

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140022.1 + phase: 0 /pseudo
(583 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
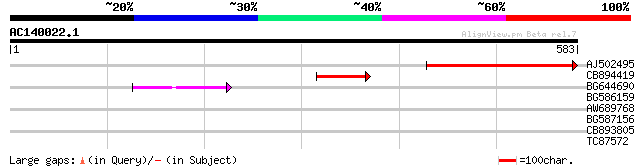
Score E
Sequences producing significant alignments: (bits) Value
AJ502495 weakly similar to GP|18071369|g putative gag-pol polypr... 262 2e-70
CB894419 weakly similar to GP|10177935|db copia-type polyprotein... 96 5e-20
BG644690 weakly similar to GP|18542179|gb putative pol protein {... 53 3e-07
BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2... 41 0.001
AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsi... 41 0.001
BG587156 similar to PIR|G85055|G8 probable polyprotein [imported... 32 0.89
CB893805 similar to GP|10177935|d copia-type polyprotein {Arabid... 29 5.7
TC87572 similar to GP|18958499|gb|AAL82617.1 elongation factor 1... 28 9.8
>AJ502495 weakly similar to GP|18071369|g putative gag-pol polyprotein {Oryza
sativa}, partial (9%)
Length = 542
Score = 262 bits (670), Expect = 2e-70
Identities = 144/155 (92%), Positives = 144/155 (92%)
Frame = +3
Query: 429 VIGQEIQKQEKARQGTHFI*EPVQYHGLRRNNLWSLFQQQK*NI**QPVVLLKQCG*EEF 488
VIGQEIQKQEKARQGT FI*EPVQYHGLRRNNLWSLF QQK NI* Q VVLLKQCG*EEF
Sbjct: 3 VIGQEIQKQEKARQGTRFI*EPVQYHGLRRNNLWSLFPQQKQNI*HQLVVLLKQCG*EEF 182
Query: 489 *K*CIMSRTLLQRYIVITSQQLH*AKIQFFMDGPSILTSGFTRYES*LLRKKW*SSTVPL 548
*K*CIMSRTLLQRYIVITSQQLH*AKIQFFMDGPSILTS FTRYES*L RKKW*SS VPL
Sbjct: 183 *K*CIMSRTLLQRYIVITSQQLH*AKIQFFMDGPSILTSSFTRYES*LPRKKW*SSIVPL 362
Query: 549 KSKLQIFLQSH*RLSHFTN*RKCLE**KLDLREAM 583
K KLQI LQSH*RLS FTN*RKCLE**KLDLREAM
Sbjct: 363 KRKLQISLQSH*RLSRFTN*RKCLE**KLDLREAM 467
>CB894419 weakly similar to GP|10177935|db copia-type polyprotein
{Arabidopsis thaliana}, partial (2%)
Length = 170
Score = 95.5 bits (236), Expect = 5e-20
Identities = 54/56 (96%), Positives = 55/56 (97%)
Frame = +1
Query: 316 RRSMQVIF*RNLRWSIQSQFPRRLKKS*S*QEKAMVKG*TQLITKV*LEV*DI*LQ 371
RRS+QVIF*RN RWSIQSQFPRRLKKS*S*QEKAMVKG*TQLITKV*LEV*DI*LQ
Sbjct: 1 RRSVQVIF*RNSRWSIQSQFPRRLKKS*S*QEKAMVKG*TQLITKV*LEV*DI*LQ 168
>BG644690 weakly similar to GP|18542179|gb putative pol protein {Zea mays},
partial (22%)
Length = 629
Score = 53.1 bits (126), Expect = 3e-07
Identities = 36/105 (34%), Positives = 56/105 (53%), Gaps = 3/105 (2%)
Frame = -3
Query: 127 LIAIKRGWWLKATNKNQVLIILKYLLLLQD*--IQFA-CLFHSQLKITGKYIKWMLSLHF 183
L+ I WW K T K + +++ LL + ++F L HS KWM +H
Sbjct: 423 LLEINPSWWCKDTIKKKE*TMMRLFHLLPEWKLLEF**LLLHSW---GSSCTKWM*RVHL 253
Query: 184 LMVLWKKKCMLSSLQDMWLEERRIKYID*RKHCMA*SRRQEHGTK 228
LM + K++C+ S+L D+ ++ +I D* +H M *S+ QEHG K
Sbjct: 252 LMEISKRRCLSSNLLDLKMQRYQIMCSD*IRHYMV*SKLQEHGMK 118
>BG586159 weakly similar to PIR|T47841|T4 hypothetical protein T2O9.150 -
Arabidopsis thaliana, partial (11%)
Length = 732
Score = 40.8 bits (94), Expect = 0.001
Identities = 35/117 (29%), Positives = 54/117 (45%), Gaps = 10/117 (8%)
Frame = +2
Query: 427 QIVIGQEIQKQEKARQGTHFI*EPVQYHGLRRNNLWSLFQQQK*NI**QPVVLLKQCG*E 486
QIVI Q I EK + VQ HG ++N W L++Q K N+ P VL+K G E
Sbjct: 374 QIVIMQVI*MIEKVLLDMYLCLVQVQSHGHLKSNRWLLYRQLKQNLLQLPFVLVKVFGCE 553
Query: 487 EF*K*CIMSRTLLQRYIVITSQQLH*AKIQFFMDG----------PSILTSGFTRYE 533
EF + + I T+QQ + + FM PS++++ +R++
Sbjct: 554 EFSRNLVTLNLEASLCIATTTQQSSFQRTRCFMGRSKHI*CSLPFPSVISASNSRFK 724
>AW689768 weakly similar to GP|10177485|d polyprotein {Arabidopsis thaliana},
partial (9%)
Length = 675
Score = 40.8 bits (94), Expect = 0.001
Identities = 40/133 (30%), Positives = 63/133 (47%), Gaps = 1/133 (0%)
Frame = +2
Query: 97 KHGS*LNYRKTRSQ*E*SGCTRQSTNPVVRLIAIKRGWWLKATNKNQVLIILKYLLLLQD 156
KHG + TR SG + L +K WL+ + K+ +I LK LL +
Sbjct: 143 KHGILFLFLHTRKLLAASGFIELKKTQMDLLTNLKPD*WLRVSVKHLDVITLKLSLL**N 322
Query: 157 *IQFACLFHSQLK-ITGKYIKWMLSLHFLMVLWKKKCMLSSLQDMWLEERRIKYID*RKH 215
+ + F L I GK+ K + ++ F MV +K+KC+ +L+D+ L + * H
Sbjct: 323 -LSLSGSFSPLLSLINGKFNKLISTMPF*MVFFKRKCICLNLRDLKL-LINLWCAS*TNH 496
Query: 216 CMA*SRRQEHGTK 228
MA*S+ HG +
Sbjct: 497 SMA*SKHHVHGMR 535
>BG587156 similar to PIR|G85055|G8 probable polyprotein [imported] -
Arabidopsis thaliana, partial (17%)
Length = 618
Score = 31.6 bits (70), Expect = 0.89
Identities = 41/134 (30%), Positives = 58/134 (42%), Gaps = 1/134 (0%)
Frame = -2
Query: 95 RIKHGS*LNYRKTRSQ*E*SGCTRQSTNPVVRLIAIKRGWWLKATNKNQVLIILKYLLLL 154
R+ HG +NY+K R Q G + ST + RL K + + I L++L
Sbjct: 482 RMIHGMRVNYQKGRRQCLVDGSLQSSTRLMGRLRGRKLD**QEGLL*HMERITLRHLHQ* 303
Query: 155 QD*IQFACLFHSQL-KITGKYIKWMLSLHFLMVLWKKKCMLSSLQDMWLEERRIKYID*R 213
Q LF + L + G KWM +HF + K + Q + + R Y *R
Sbjct: 302 PSYTQLE-LF*AWL*TLDGDCGKWM*RMHFYKENLRMKSICILHQVLNI**REGMY*G*R 126
Query: 214 KHCMA*SRRQEHGT 227
+ M *S QEHGT
Sbjct: 125 RLSMG*SNHQEHGT 84
>CB893805 similar to GP|10177935|d copia-type polyprotein {Arabidopsis
thaliana}, partial (14%)
Length = 778
Score = 28.9 bits (63), Expect = 5.7
Identities = 54/204 (26%), Positives = 78/204 (37%), Gaps = 20/204 (9%)
Frame = +1
Query: 92 QLRRIKHGS*LNYRKTRSQ*E*SGCTRQSTNPVVRLIAIKRGWWLKATNKNQVLIILKYL 151
Q + I GS* Y + + + +G +QS + RL +IK W + T N VL K+L
Sbjct: 109 QQKEIILGS*QTYDQVQKPLDSNGSLKQS*MRMERLRSIKLDLWPRGTLNNMVLTTPKFL 288
Query: 152 LLLQD*IQFACLFHSQLKITGKYIKWMLSLHFLMVLWKKKCMLSSLQDMWLEERRIKYID 211
H L G +W L L L L +C+ + + L R I+
Sbjct: 289 -------------HQWLD--GTRSEW*LLL--LPKLKGMECVSARCKKRILVWR----IE 405
Query: 212 *RKHC--------------------MA*SRRQEHGTKRLILILFKTVFRDVHSSTHSTSN 251
* C MA ++ EHGT LI + R VH +T +
Sbjct: 406 *GSFC*STTGLCEEG**AEGLKGLCMALNKHHEHGTAASKLISPRKDSRSVHMNTPCL*S 585
Query: 252 SLILEMFLLCASMSMI*YSPVTIQ 275
+ F L M MI*Y V ++
Sbjct: 586 *VKEVKFSLLVFMLMI*YLSVMMR 657
>TC87572 similar to GP|18958499|gb|AAL82617.1 elongation factor 1-gamma
{Glycine max}, partial (55%)
Length = 743
Score = 28.1 bits (61), Expect = 9.8
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 7/75 (9%)
Frame = +3
Query: 148 LKYLLLLQD*IQFACLFHSQLKITGKYIKWMLSLHFLMVLWKKKCMLSSL--QDM----- 200
+++L +L D ++ CL H L I+ L+ + +LW+ L S QD+
Sbjct: 228 MRWLAMLLDWVKATCLDHLPL------IRPTLNNGLIFLLWRLMTTL*SCIGQDVDFPPT 389
Query: 201 WLEERRIKYID*RKH 215
WL+ RR+++ *R+H
Sbjct: 390 WLQ*RRLQFPH*REH 434
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.369 0.164 0.579
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,760,092
Number of Sequences: 36976
Number of extensions: 251399
Number of successful extensions: 3378
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1049
Number of HSP's successfully gapped in prelim test: 142
Number of HSP's that attempted gapping in prelim test: 2288
Number of HSP's gapped (non-prelim): 1252
length of query: 583
length of database: 9,014,727
effective HSP length: 101
effective length of query: 482
effective length of database: 5,280,151
effective search space: 2545032782
effective search space used: 2545032782
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC140022.1


