|E86410 protein F3M18.14 [imported]... 59 8e-09
BG644470 weakly similar to GP|9759519|dbj gb|AAF16763.1~gene_id:... 47 3e-05
TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed pr... 33 0.76
TC81765 similar to GP|12323731|gb|AAG51827.1 hypothetical protei... 32 0.99
TC80809 similar to GP|22136852|gb|AAM91770.1 unknown protein {Ar... 32 1.7
AA660309 similar to PIR|H84545|H84 probable ubiquitin-conjugatin... 32 1.7
AW690845 similar to PIR|D96715|D967 protein F4N2.10 [imported] -... 31 2.9
BQ141027 weakly similar to GP|11320891|gb| pinin {Homo sapiens},... 30 3.8
TC78236 weakly similar to GP|15451010|gb|AAK96776.1 Unknown prot... 30 3.8
TC80296 similar to PIR|H86265|H86265 protein F3F19.18 [imported]... 30 4.9
TC93791 similar to GP|15810373|gb|AAL07074.1 unknown protein {Ar... 30 4.9
TC85579 homologue to GP|13366196|dbj|BAB39419. putative H+-trans... 30 4.9
TC81667 similar to GP|8777488|dbj|BAA97068.1 contains similarity... 29 8.4
TC92950 similar to GP|21928151|gb|AAM78103.1 AT3g53540/F4P12_240... 29 8.4
TC78200 similar to PIR|A84578|A84578 probable WD-40 repeat prote... 29 8.4
TC87587 similar to GP|20259233|gb|AAM14332.1 putative ankyrin pr... 29 8.4
>TC89226 weakly similar to GP|9759519|dbj|BAB10985.1
gb|AAF16763.1~gene_id:MLN1.10~strong similarity to
unknown protein {Arabidopsis thaliana}, partial (5%)
Length = 1822
Score = 65.1 bits (157), Expect(2) = 1e-15
Identities = 50/190 (26%), Positives = 87/190 (45%), Gaps = 11/190 (5%)
Frame = +2
Query: 754 ALLSVLDDRGKREA------LLIESLERRQTSLCRSMSRIKVSNIGMGCMSHSDQSELDR 807
ALL+ LD RG RE+ L IE++ + ++ K+ N C+ + + E D
Sbjct: 8 ALLNSLDSRGIRESHLRLMLLKIENIFKENVQ--KNAKCAKIGNTDEICVKN-EAEETDS 178
Query: 808 VAED---SCSPVSDVDNLNLTEITDYLPSPGAVVIEAGKKEEEQLHKWIRVQEYDSWIWN 864
+ S SP S + L+ +D + + IE GK E ++ R Q++ W+W
Sbjct: 179 SPDHHTRSDSPSSTLCGLS----SDTSETSASFTIELGKSESDKKASLRRYQDFQKWMWK 346
Query: 865 SFYLD--LNVVKYGRRSYLDSLARCRSCHDLYWRDERHCKICHMTFELDFDLEEKYAIHI 922
Y L +KYG++ + C C +LY ++ HC CH+TF + + ++ H+
Sbjct: 347 ECYNPSILCAMKYGKKRCKPQVDICDICLNLYCLEDSHCSYCHLTFPSNDGFD--FSKHV 520
Query: 923 AMCREKEDSN 932
C +K +
Sbjct: 521 IQCGDKSSKD 550
Score = 37.0 bits (84), Expect(2) = 1e-15
Identities = 29/105 (27%), Positives = 45/105 (42%), Gaps = 16/105 (15%)
Frame = +1
Query: 954 IESVMPEDALVGAWRKSAHNLWIKRLRRTSTLVELLQVLADFVGAFNDSWLFQCKFPDG- 1012
IE +P +A W + LW +L R+S+ ELLQ+L F A +L G
Sbjct: 616 IEGSVPPEAFQSVWTEDIRRLWGVKLSRSSSAEELLQMLTLFERALKRDFLSSPFSTTGD 795
Query: 1013 ---------------VVEETIASFASMPHTSSALALWLVKLDAII 1042
+ E++ +P T+SA++L L +LD I
Sbjct: 796 LLGMNAMSESAAHTSMDLESVTVLPWVPRTTSAVSLRLFELDTSI 930
>TC83661 similar to PIR|E86410|E86410 protein F3M18.14 [imported] -
Arabidopsis thaliana, partial (12%)
Length = 987
Score = 77.4 bits (189), Expect = 3e-14
Identities = 59/207 (28%), Positives = 94/207 (44%), Gaps = 37/207 (17%)
Frame = +1
Query: 251 EKCELALDSSISDAGVDQISMLIDDEELELREIQEGSNLLICSDQLAANGMLGGSLCPDV 310
E+ + AL+ + + + LI+DE+LE+ E+ S L +L + + D
Sbjct: 34 ERRKAALEKATARKIAKESLELIEDEQLEMMELAASSKGLSSIIRLDFDTLQNIESFRDS 213
Query: 311 LVKFPPGDVKMKKPIHLQPWDSSPELVKKLFK---------------------------- 342
L FPP VK++KP +QPW +S + V L
Sbjct: 214 LCLFPPESVKLRKPFAIQPWINSEDNVGNLLMVWRFLITFADVLELWPFTLDEFVQAFHD 393
Query: 343 -DSMLLGQIHVALLTLLLSDIEVELSNGFCPHLNKSCNFLA--------LLHSVENQEYS 393
DS LLG+IHVALL +++ DIE +++ C L + N A ++ +
Sbjct: 394 YDSRLLGEIHVALLKVIIKDIE-DVARTPCTGLGMNQNGAANSGGGHPEIVEGAYAWGFD 570
Query: 394 LDAWRRSLNPLTWIEILRQVLVAAGFG 420
+ W + LN LTW EI RQ+ ++AG+G
Sbjct: 571 IRNWHKHLNQLTWPEIFRQLALSAGYG 651
>BE204956 similar to PIR|E86410|E86 protein F3M18.14 [imported] - Arabidopsis
thaliana, partial (3%)
Length = 485
Score = 62.0 bits (149), Expect = 1e-09
Identities = 28/47 (59%), Positives = 37/47 (78%)
Frame = +1
Query: 580 EIDESHAGEVWLLGLMDSEYSDLKIEEKLNALAALTGLLSSGSSIRM 626
EIDES +GE W+ GL + EYSDL +EE+LNAL AL G+ + G+SIR+
Sbjct: 43 EIDESKSGEPWVQGLTEGEYSDLSVEERLNALVALVGVANEGNSIRI 183
>TC90840 similar to PIR|E86410|E86410 protein F3M18.14 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 716
Score = 59.3 bits (142), Expect = 8e-09
Identities = 26/47 (55%), Positives = 35/47 (74%)
Frame = +3
Query: 580 EIDESHAGEVWLLGLMDSEYSDLKIEEKLNALAALTGLLSSGSSIRM 626
E+DES +GE W+ GL + EYSDL +EE+LNAL L GL + G+ IR+
Sbjct: 252 EVDESKSGESWIQGLTEGEYSDLSVEERLNALVVLVGLANEGNLIRV 392
>BG644470 weakly similar to GP|9759519|dbj gb|AAF16763.1~gene_id:MLN1.10~strong
similarity to unknown protein {Arabidopsis thaliana},
partial (4%)
Length = 814
Score = 47.4 bits (111), Expect = 3e-05
Identities = 42/170 (24%), Positives = 73/170 (42%), Gaps = 16/170 (9%)
Frame = +3
Query: 890 CHDLYWRDERHCKICHMTFELDFDLEEKYAIHIAMCREKEDSNTFPNHKVLPSQIQSLKA 949
C D Y+ ++ HC CH TF + E + H C + + LP + + LK
Sbjct: 3 CLDPYFMEDSHCNSCHQTF--PSNNEFNISKHTFQCVGNLSKDIMEHS--LPLRTRLLKV 170
Query: 950 AIYAIESVMPEDALVGAWRKSAHNLWIKRLRRTSTLVELLQVLADFVGAFNDSWL-FQCK 1008
+ +E+ + +A W W +L ++ST+ ELLQ+L F A +L
Sbjct: 171 LLSCMEASVLSEAFGTIWTTDFRKHWGVKLNKSSTVEELLQMLTLFEKALRRDFLSSNFS 350
Query: 1009 FPDGVV---------------EETIASFASMPHTSSALALWLVKLDAIIA 1043
D ++ E++A +P T++AL+L L + D+ I+
Sbjct: 351 TTDELLGLSSMSKSAAHVSADPESVALLPWVPLTTAALSLRLFEFDSSIS 500
>TC89427 weakly similar to GP|4557063|gb|AAD22502.1| expressed protein
{Arabidopsis thaliana}, partial (50%)
Length = 753
Score = 32.7 bits (73), Expect = 0.76
Identities = 30/108 (27%), Positives = 51/108 (46%), Gaps = 14/108 (12%)
Frame = +1
Query: 456 LSERGNNGCKVSELAKSMQIAELNLSSTTEELESLIYSTLSSDITLFEKISSSAYRLRMS 515
L+ R NN VS A+++ + +L +T + L L+ +T S + IS R +
Sbjct: 82 LAFRNNNTISVSSFAEALILQ--SLIATAKSLAYLLIATGSLLTDVNHLISPMDGRFPLD 255
Query: 516 TVAK--------------DDDDSQSDTEDSGSVDDELNDSDTCSSGDD 549
++K +DDD + D +D V+DE +D+D SGD+
Sbjct: 256 QLSKGNHTCEENKDGSETEDDDDEDDDDD---VNDEDDDNDEDFSGDE 390
>TC81765 similar to GP|12323731|gb|AAG51827.1 hypothetical protein;
55509-55015 {Arabidopsis thaliana}, partial (25%)
Length = 710
Score = 32.3 bits (72), Expect = 0.99
Identities = 16/33 (48%), Positives = 23/33 (69%), Gaps = 4/33 (12%)
Frame = +1
Query: 521 DDDSQSDTEDSGSVDDE--LNDS--DTCSSGDD 549
DDDS++D+E+ S DD+ NDS D C +G+D
Sbjct: 475 DDDSETDSEEESSTDDDGGENDSNFDDCDNGND 573
>TC80809 similar to GP|22136852|gb|AAM91770.1 unknown protein {Arabidopsis
thaliana}, partial (11%)
Length = 681
Score = 31.6 bits (70), Expect = 1.7
Identities = 18/40 (45%), Positives = 22/40 (55%)
Frame = +3
Query: 515 STVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGS 554
S ++ D DD +D GS DDE ND+D G D SGS
Sbjct: 198 SDLSSDGDDQLADDFLQGS-DDEDNDNDNDDEGSDLESGS 314
Score = 30.4 bits (67), Expect = 3.8
Identities = 26/66 (39%), Positives = 33/66 (49%), Gaps = 1/66 (1%)
Frame = +3
Query: 520 DDDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKLRRHNSRKAKHNKLKVYT 579
D+DD SD E SGS DE DSD+ S +D S + RK R + A + + T
Sbjct: 276 DNDDEGSDLE-SGSGSDEDEDSDSESGEEDIARKSKLID-RKRERDSQAAADELQTNIQT 449
Query: 580 EI-DES 584
I DES
Sbjct: 450 NIQDES 467
>AA660309 similar to PIR|H84545|H84 probable ubiquitin-conjugating enzyme
[imported] - Arabidopsis thaliana, partial (4%)
Length = 600
Score = 31.6 bits (70), Expect = 1.7
Identities = 25/67 (37%), Positives = 35/67 (51%), Gaps = 1/67 (1%)
Frame = +1
Query: 521 DDDSQSDTEDSGSVDDELNDSDTCSSGDDFGSGSIHSNIRKLRRHNSRKA-KHNKLKVYT 579
D DS SDT DSG DDE + D GD+ +G +S+ H A + N+L+V
Sbjct: 295 DSDSDSDT-DSGVTDDEGEEGDV-EEGDESNNGCRNSDRNDASVHRKTAALQTNELRVLW 468
Query: 580 EIDESHA 586
+DES +
Sbjct: 469 -MDESES 486
>AW690845 similar to PIR|D96715|D967 protein F4N2.10 [imported] - Arabidopsis
thaliana, partial (21%)
Length = 581
Score = 30.8 bits (68), Expect = 2.9
Identities = 16/54 (29%), Positives = 27/54 (49%)
Frame = +1
Query: 129 NPNCKMKAPVKRHGMGKGLATNPNCNMKAPVKRHGMGKGLAANPNSNMKAPVKR 182
NP +K V+R + NPN +K ++R + NPN+ +K P++R
Sbjct: 241 NPYNTIKPSVRRLRLTTSRPFNPNNTIKPTIRRLRLTTIRPYNPNNTIKPPIRR 402
>BQ141027 weakly similar to GP|11320891|gb| pinin {Homo sapiens}, partial
(4%)
Length = 633
Score = 30.4 bits (67), Expect = 3.8
Identities = 14/51 (27%), Positives = 26/51 (50%)
Frame = +2
Query: 627 KDPVKVTADCSSSIQLRGSGAKIKRSVNPIEQMQCTKEVHMNSHACPVDSS 677
+ P+ +TA CS + + + S P + +CT+ ++ ACP +SS
Sbjct: 71 RSPMALTA-CSKNYSKQTLACAVASSTTPNSRAECTRATSSSTTACPTNSS 220
>TC78236 weakly similar to GP|15451010|gb|AAK96776.1 Unknown protein
{Arabidopsis thaliana}, partial (61%)
Length = 1303
Score = 30.4 bits (67), Expect = 3.8
Identities = 17/39 (43%), Positives = 19/39 (48%)
Frame = +3
Query: 143 MGKGLATNPNCNMKAPVKRHGMGKGLAANPNSNMKAPVK 181
+G GL NCN K P K H G G+ NS K P K
Sbjct: 210 LGNGLLPGFNCNGK*PFKAH--GNGILTGFNSTGKRPFK 320
>TC80296 similar to PIR|H86265|H86265 protein F3F19.18 [imported] -
Arabidopsis thaliana, partial (34%)
Length = 1734
Score = 30.0 bits (66), Expect = 4.9
Identities = 25/105 (23%), Positives = 45/105 (42%), Gaps = 2/105 (1%)
Frame = +3
Query: 447 TLKCELFKILSERGNNGCKVSELAKSMQIAELNLSSTTE--ELESLIYSTLSSDITLFEK 504
TL E+ L + + G AK E+N+++ EL +I + + + +
Sbjct: 615 TLFREVCPSLLIKKDRGRPTDPKAKPKAYGEVNVAADVPGAELLQIIDDDVEQESSHSDD 794
Query: 505 ISSSAYRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSGDD 549
S + DDD+Q ++++GS DDE D D S ++
Sbjct: 795 CGSDNAQEDDQVSLNSDDDNQLGSDNTGSDDDEAEDHDGVSDDEN 929
>TC93791 similar to GP|15810373|gb|AAL07074.1 unknown protein {Arabidopsis
thaliana}, partial (40%)
Length = 866
Score = 30.0 bits (66), Expect = 4.9
Identities = 46/186 (24%), Positives = 80/186 (42%), Gaps = 9/186 (4%)
Frame = +1
Query: 245 QNQLPIEKCELALDSSISDAGVDQISMLIDDEELELREIQEGSNLLICSDQLAANGMLGG 304
Q+QL K ++ + + +Q+ L E E Q +NLL+ + A +LG
Sbjct: 262 QDQLQSAKESVSNMKKLHELAQNQLFELRAQSEEERAAKQAEANLLMDEVERAQTMLLGL 441
Query: 305 SLCPDVL---VKFPPGDVKMKKPIHLQPWDSSPELVKKLFKDSMLLGQIHVALLTLLLSD 361
VL ++ D + KK +L DS+ L L L+ ++++ L +
Sbjct: 442 EREKGVLRSQLQTANEDSETKKSDNL---DSNNALENSLIAKEKLISELNMEL-----HN 597
Query: 362 IEVELSNGFCPHLNKSCNFLALLHSVENQEYSLDAWRRSLNPLTWIEILR------QVLV 415
IE LSN H+N F A+L+ +E S+ A ++ L +I+ ++L
Sbjct: 598 IETTLSNEREEHINDVKKFTAMLN---EKEASIVAMKKELQTRPTEKIVDDLRKKVKILQ 768
Query: 416 AAGFGS 421
A G+ S
Sbjct: 769 AVGYNS 786
>TC85579 homologue to GP|13366196|dbj|BAB39419. putative H+-transporting ATP
synthase {Oryza sativa (japonica cultivar-group)},
partial (97%)
Length = 1934
Score = 30.0 bits (66), Expect = 4.9
Identities = 21/73 (28%), Positives = 35/73 (47%), Gaps = 1/73 (1%)
Frame = -3
Query: 609 NALAA-LTGLLSSGSSIRMKDPVKVTADCSSSIQLRGSGAKIKRSVNPIEQMQCTKEVHM 667
N LAA + + SSG S D T +C Q++ G + R++ + +H+
Sbjct: 975 NMLAAFICQIFSSGKSNTRSDD---TLNCRIICQIKEKGHSLHRAILFKVTFEKLCSLHV 805
Query: 668 NSHACPVDSSLLV 680
NSH C +S ++V
Sbjct: 804 NSHGCKHNSKIVV 766
>TC81667 similar to GP|8777488|dbj|BAA97068.1 contains similarity to SF16
protein~gene_id:K15M2.19 {Arabidopsis thaliana}, partial
(34%)
Length = 1216
Score = 29.3 bits (64), Expect = 8.4
Identities = 28/126 (22%), Positives = 50/126 (39%), Gaps = 23/126 (18%)
Frame = +1
Query: 89 QQDQEPVKRRKASKSAI--QSHPNCNMKAPVERHGMGKG----------LATNP---NCK 133
Q+++ +KR +A A Q PNCN + +GK +A P +
Sbjct: 589 QREEAAIKRERAMAYAFSHQWRPNCNQYFGQASYSLGKESWGWSWMERWVAARPWEVRVQ 768
Query: 134 MKAPVKRHGMGKGLAT--------NPNCNMKAPVKRHGMGKGLAANPNSNMKAPVKRHGM 185
+++P K G+ T + +K+P+ + GM G S +K+P+ + M
Sbjct: 769 VQSPKKNKLNGQQQKTKLDKMNHNDTKSPLKSPLTKAGMSNGKENETKSPLKSPLAKASM 948
Query: 186 GKGLMT 191
T
Sbjct: 949 SNAQET 966
>TC92950 similar to GP|21928151|gb|AAM78103.1 AT3g53540/F4P12_240
{Arabidopsis thaliana}, partial (3%)
Length = 707
Score = 29.3 bits (64), Expect = 8.4
Identities = 16/55 (29%), Positives = 26/55 (47%)
Frame = +2
Query: 40 WKKNRKRKSVQELYTTGDIVNTVLLNDAPTLGSEFDSLPSGPKNYNSACQQDQEP 94
WK K + VQ + + + + G+ FDSLPSG + +N ++ EP
Sbjct: 122 WKMAHKSQEVQGTSRSSTLADMLAFPGKRMKGTRFDSLPSG-EGFNDKFARNGEP 283
>TC78200 similar to PIR|A84578|A84578 probable WD-40 repeat protein
[imported] - Arabidopsis thaliana, partial (80%)
Length = 1824
Score = 29.3 bits (64), Expect = 8.4
Identities = 14/44 (31%), Positives = 25/44 (56%)
Frame = +1
Query: 504 KISSSAYRLRMSTVAKDDDDSQSDTEDSGSVDDELNDSDTCSSG 547
K+S+ + + R ++ DDS+ D EDS S DD ++ + + G
Sbjct: 361 KVSNISGKRREPVPKRETDDSEMDGEDSDSEDDSEDEEEGGAGG 492
>TC87587 similar to GP|20259233|gb|AAM14332.1 putative ankyrin protein
{Arabidopsis thaliana}, partial (70%)
Length = 1641
Score = 29.3 bits (64), Expect = 8.4
Identities = 15/62 (24%), Positives = 29/62 (46%), Gaps = 4/62 (6%)
Frame = -2
Query: 764 KREALLIESLERRQTSLCRSMSRIKVSNIGM----GCMSHSDQSELDRVAEDSCSPVSDV 819
KR + ESL + T +CR+ S N+ + GC H + ++ +C+ +++
Sbjct: 311 KRSTWVRESLGSQDTKICRNSSF*SKLNLILMHIIGCQGHCNDIRCQKIRPKTCNTTANI 132
Query: 820 DN 821
N
Sbjct: 131 KN 126
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.316 0.132 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,562,090
Number of Sequences: 36976
Number of extensions: 510845
Number of successful extensions: 2777
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 2615
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2744
length of query: 1060
length of database: 9,014,727
effective HSP length: 106
effective length of query: 954
effective length of database: 5,095,271
effective search space: 4860888534
effective search space used: 4860888534
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC140023.16


