

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
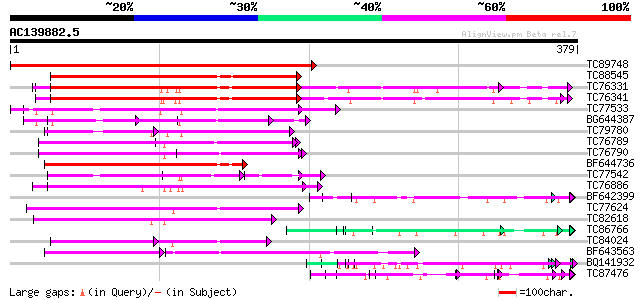
Query= AC139882.5 - phase: 0
(379 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC89748 homologue to PIR|B84565|B84565 probable spliceosome asso... 412 e-116
TC88545 similar to GP|13560783|gb|AAK30205.1 poly(A)-binding pro... 121 5e-28
TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding pro... 120 7e-28
TC76341 similar to GP|7528270|gb|AAF63202.1| poly(A)-binding pro... 120 7e-28
TC77533 similar to GP|6996560|emb|CAB75429.1 oligouridylate bind... 112 3e-25
BG644387 homologue to GP|6996560|emb oligouridylate binding prot... 101 5e-22
TC79780 similar to GP|18377731|gb|AAL67015.1 putative DNA bindin... 99 3e-21
TC76789 similar to GP|15294254|gb|AAK95304.1 AT4g24770/F22K18_30... 95 5e-20
TC76790 similar to PIR|T06817|T06817 RNA-binding protein - garde... 92 3e-19
BF644736 similar to GP|7528270|gb| poly(A)-binding protein {Cucu... 91 1e-18
TC77542 similar to GP|6996560|emb|CAB75429.1 oligouridylate bind... 77 4e-17
TC76886 similar to PIR|T01932|T01932 RNA binding protein homolog... 84 1e-16
BF642399 similar to PIR|T10863|T10 extensin precursor - kidney b... 83 2e-16
TC77624 similar to PIR|T12196|T12196 RNA-binding protein - fava ... 82 3e-16
TC82618 similar to GP|9294614|dbj|BAB02953.1 DNA/RNA binding pro... 79 2e-15
TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E... 79 2e-15
TC84024 similar to PIR|S77714|S77714 RNA-binding protein precurs... 79 4e-15
BF643563 similar to GP|13877739|gb unknown protein {Arabidopsis ... 78 5e-15
BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich gl... 78 6e-15
TC87476 homologue to PIR|T11622|T11622 extensin class 1 precurso... 70 2e-14
>TC89748 homologue to PIR|B84565|B84565 probable spliceosome associated
protein [imported] - Arabidopsis thaliana, partial (56%)
Length = 707
Score = 412 bits (1060), Expect = e-116
Identities = 204/205 (99%), Positives = 204/205 (99%)
Frame = +2
Query: 1 MTTRIAPGVGANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDR 60
MTTRIAPGVGANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDR
Sbjct: 92 MTTRIAPGVGANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDR 271
Query: 61 VTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGANLFIGNLD 120
VTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGANLFIGNLD
Sbjct: 272 VTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGANLFIGNLD 451
Query: 121 PDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLC 180
PDVDEKLLYDTFSAFGVIVTNPKIMRDPDT NSRGFGFISYDSFEASDSAIEAMNGQYLC
Sbjct: 452 PDVDEKLLYDTFSAFGVIVTNPKIMRDPDTRNSRGFGFISYDSFEASDSAIEAMNGQYLC 631
Query: 181 NRQITVSYAYKKDTKGERHGTPAER 205
NRQITVSYAYKKDTKGERHGTPAER
Sbjct: 632 NRQITVSYAYKKDTKGERHGTPAER 706
>TC88545 similar to GP|13560783|gb|AAK30205.1 poly(A)-binding protein
{Daucus carota}, partial (35%)
Length = 866
Score = 121 bits (303), Expect = 5e-28
Identities = 68/169 (40%), Positives = 105/169 (61%), Gaps = 1/169 (0%)
Frame = +2
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YVG+LD +++ L++LF Q G VV+V + +D + Q GYG+V F + DA A+ VLN
Sbjct: 188 YVGDLDHDVTDSQLYDLFNQIGQVVSVRICRDLASQQSLGYGYVNFSNPHDAAKAMDVLN 367
Query: 88 MIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMR 146
L KPIR+ + +D G AN+FI NLD +D K LYDTFS FG I+ + KI
Sbjct: 368 FTPLNNKPIRIMYSHRDPSVRKSGAANIFIKNLDRAIDHKALYDTFSIFGNIL-SCKIAM 544
Query: 147 DPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTK 195
D +G S+G+GF+ +++ E++ SAI+ +NG L ++ + V + +K +
Sbjct: 545 DA-SGLSKGYGFVQFENEESAQSAIDKLNGMLLNDKPVYVGHFQRKQDR 688
>TC76331 similar to GP|7673355|gb|AAF66823.1| poly(A)-binding protein
{Nicotiana tabacum}, partial (97%)
Length = 2581
Score = 120 bits (302), Expect = 7e-28
Identities = 67/169 (39%), Positives = 105/169 (61%), Gaps = 1/169 (0%)
Frame = +3
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YVG+LD +++ L++LF Q G VV+V V +D T + GYG+V + + +DA A+ VLN
Sbjct: 453 YVGDLDMNVTDSQLYDLFNQLGQVVSVRVCRDLTTRRSLGYGYVNYSNPQDAARALDVLN 632
Query: 88 MIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMR 146
L +PIR+ + +D G N+FI NLD +D K L+DTFS+FG I+ + K+
Sbjct: 633 FTPLNNRPIRIMYSHRDPSIRKSGQGNIFIKNLDKAIDHKALHDTFSSFGNIL-SCKVAV 809
Query: 147 DPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTK 195
D +G S+G+GF+ +D+ EA+ AIE +NG L ++Q+ V +K +
Sbjct: 810 D-GSGQSKGYGFVQFDTEEAAQKAIEKLNGMLLNDKQVYVGPFLRKQER 953
Score = 102 bits (254), Expect = 2e-22
Identities = 93/341 (27%), Positives = 159/341 (46%), Gaps = 26/341 (7%)
Frame = +3
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRS 75
+ + +R + +V NL +++ L + F + G + + V +D + + +GFV F S
Sbjct: 954 ESTGDRAKFNNVFVKNLSESTTDDELKKTFGEFGTITSAVVMRDG-DGKSKCFGFVNFES 1130
Query: 76 EEDADYAIKVLNMIKLYGKPIRVNKASQD---------------KKSLDV--GANLFIGN 118
+DA A++ LN K+ K V KA + K++ D GANL++ N
Sbjct: 1131 TDDAARAVEALNGKKIDDKEWYVGKAQKKSEREHELKIKFEQSMKEAADKYQGANLYVKN 1310
Query: 119 LDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQY 178
LD + ++ L + FS++G I T+ K+MRDP+ G SRG GF+++ + E + A+ MNG+
Sbjct: 1311 LDDSIADEKLKELFSSYGTI-TSCKVMRDPN-GVSRGSGFVAFSTPEEASRALLEMNGKM 1484
Query: 179 LCNRQITVSYAYKKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANG 238
+ ++ + V+ A +K+ + R ++ S P + R ++ G P +
Sbjct: 1485 VASKPLYVTLAQRKEDRRARLQAQFAQMRPVSMPPSVAPR-MPMYPPGGPGMGQQIFYGQ 1661
Query: 239 TIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAF------QPMQMPPPGQQVW---HQQQQ 289
PA +P +P G + ++P P F Q Q PG + Q QQ
Sbjct: 1662 GPPAIIPSQP-GFGYQQQLVPGMRPGGAPVPNFFVPMVQQGQQGQRPGGRRGGGVQQSQQ 1838
Query: 290 GQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLP 330
P+M Q M P + +R P MP P P G+ G +P
Sbjct: 1839 PVPLMPQQMLPRGRVYRYPPGRGMPDGPMP-GVAGGMYSVP 1958
Score = 94.0 bits (232), Expect = 9e-20
Identities = 105/386 (27%), Positives = 162/386 (41%), Gaps = 27/386 (6%)
Frame = +3
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
S ++ ++ NLD I + L + F G +++ V D + Q +GYGFV+F +EE
Sbjct: 687 SIRKSGQGNIFIKNLDKAIDHKALHDTFSSFGNILSCKVAVDG-SGQSKGYGFVQFDTEE 863
Query: 78 DADYAIKVLNMIKLYGKPIRVNK--ASQDKKSLDVGA---NLFIGNLDPDVDEKLLYDTF 132
A AI+ LN + L K + V Q+++S A N+F+ NL + L TF
Sbjct: 864 AAQKAIEKLNGMLLNDKQVYVGPFLRKQERESTGDRAKFNNVFVKNLSESTTDDELKKTF 1043
Query: 133 SAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
FG I T+ +MRD D G S+ FGF++++S + + A+EA+NG+ + +++ V A K
Sbjct: 1044 GEFGTI-TSAVVMRDGD-GKSKCFGFVNFESTDDAARAVEALNGKKIDDKEWYVGKAQK- 1214
Query: 193 DTKGERHGTPAERVLAASNPTAQKSRPHTLFAS------GPPSLPNAPQANGTIPAPVPP 246
K ER + + A K + L+ L + GTI +
Sbjct: 1215 --KSEREHELKIKFEQSMKEAADKYQGANLYVKNLDDSIADEKLKELFSSYGTITSCKVM 1388
Query: 247 RPFANGVAPPPIHVIQPPPPQAP-----------AFQPMQMPPPGQQVWHQQQQGQPMMQ 295
R NGV+ V P +A A +P+ + Q+ ++ + Q
Sbjct: 1389 RD-PNGVSRGSGFVAFSTPEEASRALLEMNGKMVASKPLYV-TLAQRKEDRRARLQAQFA 1562
Query: 296 QGMPPPMQQFRPPHSMQMPPPPPPQGM-----QGPPRPLPPPSVMAGQQQVWRPPPPPQQ 350
Q P M P PP P G QGPP +P QQQ+ P +
Sbjct: 1563 QMRPVSMPPSVAPRMPMYPPGGPGMGQQIFYGQGPPAIIPSQPGFGYQQQL----VPGMR 1730
Query: 351 QGGHPMGYPQNSMPPPPPPNHHGMRP 376
GG P+ N P G RP
Sbjct: 1731 PGGAPV---PNFFVPMVQQGQQGQRP 1799
>TC76341 similar to GP|7528270|gb|AAF63202.1| poly(A)-binding protein
{Cucumis sativus}, partial (90%)
Length = 2501
Score = 120 bits (302), Expect = 7e-28
Identities = 65/169 (38%), Positives = 108/169 (63%), Gaps = 1/169 (0%)
Frame = +3
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YVG+L+ +++ L++LF Q G VV+V V +D T + GYG+V F + +DA A+ VLN
Sbjct: 303 YVGDLEVNVNDSQLYDLFNQVGQVVSVRVCRDLATRRSLGYGYVNFTNPQDAARALDVLN 482
Query: 88 MIKLYGKPIRVNKASQDKKSLDVG-ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMR 146
+ K IRV + +D S G AN+FI NLD +D K L+DTFS+FG I+ + KI
Sbjct: 483 FTPMNNKSIRVMYSHRDPSSRKSGTANIFIKNLDKTIDHKALHDTFSSFGQIM-SCKIAT 659
Query: 147 DPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTK 195
D +G S+G+GF+ +++ +++ +AI+ +NG + ++Q+ V + +K +
Sbjct: 660 D-GSGQSKGYGFVQFEAEDSAQNAIDKLNGMLINDKQVFVGHFLRKQDR 803
Score = 107 bits (266), Expect = 1e-23
Identities = 109/380 (28%), Positives = 161/380 (41%), Gaps = 36/380 (9%)
Frame = +3
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YV NL +E+ L F G + + + +D + + +GFV F + EDA A++ LN
Sbjct: 840 YVKNLSESFTEDDLKNEFGAYGTITSAVLMRD-ADGRSKCFGFVNFENAEDAAKAVEALN 1016
Query: 88 MIKLYGKPIRVNKA---SQDKKSLD---------------VGANLFIGNLDPDVDEKLLY 129
K+ K V KA S+ ++ L G NL++ NLD + ++ L
Sbjct: 1017 GKKVDDKEWYVGKAQKKSEREQELKGRFEQTVKESVVDKFQGLNLYLKNLDDSITDEKLK 1196
Query: 130 DTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYA 189
+ FS FG I T+ KIMRDP+ G SRG GF+++ + E + A+ MNG+ + ++ + V+ A
Sbjct: 1197 EMFSEFGTI-TSYKIMRDPN-GVSRGSGFVAFSTPEEASRALGEMNGKMIVSKPLYVAVA 1370
Query: 190 YKKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPF 249
+K+ + R + RP + S P +P P P + F
Sbjct: 1371 QRKEDRRAR-----------LQAQFSQMRPVAITPSVAPRMPLYPPG-----TPGLGQQF 1502
Query: 250 ANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPH 309
G PP + MPP Q + QQQ P M+ G PM + P
Sbjct: 1503 MYGQGPPAM-----------------MPP--QAGFGYQQQLVPGMRPG-GGPMPSYFVPM 1622
Query: 310 SMQMPPPPPPQGMQGPPRPLPPPSV------MAGQQQVWRPPP--------PPQQQGGHP 355
Q P G +G P PP V M +Q+V+R PP P Q GG
Sbjct: 1623 VQQGQQGQRPGGRRGGPGQQPPQQVPMMQQQMLPRQRVYRYPPGRSNIQDAPVQNIGGGM 1802
Query: 356 MGYPQNSMP----PPPPPNH 371
M Y +P PP P H
Sbjct: 1803 MSYDMGGLPLRDVVPPMPIH 1862
Score = 89.4 bits (220), Expect = 2e-18
Identities = 99/386 (25%), Positives = 164/386 (41%), Gaps = 27/386 (6%)
Frame = +3
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEE 77
S+ ++ A ++ NLD I + L + F G +++ + D + Q +GYGFV+F +E+
Sbjct: 537 SSRKSGTANIFIKNLDKTIDHKALHDTFSSFGQIMSCKIATDG-SGQSKGYGFVQFEAED 713
Query: 78 DADYAIKVLNMIKLYGKPIRVNK--ASQDKKSLDVGA---NLFIGNLDPDVDEKLLYDTF 132
A AI LN + + K + V QD+ ++ N+++ NL E L + F
Sbjct: 714 SAQNAIDKLNGMLINDKQVFVGHFLRKQDRDNVLSKTKFNNVYVKNLSESFTEDDLKNEF 893
Query: 133 SAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
A+G I T+ +MRD D G S+ FGF+++++ E + A+EA+NG+ + +++ V A KK
Sbjct: 894 GAYGTI-TSAVLMRDAD-GRSKCFGFVNFENAEDAAKAVEALNGKKVDDKEWYVGKAQKK 1067
Query: 193 DTKGERHGTPAERVLAASNPTAQKSRPHTLFASG------PPSLPNAPQANGTIPAPVPP 246
+ + E+ + S K + L+ L GTI +
Sbjct: 1068SEREQELKGRFEQTVKES--VVDKFQGLNLYLKNLDDSITDEKLKEMFSEFGTITSYKIM 1241
Query: 247 RPFANGVAPPPIHVIQPPPPQAP-----------AFQPMQMPPPGQQVWHQQQQGQPMMQ 295
R NGV+ V P +A +P+ + Q+ ++ + Q
Sbjct: 1242RD-PNGVSRGSGFVAFSTPEEASRALGEMNGKMIVSKPLYV-AVAQRKEDRRARLQAQFS 1415
Query: 296 QGMPPPMQQFRPPHSMQMPPPPPPQGM-----QGPPRPLPPPSVMAGQQQVWRPPPPPQQ 350
Q P + P PP P G QGPP +PP + QQQ+ P +
Sbjct: 1416QMRPVAITPSVAPRMPLYPPGTPGLGQQFMYGQGPPAMMPPQAGFGYQQQL----VPGMR 1583
Query: 351 QGGHPMGYPQNSMPPPPPPNHHGMRP 376
GG PM + P G RP
Sbjct: 1584PGGGPM---PSYFVPMVQQGQQGQRP 1652
>TC77533 similar to GP|6996560|emb|CAB75429.1 oligouridylate binding protein
{Nicotiana plumbaginifolia}, partial (79%)
Length = 1621
Score = 112 bits (279), Expect = 3e-25
Identities = 71/224 (31%), Positives = 124/224 (54%), Gaps = 3/224 (1%)
Frame = +1
Query: 1 MTTRIAPGVGANLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDR 60
+T +I P + NL + + + YVGN+ PQ+SE LL ELF AG + +
Sbjct: 154 ITPQIEPILSGNL--PPGFDSSTCRSVYVGNIHPQVSEPLLQELFSSAGALEGCKL---- 315
Query: 61 VTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSLDVGAN--LFIGN 118
+ + YGFV++ A AI LN ++G+ I+VN A + D + +F+G+
Sbjct: 316 IRKEKSSYGFVDYFDRSSAAIAIVTLNGRNIFGQSIKVNWAYTRGQREDTSGHFHIFVGD 495
Query: 119 LDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQY 178
L P+V + LY FSA+ ++ ++M D TG SRGFGF+S+ + + + SAI + G++
Sbjct: 496 LSPEVTDATLYACFSAYSSC-SDARVMWDQKTGRSRGFGFVSFRNQQEAQSAINDLTGKW 672
Query: 179 LCNRQITVSYAYK-KDTKGERHGTPAERVLAASNPTAQKSRPHT 221
L +RQI ++A K + GE + ++ V+ ++ T+++++ T
Sbjct: 673 LGSRQIRCNWATKGANMNGENQSSESKSVVELTSGTSEEAQEMT 804
Score = 64.7 bits (156), Expect = 6e-11
Identities = 55/233 (23%), Positives = 103/233 (43%), Gaps = 46/233 (19%)
Frame = +1
Query: 10 GANLLGQ-------HSAERNQDATA----YVGNLDPQISEELLWELFVQAGPVVNVYVPK 58
G N+ GQ ++ + +D + +VG+L P++++ L+ F + V
Sbjct: 397 GRNIFGQSIKVNWAYTRGQREDTSGHFHIFVGDLSPEVTDATLYACFSAYSSCSDARVMW 576
Query: 59 DRVTNQHQGYGFVEFRSEEDADYAIKVLNMIKLYGKPIRVN------------KASQDKK 106
D+ T + +G+GFV FR++++A AI L L + IR N ++S+ K
Sbjct: 577 DQKTGRSRGFGFVSFRNQQEAQSAINDLTGKWLGSRQIRCNWATKGANMNGENQSSESKS 756
Query: 107 SLDVGAN----------------------LFIGNLDPDVDEKLLYDTFSAFGV-IVTNPK 143
+++ + +++GNL P+V L+ F A GV + + +
Sbjct: 757 VVELTSGTSEEAQEMTSDDSPEKNPQYTTVYVGNLAPEVTSVDLHHHFHALGVGTIEDVR 936
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKG 196
+ RD +GFGF+ Y + + AI+ N ++L + I S+ K G
Sbjct: 937 VQRD------KGFGFVRYSTHGEAALAIQMGNTRFLFGKPIKCSWGSKPTPPG 1077
>BG644387 homologue to GP|6996560|emb oligouridylate binding protein
{Nicotiana plumbaginifolia}, partial (53%)
Length = 717
Score = 101 bits (251), Expect = 5e-22
Identities = 57/153 (37%), Positives = 88/153 (57%), Gaps = 2/153 (1%)
Frame = +2
Query: 26 TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKV 85
+ YVGN+ PQ++E LL E+F GP+ + K + YGFV++ A AI
Sbjct: 122 SVYVGNIHPQVTEPLLQEVFSSTGPLEGCKLIK----KEKSSYGFVDYFDRRSAALAIVT 289
Query: 86 LNMIKLYGKPIRVNKASQDKKSLDVGA--NLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
LN L+G+PI+VN A + D + N+F+G+L P+V + LY FS + ++ K
Sbjct: 290 LNGRNLFGQPIKVNWAYTSAQREDTSSHFNIFVGDLSPEVTDATLYACFSVYPSC-SDAK 466
Query: 144 IMRDPDTGNSRGFGFISYDSFEASDSAIEAMNG 176
+M D +G SRGFGF+S+ + + + SAI +NG
Sbjct: 467 VMWDQKSGRSRGFGFVSFRNQQEAQSAINDLNG 565
Score = 43.5 bits (101), Expect = 1e-04
Identities = 26/89 (29%), Positives = 47/89 (52%)
Frame = +2
Query: 113 NLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIE 172
++++GN+ P V E LL + FS+ G + I ++ + +GF+ Y ++ AI
Sbjct: 122 SVYVGNIHPQVTEPLLQEVFSSTGPLEGCKLIKKEKSS-----YGFVDYFDRRSAALAIV 286
Query: 173 AMNGQYLCNRQITVSYAYKKDTKGERHGT 201
+NG+ L + I V++AY T +R T
Sbjct: 287 TLNGRNLFGQPIKVNWAY---TSAQREDT 364
Score = 43.5 bits (101), Expect = 1e-04
Identities = 25/89 (28%), Positives = 49/89 (54%), Gaps = 11/89 (12%)
Frame = +2
Query: 10 GANLLGQ-------HSAERNQDATA----YVGNLDPQISEELLWELFVQAGPVVNVYVPK 58
G NL GQ +++ + +D ++ +VG+L P++++ L+ F + V
Sbjct: 296 GRNLFGQPIKVNWAYTSAQREDTSSHFNIFVGDLSPEVTDATLYACFSVYPSCSDAKVMW 475
Query: 59 DRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
D+ + + +G+GFV FR++++A AI LN
Sbjct: 476 DQKSGRSRGFGFVSFRNQQEAQSAINDLN 562
>TC79780 similar to GP|18377731|gb|AAL67015.1 putative DNA binding protein
ACBF {Arabidopsis thaliana}, partial (67%)
Length = 1649
Score = 99.0 bits (245), Expect = 3e-21
Identities = 54/171 (31%), Positives = 94/171 (54%), Gaps = 6/171 (3%)
Frame = +3
Query: 26 TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKV 85
T ++G+L + E L F G V+++ + ++++T Q +GYGF+EF S A+ ++
Sbjct: 102 TLWIGDLQYWVDENYLTHCFSHTGEVISIKIIRNKITGQPEGYGFIEFVSHSAAERVLQT 281
Query: 86 LNMIKLYG--KPIRVNKASQD--KKSLDVGAN--LFIGNLDPDVDEKLLYDTFSAFGVIV 139
N ++ G + R+N AS ++ D G + +F+G+L PDV + LL +TF V
Sbjct: 282 YNGTQMPGTEQTFRLNWASFGIGERRPDAGPDHSIFVGDLAPDVTDYLLQETFRTHYGSV 461
Query: 140 TNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAY 190
K++ DP+TG S+G+GF+ + + A+ MNG Y R + +S Y
Sbjct: 462 RGAKVVTDPNTGRSKGYGFVKFSDESERNRAMSEMNGVYCSTRPMRISCCY 614
Score = 41.6 bits (96), Expect = 5e-04
Identities = 28/76 (36%), Positives = 43/76 (55%)
Frame = +2
Query: 24 DATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAI 83
+ T YVGNLD +SEE L + F+Q G +V+V V + + GFV+F + A+ AI
Sbjct: 755 NTTIYVGNLDLNVSEEELKQNFLQFGEIVSVKV------HPGKACGFVQFGARASAEEAI 916
Query: 84 KVLNMIKLYGKPIRVN 99
+ + L + IRV+
Sbjct: 917 QKMQGKILGQQVIRVS 964
>TC76789 similar to GP|15294254|gb|AAK95304.1 AT4g24770/F22K18_30
{Arabidopsis thaliana}, partial (62%)
Length = 1294
Score = 94.7 bits (234), Expect = 5e-20
Identities = 58/180 (32%), Positives = 98/180 (54%), Gaps = 7/180 (3%)
Frame = +3
Query: 20 ERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDA 79
E ++D +VGNL + E L +LF Q+G V V +R T++ +G+GFV + E+
Sbjct: 438 EPSEDLKIFVGNLPFDVDSEKLAQLFEQSGTVEIAEVIYNRDTDRSRGFGFVTMSTSEEV 617
Query: 80 DYAIKVLNMIKLYGKPIRVNKAS-------QDKKSLDVGANLFIGNLDPDVDEKLLYDTF 132
+ A+ + +L G+ + VNKA+ + ++ + G ++GNL DVD L F
Sbjct: 618 ERAVNKFSGFELDGRLLTVNKAAPRGTPRLRQPRTFNSGLRAYVGNLPWDVDNSSLEQLF 797
Query: 133 SAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKK 192
S G V + +++ D +TG SRGFGF++ + + AI A++GQ R I V+ A ++
Sbjct: 798 SEHGK-VESAQVVYDRETGRSRGFGFVTMSNEAEMNDAIAALDGQSFNGRAIRVNVAEER 974
Score = 49.7 bits (117), Expect = 2e-06
Identities = 32/97 (32%), Positives = 52/97 (52%)
Frame = +3
Query: 98 VNKASQDKKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFG 157
V AS ++ S D+ +F+GNL DVD + L F G + +++ + DT SRGFG
Sbjct: 417 VEVASFEEPSEDL--KIFVGNLPFDVDSEKLAQLFEQSGTVEI-AEVIYNRDTDRSRGFG 587
Query: 158 FISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDT 194
F++ + E + A+ +G L R +TV+ A + T
Sbjct: 588 FVTMSTSEEVERAVNKFSGFELDGRLLTVNKAAPRGT 698
>TC76790 similar to PIR|T06817|T06817 RNA-binding protein - garden pea,
partial (79%)
Length = 1141
Score = 92.0 bits (227), Expect = 3e-19
Identities = 62/187 (33%), Positives = 96/187 (51%), Gaps = 10/187 (5%)
Frame = +1
Query: 20 ERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDA 79
E +DA +VGN + E L LF QAG V V +R T+ +G+GFV + E+A
Sbjct: 346 EPPEDAKLFVGNFPFDVDSEKLAMLFGQAGTVEIAEVIYNRQTDLSRGFGFVTMSTVEEA 525
Query: 80 DYAIKVLNMIKLYGKPIRVNKAS----------QDKKSLDVGANLFIGNLDPDVDEKLLY 129
+ A++ N G+ + VNKAS + ++ + +++ NL +VD L
Sbjct: 526 ESAVEKFNGYDYNGRSLVVNKASPKGSRPERTERAPRTFEPVLRIYVANLAWEVDNSRLE 705
Query: 130 DTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYA 189
FS G IV + +++ D +TG SRGFGF++ + AI A++GQ L R I VS A
Sbjct: 706 QVFSEHGKIV-SARVVYDRETGRSRGFGFVTMSDETEMNDAIAALDGQSLEGRTIRVSVA 882
Query: 190 YKKDTKG 196
+ +G
Sbjct: 883 EDRPRRG 903
Score = 52.4 bits (124), Expect = 3e-07
Identities = 30/87 (34%), Positives = 47/87 (53%)
Frame = +1
Query: 112 ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAI 171
A LF+GN DVD + L F G + +++ + T SRGFGF++ + E ++SA+
Sbjct: 361 AKLFVGNFPFDVDSEKLAMLFGQAGTVEI-AEVIYNRQTDLSRGFGFVTMSTVEEAESAV 537
Query: 172 EAMNGQYLCNRQITVSYAYKKDTKGER 198
E NG R + V+ A K ++ ER
Sbjct: 538 EKFNGYDYNGRSLVVNKASPKGSRPER 618
>BF644736 similar to GP|7528270|gb| poly(A)-binding protein {Cucumis
sativus}, partial (20%)
Length = 674
Score = 90.5 bits (223), Expect = 1e-18
Identities = 51/137 (37%), Positives = 83/137 (60%), Gaps = 1/137 (0%)
Frame = +1
Query: 24 DATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAI 83
+A+ YVG+L+ ++E L++LF Q VV++ V +D+ GY +V F + +DA A
Sbjct: 265 NASLYVGDLEGSVNEGQLFDLFSQVAQVVSIRVCRDQTRRSSLGYAYVNFTNAQDAANAK 444
Query: 84 KVLNMIKLYGKPIRVNKASQDKKSLDVGA-NLFIGNLDPDVDEKLLYDTFSAFGVIVTNP 142
++LN L KPIR+ + +D G+ N+FI NLD +D K L+DTF+AFG ++ +
Sbjct: 445 ELLNFTPLNEKPIRIMFSQRDPSVRKSGSGNVFIKNLDTSIDHKALHDTFAAFGPVL-SC 621
Query: 143 KIMRDPDTGNSRGFGFI 159
K+ D G S+ +GF+
Sbjct: 622 KVALD-GNGQSKXYGFV 669
Score = 32.3 bits (72), Expect = 0.31
Identities = 17/55 (30%), Positives = 27/55 (48%)
Frame = +1
Query: 18 SAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVE 72
S ++ ++ NLD I + L + F GPV++ V D Q + YGFV+
Sbjct: 511 SVRKSGSGNVFIKNLDTSIDHKALHDTFAAFGPVLSCKVALDG-NGQSKXYGFVQ 672
>TC77542 similar to GP|6996560|emb|CAB75429.1 oligouridylate binding protein
{Nicotiana plumbaginifolia}, partial (74%)
Length = 1840
Score = 77.0 bits (188), Expect(2) = 4e-17
Identities = 42/134 (31%), Positives = 74/134 (54%), Gaps = 2/134 (1%)
Frame = +2
Query: 26 TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKV 85
+ YVGN+ ++++LL E+F AGP+ + + + YGFV++ A AI
Sbjct: 380 SVYVGNIHVNVTDKLLAEVFQSAGPLAGCKL----IRKEKSSYGFVDYHDRASAALAIMT 547
Query: 86 LNMIKLYGKPIRVNKASQDKKSLDVGA--NLFIGNLDPDVDEKLLYDTFSAFGVIVTNPK 143
L+ +LYG+ ++VN A + D N+F+G+L P+V + L+ FS + ++ +
Sbjct: 548 LHGRQLYGQALKVNWAYANSSREDTSGHFNVFVGDLSPEVTDATLFACFSVY-TTCSDAR 724
Query: 144 IMRDPDTGNSRGFG 157
+M D TG S+G+G
Sbjct: 725 VMWDHKTGRSKGYG 766
Score = 40.8 bits (94), Expect = 9e-04
Identities = 26/100 (26%), Positives = 47/100 (47%), Gaps = 6/100 (6%)
Frame = +1
Query: 103 QDKKSLDVGAN------LFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGF 156
QD + D N +++GNL DV + L+ F A G ++ + + +GF
Sbjct: 955 QDNSNEDAPENNPSYTTVYVGNLPHDVTQAELHCQFHALGA-----GVLEEVRVQSGKGF 1119
Query: 157 GFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKG 196
GF+ Y++ E + AI+ NG+ + + + S+ K G
Sbjct: 1120 GFVRYNTHEEAAMAIQMANGRPVRGKTMKCSWGSKPTPPG 1239
Score = 40.0 bits (92), Expect = 0.001
Identities = 20/88 (22%), Positives = 47/88 (52%)
Frame = +2
Query: 113 NLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIE 172
++++GN+ +V +KLL + F + G + I ++ + +GF+ Y ++ AI
Sbjct: 380 SVYVGNIHVNVTDKLLAEVFQSAGPLAGCKLIRKEKSS-----YGFVDYHDRASAALAIM 544
Query: 173 AMNGQYLCNRQITVSYAYKKDTKGERHG 200
++G+ L + + V++AY ++ + G
Sbjct: 545 TLHGRQLYGQALKVNWAYANSSREDTSG 628
Score = 28.5 bits (62), Expect(2) = 4e-17
Identities = 20/54 (37%), Positives = 30/54 (55%)
Frame = +1
Query: 158 FISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERHGTPAERVLAASN 211
F+S D +A SAI M G++L NRQI ++A TKG + E++ + N
Sbjct: 772 FLSRDHQDAQ-SAINDMTGKWLGNRQIRCNWA----TKGAGGSSNEEKINDSQN 918
>TC76886 similar to PIR|T01932|T01932 RNA binding protein homolog - common
tobacco (fragment), partial (67%)
Length = 1896
Score = 83.6 bits (205), Expect = 1e-16
Identities = 55/195 (28%), Positives = 99/195 (50%), Gaps = 11/195 (5%)
Frame = +2
Query: 26 TAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKV 85
T ++G+L + E L F G V + V +++ T Q +GYGFVEF + A+ ++
Sbjct: 551 TIWLGDLHHWMDETFLHNCFAHTGEVASAKVIRNKQTGQSEGYGFVEFYTRAMAEKVLQN 730
Query: 86 LN--MIKLYGKPIRVNKAS-------QDKKSLDVGANL--FIGNLDPDVDEKLLYDTFSA 134
N M+ + R+N A+ +++S + ++L F+G+L DV + +L +TF++
Sbjct: 731 FNGTMMPNTDQAFRLNWATFSAAGGGGERRSSEATSDLSVFVGDLAIDVTDAMLQETFAS 910
Query: 135 FGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVSYAYKKDT 194
+ K++ D +TG S+G+GF+ + A+ MNG Y +R + V A K T
Sbjct: 911 KFSSIKGAKVVIDSNTGRSKGYGFVRFGDESERTRAMTEMNGVYCSSRPMRVGVATPKKT 1090
Query: 195 KGERHGTPAERVLAA 209
G ++ V+ A
Sbjct: 1091YGNPQQYSSQAVVLA 1135
Score = 56.6 bits (135), Expect = 2e-08
Identities = 48/217 (22%), Positives = 90/217 (41%), Gaps = 33/217 (15%)
Frame = +2
Query: 16 QHSAERNQDATAYVGNLDPQISEELLWELFVQA-GPVVNVYVPKDRVTNQHQGYGFVEFR 74
+ S+E D + +VG+L +++ +L E F + V D T + +GYGFV F
Sbjct: 815 RRSSEATSDLSVFVGDLAIDVTDAMLQETFASKFSSIKGAKVVIDSNTGRSKGYGFVRFG 994
Query: 75 SEEDADYAIKVLNMIKLYGKPIRVNKASQDKK---------------------------- 106
E + A+ +N + +P+RV A+ K
Sbjct: 995 DESERTRAMTEMNGVYCSSRPMRVGVATPKKTYGNPQQYSSQAVVLAGGGGHSNGAMAQG 1174
Query: 107 SLDVG----ANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYD 162
S G +F+G LD D+ ++ L F FG +++ ++ P +G GF+
Sbjct: 1175 SQSEGDSNNTTIFVGGLDSDISDEDLRQPFLQFGDVIS----VKIPV---GKGCGFVQLA 1333
Query: 163 SFEASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERH 199
+ ++ AI+ +NG + + + +S+ K R+
Sbjct: 1334 DRKNAEEAIQGLNGTVIGKQTVRLSWGRSPGNKHWRN 1444
Score = 32.3 bits (72), Expect = 0.31
Identities = 21/66 (31%), Positives = 25/66 (37%), Gaps = 3/66 (4%)
Frame = +2
Query: 262 QPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMP---PPMQQFRPPHSMQMPPPPP 318
Q P + MQ P + QQ P Q MP P Q +PPH Q+ PPP
Sbjct: 296 QQPHQYQQQWMSMQYPATAMAMMQQQMMMYP--QHYMPYVHPHHYQQQPPHQQQVSPPPS 469
Query: 319 PQGMQG 324
G
Sbjct: 470 AHSHHG 487
Score = 30.8 bits (68), Expect = 0.90
Identities = 27/86 (31%), Positives = 28/86 (32%), Gaps = 6/86 (6%)
Frame = +2
Query: 275 QMPPPGQQVWHQQQQ---GQPMMQQGMPPPMQQFRP---PHSMQMPPPPPPQGMQGPPRP 328
Q P QQ W Q MMQQ M Q + P PH Q PP
Sbjct: 296 QQPHQYQQQWMSMQYPATAMAMMQQQMMMYPQHYMPYVHPHHYQQQPPH----------- 442
Query: 329 LPPPSVMAGQQQVWRPPPPPQQQGGH 354
QQQV PP GGH
Sbjct: 443 ---------QQQVSPPPSAHSHHGGH 493
Score = 30.4 bits (67), Expect = 1.2
Identities = 32/93 (34%), Positives = 35/93 (37%), Gaps = 4/93 (4%)
Frame = +2
Query: 285 HQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPP----PPQGMQGPPRPLPPPSVMAGQQQ 340
HQQQQ Q QQ QQ+ SMQ P Q M P +P QQQ
Sbjct: 263 HQQQQQQQWAQQQPHQYQQQWM---SMQYPATAMAMMQQQMMMYPQHYMPYVHPHHYQQQ 433
Query: 341 VWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHG 373
PP QQQ PPP +HHG
Sbjct: 434 -----PPHQQQVS----------PPPSAHSHHG 487
>BF642399 similar to PIR|T10863|T10 extensin precursor - kidney bean, partial
(33%)
Length = 653
Score = 83.2 bits (204), Expect = 2e-16
Identities = 56/192 (29%), Positives = 81/192 (42%), Gaps = 14/192 (7%)
Frame = +3
Query: 201 TPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHV 260
T A + +S P K P+ + PP P + + P P P +P+ PPP++
Sbjct: 78 TSANSYIYSSPPPPPK--PYYYHSPPPPVHSPPPPYHYSSPPPPPKKPYKYASPPPPVYK 251
Query: 261 IQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPP-- 318
+ PPP P ++ PPP ++ + P+ + PPP P + + PPPPP
Sbjct: 252 YKSPPP--PVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPPP-----PVYKYKSPPPPPKK 410
Query: 319 PQGMQGPPRPL-----PPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPP-------P 366
P PP P+ PPP V + V++ PP P Y S PP P
Sbjct: 411 PYKYPSPPPPVYKYKSPPPPVYSPPPPVYKYKSPPPPVHSPPXPYKYKSPPPXPYKYPSP 590
Query: 367 PPPNHHGMRPPS 378
PPP + PPS
Sbjct: 591 PPPVYKYKSPPS 626
Score = 75.5 bits (184), Expect = 3e-14
Identities = 54/166 (32%), Positives = 74/166 (44%), Gaps = 17/166 (10%)
Frame = +3
Query: 229 SLPNAPQANGTIPA--PVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQ 286
+LP+ AN I + P PP+P+ PPP+H PPPP + PPP ++ +
Sbjct: 63 TLPSQTSANSYIYSSPPPPPKPYYYHSPPPPVH--SPPPP----YHYSSPPPPPKKPYKY 224
Query: 287 QQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPP--PQGMQGPPRPL-----PPPSVMAGQQ 339
P+ + PPP P + + PPPPP P PP P+ PPP V +
Sbjct: 225 ASPPPPVYKYKSPPP-----PVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKS 389
Query: 340 QVWRPPPPPQQQGGHPMG----YPQNSMPP----PPPPNHHGMRPP 377
PPPPP++ +P Y S PP PPPP + PP
Sbjct: 390 ----PPPPPKKPYKYPSPPPPVYKYKSPPPPVYSPPPPVYKYKSPP 515
Score = 60.1 bits (144), Expect = 1e-09
Identities = 44/169 (26%), Positives = 66/169 (39%), Gaps = 13/169 (7%)
Frame = +3
Query: 210 SNPTAQKSRPHTLFASGPPSL-----PNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPP 264
S+P +P+ +AS PP + P P P P P +P+ PPP++ + P
Sbjct: 186 SSPPPPPKKPYK-YASPPPPVYKYKSPPPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSP 362
Query: 265 PPQA--------PAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPP 316
PP P +P + P P V+ + P+ PPP+ +++ P PP
Sbjct: 363 PPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPPPPVYSP--PPPVYKYKSPPPPVHSPP 536
Query: 317 PPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPP 365
P + PP P PS PPPP + P + PP
Sbjct: 537 XPYKYKSPPPXPYKYPS----------PPPPVYKYKSPPSPVYKYKSPP 653
Score = 36.2 bits (82), Expect = 0.021
Identities = 28/135 (20%), Positives = 46/135 (33%), Gaps = 13/135 (9%)
Frame = +3
Query: 198 RHGTPAERVLAASNPTAQKSRPHTLFASGPP----SLPNAPQANGTIPAPVPPRPFANGV 253
++ +P V +P +P+ + PP P P P P P +P+
Sbjct: 249 KYKSPPPPVYKYKSPPPPPKKPYKYPSPPPPVYKYKSPPPPVYKYKSPPPPPKKPYKYPS 428
Query: 254 APPPIH---------VIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQ 304
PPP++ PPP P + P ++ P PPP+ +
Sbjct: 429 PPPPVYKYKSPPPPVYSPPPPVYKYKSPPPPVHSPPXPYKYKSPPPXPYKYPSPPPPVYK 608
Query: 305 FRPPHSMQMPPPPPP 319
++ P S PP
Sbjct: 609 YKSPPSPVYKYKSPP 653
>TC77624 similar to PIR|T12196|T12196 RNA-binding protein - fava bean
(fragment), partial (90%)
Length = 1214
Score = 82.0 bits (201), Expect = 3e-16
Identities = 58/189 (30%), Positives = 91/189 (47%), Gaps = 4/189 (2%)
Frame = +1
Query: 12 NLLGQHSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFV 71
N+ G+ AE + Y GNL + LL L + G + V DR T + +G+ FV
Sbjct: 352 NIGGETVAEVDTRTKLYFGNLPYSVDSALLAGLIEEYGSAELIEVLYDRDTGKSRGFAFV 531
Query: 72 EFRSEEDADYAIKVLNMIKLYGKPIRVNKASQDKKSL----DVGANLFIGNLDPDVDEKL 127
ED + I+ L+ + G+ +RVN + + K + LF+GNL V +
Sbjct: 532 TMSCVEDCNAVIQNLDGKEFMGRTLRVNFSDKPKPKEPLYPETEYKLFVGNLAWTVTSES 711
Query: 128 LYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQYLCNRQITVS 187
L F G +V +++ D +TG SRG+GF+SY + D+A+ M+ L R + VS
Sbjct: 712 LTQAFQEHGTVV-GARVLFDGETGKSRGYGFVSYATKSEMDTALAIMDNVELEGRTLRVS 888
Query: 188 YAYKKDTKG 196
A K + G
Sbjct: 889 LAQGKRS*G 915
>TC82618 similar to GP|9294614|dbj|BAB02953.1 DNA/RNA binding protein-like
{Arabidopsis thaliana}, partial (35%)
Length = 850
Score = 79.3 bits (194), Expect = 2e-15
Identities = 50/169 (29%), Positives = 88/169 (51%), Gaps = 7/169 (4%)
Frame = +1
Query: 17 HSAERNQDATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSE 76
H + + T +VG+L + E L F G + ++ V +++ T Q +GYGFVEF S
Sbjct: 343 HGSSAADNKTLWVGDLHHWMDENYLHRCFASTGEIFSIKVIRNKQTCQTEGYGFVEFTSH 522
Query: 77 EDADYAIKVLNMIKLYG--KPIRVNKAS----QDKKSLDV-GANLFIGNLDPDVDEKLLY 129
A+ ++ + + +P R+N A+ K+S +V ++F+G+L DV + +L
Sbjct: 523 GTAEKVLQTYAGMLMPNTEQPFRLNWATFSTGDHKRSDNVPDLSIFVGDLAADVTDTMLL 702
Query: 130 DTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMNGQY 178
+TFS V K++ D +TG S+G+GF+ + A+ MNG +
Sbjct: 703 ETFSDKYPSVKAAKVVFDANTGRSKGYGFVRFGDDGERSKALNEMNGVF 849
Score = 37.7 bits (86), Expect = 0.007
Identities = 24/68 (35%), Positives = 27/68 (39%)
Frame = +1
Query: 287 QQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPP 346
+Q QQ PPP PHS+ PPP MQ PP + P M Q PPP
Sbjct: 127 EQSSNNRPQQKTPPPPS----PHSLTFPPPQQWVPMQYPPAAMVMPHHMLPPQHY--PPP 288
Query: 347 PPQQQGGH 354
P H
Sbjct: 289 PHHYMAYH 312
Score = 32.7 bits (73), Expect = 0.24
Identities = 24/73 (32%), Positives = 29/73 (38%)
Frame = +1
Query: 300 PPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYP 359
P Q + PPPP P + PPP QQ V PP H M P
Sbjct: 121 PTEQSSNNRPQQKTPPPPSPHSL-----TFPPP-----QQWVPMQYPPAAMVMPHHMLPP 270
Query: 360 QNSMPPPPPPNHH 372
Q+ PPPP+H+
Sbjct: 271 QHY---PPPPHHY 300
Score = 30.8 bits (68), Expect = 0.90
Identities = 17/46 (36%), Positives = 20/46 (42%)
Frame = +1
Query: 264 PPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPH 309
PPP +P + PPP Q V Q +M M PP PPH
Sbjct: 163 PPPPSP--HSLTFPPPQQWVPMQYPPAAMVMPHHMLPPQHYPPPPH 294
>TC86766 weakly similar to PIR|T06291|T06291 extensin homolog T9E8.80 -
Arabidopsis thaliana, partial (18%)
Length = 1260
Score = 79.3 bits (194), Expect = 2e-15
Identities = 55/155 (35%), Positives = 61/155 (38%), Gaps = 2/155 (1%)
Frame = +2
Query: 225 SGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVW 284
S PP PN+P P P PP P + P P + PPPP PA P PPP
Sbjct: 26 SPPPPPPNSP------PPPPPPAPVFSPPPPVPYYYSSPPPP--PAHSP--PPPPXSPPP 175
Query: 285 HQQQQGQPMMQQ-GMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWR 343
P+ PPP PP + PPPP P PP P PPP V
Sbjct: 176 PPHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPPPPPCV--------- 328
Query: 344 PPPPPQQQGGHPMGY-PQNSMPPPPPPNHHGMRPP 377
PPPP H Y P S PPP P + PP
Sbjct: 329 EPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPP 433
Score = 77.8 bits (190), Expect = 6e-15
Identities = 55/171 (32%), Positives = 67/171 (39%), Gaps = 12/171 (7%)
Frame = +2
Query: 219 PHTLFASGPP-----SLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQP 273
P +F+ PP S P P A+ P P P P + PP + PPPP P
Sbjct: 71 PAPVFSPPPPVPYYYSSPPPPPAHSPPPPPXSPPPPPHSPTPPVYPYLSPPPPP-----P 235
Query: 274 MQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPS 333
+ PPP V+ P + PPP PP ++ PPP P Q P P P PS
Sbjct: 236 VHSPPP--PVYSPPPPSPPPCVEPPPPP-----PPPCVEPPPPSSPAPHQTPYHPPPSPS 394
Query: 334 VMAGQQQVWRPPPPPQQQGGHP---MGYPQNSMP----PPPPPNHHGMRPP 377
+ PPPP P YP P PPPPP + G PP
Sbjct: 395 PPPSPVYAYPSPPPPVYTSPPPSPVYAYPSPPPPVYSSPPPPPVYEGPIPP 547
Score = 76.6 bits (187), Expect = 1e-14
Identities = 52/157 (33%), Positives = 62/157 (39%), Gaps = 11/157 (7%)
Frame = +2
Query: 224 ASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIH-----VIQPPPPQAP-AFQPMQMP 277
A PP P +P P P P P+ + PPP+H V PPPP P +P P
Sbjct: 137 AHSPPPPPXSPPPPPHSPTP-PVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEPPPPP 313
Query: 278 PPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPL-----PPP 332
PP P PPP P P PPPP PP P+ PPP
Sbjct: 314 PPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPPVYTSPPPSPVYAYPSPPP 493
Query: 333 SVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPP 369
V + PPPPP +G P + + PPPPP
Sbjct: 494 PVYSS------PPPPPVYEGPIPPVFGISYASPPPPP 586
Score = 68.9 bits (167), Expect = 3e-12
Identities = 52/143 (36%), Positives = 55/143 (38%), Gaps = 7/143 (4%)
Frame = +2
Query: 243 PVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPM 302
P PP PPP + PPPP AP F P PPP P PPP
Sbjct: 2 PTPPY-CVRSPPPPPPNSPPPPPPPAPVFSP---PPP-----------VPYYYSSPPPP- 133
Query: 303 QQFRPPHSMQMPP--PPPPQGMQGPP-----RPLPPPSVMAGQQQVWRPPPPPQQQGGHP 355
P HS PP PPPP PP P PPP V + V+ PPPP P
Sbjct: 134 ----PAHSPPPPPXSPPPPPHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPSPPPCVEP 301
Query: 356 MGYPQNSMPPPPPPNHHGMRPPS 378
PPPPPP PPS
Sbjct: 302 --------PPPPPPPCVEPPPPS 346
Score = 50.8 bits (120), Expect = 8e-07
Identities = 40/162 (24%), Positives = 54/162 (32%), Gaps = 16/162 (9%)
Frame = +2
Query: 186 VSYAYKKDTKGERHGTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVP 245
V Y Y H P P + + + PP ++P P P
Sbjct: 101 VPYYYSSPPPPPAHSPPPPPXSPPPPPHSPTPPVYPYLSPPPPPPVHSPPPPVYSPPPPS 280
Query: 246 PRPFANGVAPPPIHVIQPPPPQAPA-----FQPMQMP-PPGQQVWHQQQQGQPMMQQ--- 296
P P PPP ++PPPP +PA + P P PP V+ P+
Sbjct: 281 PPPCVEPPPPPPPPCVEPPPPSSPAPHQTPYHPPPSPSPPPSPVYAYPSPPPPVYTSPPP 460
Query: 297 -------GMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPP 331
PPP+ PP + P PP G+ P PP
Sbjct: 461 SPVYAYPSPPPPVYSSPPPPPVYEGPIPPVFGISYASPPPPP 586
>TC84024 similar to PIR|S77714|S77714 RNA-binding protein precursor 33K -
wood tobacco, partial (55%)
Length = 867
Score = 78.6 bits (192), Expect = 4e-15
Identities = 49/164 (29%), Positives = 88/164 (52%), Gaps = 16/164 (9%)
Frame = +3
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
YVGNL ++ L ++F +AG VV+V + DR+T++ +G+ FV +S E+A AI++ +
Sbjct: 342 YVGNLPFSMTSSQLADIFAEAGTVVSVEIVYDRLTDRSRGFAFVTMKSAEEAKEAIRMFD 521
Query: 88 MIKLYGKPIRVNKASQDKKS----------------LDVGANLFIGNLDPDVDEKLLYDT 131
++ G+ +VN K +D ++ GNL ++ + L D
Sbjct: 522 GSQVGGRSAKVNFPEVPKGGERLVMGQKVRNSYRGFVDSPHKIYAGNLGWGMNSQDLRDA 701
Query: 132 FSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSFEASDSAIEAMN 175
F ++ KI+ + D G SRGFGF+++++ E ++A+ AMN
Sbjct: 702 FDEQPGLL-GAKIIYERDNGRSRGFGFVTFETAEHLEAALNAMN 830
Score = 43.1 bits (100), Expect = 2e-04
Identities = 19/72 (26%), Positives = 41/72 (56%)
Frame = +3
Query: 28 YVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAIKVLN 87
Y GNL ++ + L + F + ++ + +R + +G+GFV F + E + A+ +N
Sbjct: 651 YAGNLGWGMNSQDLRDAFDEQPGLLGAKIIYERDNGRSRGFGFVTFETAEHLEAALNAMN 830
Query: 88 MIKLYGKPIRVN 99
+++ G+P+R+N
Sbjct: 831 XVEVQGRPLRLN 866
>BF643563 similar to GP|13877739|gb unknown protein {Arabidopsis thaliana},
partial (18%)
Length = 637
Score = 78.2 bits (191), Expect = 5e-15
Identities = 59/173 (34%), Positives = 79/173 (45%), Gaps = 3/173 (1%)
Frame = +1
Query: 105 KKSLDVGANLFIGNLDPDVDEKLLYDTFSAFGVIVTNPKIMRDPDTGNSRGFGFISYDSF 164
KK +D NL+IG L P +++ L F FG IV K+++D TG S+G+GF+ Y
Sbjct: 112 KKEID-DTNLYIGYLPPTLEDDGLIQLFQQFGEIVM-AKVIKDRITGLSKGYGFVKYADI 285
Query: 165 EASDSAIEAMNGQYLCNRQITVSYAYKKDTKGERHGTPAERV---LAASNPTAQKSRPHT 221
+++AI AMNG L R I V A K G PA V S P + P
Sbjct: 286 TMANNAIVAMNGYRLEGRTIAVRVAGKPPQPVVPPGPPASAVPTYPVQSQPIG--AYPSQ 459
Query: 222 LFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPM 274
+ G P P + G P P P PPP + PPPP + + PM
Sbjct: 460 QYTGGGPIGNAPPGSYGGTPVPWGP------PIPPPYNPYAPPPPGSTMYPPM 600
Score = 56.2 bits (134), Expect = 2e-08
Identities = 30/80 (37%), Positives = 45/80 (55%)
Frame = +1
Query: 24 DATAYVGNLDPQISEELLWELFVQAGPVVNVYVPKDRVTNQHQGYGFVEFRSEEDADYAI 83
D Y+G L P + ++ L +LF Q G +V V KDR+T +GYGFV++ A+ AI
Sbjct: 127 DTNLYIGYLPPTLEDDGLIQLFQQFGEIVMAKVIKDRITGLSKGYGFVKYADITMANNAI 306
Query: 84 KVLNMIKLYGKPIRVNKASQ 103
+N +L G+ I V A +
Sbjct: 307 VAMNGYRLEGRTIAVRVAGK 366
Score = 37.0 bits (84), Expect = 0.013
Identities = 27/87 (31%), Positives = 31/87 (35%)
Frame = +1
Query: 291 QPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQ 350
QP++ G P P S + P Q G P PP G W PP PP
Sbjct: 373 QPVVPPGPPASAVPTYPVQSQPIGAYPSQQYTGGGPIGNAPPGSYGGTPVPWGPPIPP-- 546
Query: 351 QGGHPMGYPQNSMPPPPPPNHHGMRPP 377
P N PPPP + M PP
Sbjct: 547 --------PYNPYAPPPPGS--TMYPP 597
Score = 30.0 bits (66), Expect = 1.5
Identities = 24/71 (33%), Positives = 28/71 (38%), Gaps = 16/71 (22%)
Frame = +1
Query: 325 PPRPLPPPSVMAG-------QQQVWRPPPPPQQQGGHPMG-YPQNSM--------PPPPP 368
PP+P+ PP A Q Q P Q GG P+G P S PP PP
Sbjct: 367 PPQPVVPPGPPASAVPTYPVQSQPIGAYPSQQYTGGGPIGNAPPGSYGGTPVPWGPPIPP 546
Query: 369 PNHHGMRPPSG 379
P + PP G
Sbjct: 547 PYNPYAPPPPG 579
>BQ141932 weakly similar to GP|6523547|emb hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (30%)
Length = 1338
Score = 77.8 bits (190), Expect = 6e-15
Identities = 50/153 (32%), Positives = 68/153 (43%), Gaps = 9/153 (5%)
Frame = -1
Query: 225 SGPPSL-----PNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPP 279
S PP++ P A Q+ T+ P P PPP PP P+AP+ P PPP
Sbjct: 588 SAPPAIHHHPNPPASQSPATLNRPTSHTP----ACPPPSR--PPPHPRAPSLSP---PPP 436
Query: 280 GQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPS----VM 335
+ G+P PPP +QF P ++ PP + PP P PPPS ++
Sbjct: 435 PHPPTSPPRPGRPRSSTLAPPPHRQFHSPSALCTHCGIPPPAVPPPPPPTPPPSSPPALV 256
Query: 336 AGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPP 368
+ + + PPPP Q P P +PPPPP
Sbjct: 255 SPPPRPFLFPPPPNQPSPSPSPPPPPPLPPPPP 157
Score = 61.6 bits (148), Expect = 5e-10
Identities = 56/178 (31%), Positives = 73/178 (40%), Gaps = 18/178 (10%)
Frame = -2
Query: 220 HTLFASGPPSLPNAPQANGTIPA-PVPPRPFANGVAPP---PIHVIQPP-------PPQA 268
+TL P+ P+ P P P PP P + A P P H ++PP PP
Sbjct: 617 NTLLRFWEPAAPHPPSTTTLTPQHPNPPPP*TDPPATPLPAPPHPVRPPTHARLRSPPLR 438
Query: 269 PAFQPMQMPPPGQQVWHQQQQGQPM-MQQGMPPP----MQQFRPPHSMQMPPPPPPQGMQ 323
P P PPP V H + P + PPP + F PP S PPPPP +
Sbjct: 437 P---PTLRPPPPAPVAHGHRPSPPRRIGSSTPPPPFVPIAGFPPPPS-P--PPPPPLPLH 276
Query: 324 GPPR--PLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNHHGMRPPSG 379
PP+ LPP + + + PPPPP P + P PP P++H SG
Sbjct: 275 PPPQLSSLPPHAPSSFLLPLTSPPPPPP---------PPHPPPSPPRPHNHPNLGTSG 129
Score = 55.5 bits (132), Expect = 3e-08
Identities = 54/187 (28%), Positives = 72/187 (37%), Gaps = 7/187 (3%)
Frame = -3
Query: 199 HGTPAERVLAASNPTAQKSRPHTLFASGPP-------SLPNAPQANGTIPAPVPPRPFAN 251
H RV A + +++R + F PP S P A A T A +P F N
Sbjct: 769 HTRSCRRVSAIARRCTRRTREGSPFTPPPPRRTISGSSRPPASGAGSTGHA-IPYFVFGN 593
Query: 252 GVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSM 311
P PPPP P+ + P P HQ P PPP + F P
Sbjct: 592 RQRPTR----HPPPP*PPSIPIPRHPEPT----HQPHPCLPPPIPSAPPPTRAFALP--- 446
Query: 312 QMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPPPPPPNH 371
P PPP + PPR P+V+ + P P + +P+ PPPPPP+
Sbjct: 445 --PSAPPPSDLPPPPRS---PTVIDPRPPAASAVPLPLRPL-YPLRDSPPRRPPPPPPHS 284
Query: 372 HGMRPPS 378
+ PPS
Sbjct: 283 PSILPPS 263
Score = 53.9 bits (128), Expect = 1e-07
Identities = 52/175 (29%), Positives = 61/175 (34%), Gaps = 27/175 (15%)
Frame = -1
Query: 231 PNAPQANGTIPAPVPPRP-------------FANGVAPPPIHVI-QPPPPQAPAF----- 271
P P ++ P PRP G APP IH PP Q+PA
Sbjct: 693 PRPPHVGPSLAHPARPRPGRVLLVMQYPTSFLGTGSAPPAIHHHPNPPASQSPATLNRPT 514
Query: 272 -QPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRP-L 329
PPP + PPP R P PPP PP PPRP
Sbjct: 513 SHTPACPPPSR-----------------PPPHP--RAPSLSPPPPPHPP---TSPPRPGR 400
Query: 330 PPPSVMAGQQQVWRPPPPPQQQGGHP------MGYPQNSMPPPPPPNHHGMRPPS 378
P S +A PPP +Q P G P ++PPPPPP PP+
Sbjct: 399 PRSSTLA---------PPPHRQFHSPSALCTHCGIPPPAVPPPPPPTPPPSSPPA 262
Score = 53.9 bits (128), Expect = 1e-07
Identities = 44/144 (30%), Positives = 54/144 (36%), Gaps = 5/144 (3%)
Frame = -3
Query: 227 PPSLPNAPQANGTIPAPVPPRPFANGVAPPPIH--VIQPPPPQAPAFQPMQMPPPGQQVW 284
PP +P+AP P P P + + PPP VI P PP A A P+ + P
Sbjct: 496 PPPIPSAPPPTRAFALP-PSAPPPSDLPPPPRSPTVIDPRPPAASAV-PLPLRP-----L 338
Query: 285 HQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQ---GPPRPLPPPSVMAGQQQV 341
+ + P PP PP S PP P P P PLPPP+
Sbjct: 337 YPLRDSPPRRPPPPPPHSPSILPPSSRLSPPTPLPLSSSP*PALPLPLPPPT-------- 182
Query: 342 WRPPPPPQQQGGHPMGYPQNSMPP 365
PPPPP P+ P P
Sbjct: 181 --PPPPPPAPTTTPI*APAAGYTP 116
Score = 50.8 bits (120), Expect = 8e-07
Identities = 53/175 (30%), Positives = 64/175 (36%), Gaps = 20/175 (11%)
Frame = -1
Query: 209 ASNPTAQKSRP--HTLFA---SGPPSLPNAPQANGTIPA------PVPPRPFANGVAPPP 257
AS A +RP HT S PP P AP + P P P RP ++ +APPP
Sbjct: 549 ASQSPATLNRPTSHTPACPPPSRPPPHPRAPSLSPPPPPHPPTSPPRPGRPRSSTLAPPP 370
Query: 258 IHVIQPP---------PPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPP 308
P PP A P PPP +P + PPP Q P
Sbjct: 369 HRQFHSPSALCTHCGIPPPAVPPPPPPTPPPSSPPALVSPPPRPFL---FPPPPNQPSPS 199
Query: 309 HSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSM 363
P PPPP PP P PPP +PP Q+ G + +N M
Sbjct: 198 -----PSPPPP-----PPLPPPPP----------QPPQSRHQRLGIHLRVDKNKM 94
Score = 38.9 bits (89), Expect = 0.003
Identities = 51/192 (26%), Positives = 65/192 (33%), Gaps = 15/192 (7%)
Frame = -1
Query: 201 TPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVP-PRPF---ANGVAPP 256
+P L PT + P L A+ P S P G P P P P+P A+ A P
Sbjct: 918 SPRSAPLHLLAPTFTTNTP-PLIATRPRSTDITPTLRGQPPTPRPYPQPLRTRAHVGAYP 742
Query: 257 PI-----------HVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQF 305
+ H PP P+ P PG+ + Q + PP +
Sbjct: 741 RLRDAAHAVLVKAHRSPRPPHVGPSLAHPARPRPGRVLLVMQYPTSFLGTGSAPPAIHHH 562
Query: 306 RPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQNSMPP 365
P + Q P P PPPS RPPP P+ S+ P
Sbjct: 561 PNPPASQSPATLNRPTSHTPA--CPPPS---------RPPPHPRAP----------SLSP 445
Query: 366 PPPPNHHGMRPP 377
PPPP H PP
Sbjct: 444 PPPP-HPPTSPP 412
Score = 38.5 bits (88), Expect = 0.004
Identities = 58/243 (23%), Positives = 80/243 (32%), Gaps = 78/243 (32%)
Frame = -2
Query: 212 PTAQKSRPHTLFASGP-PSLPNAPQANGTIP--APVPPRP-----FANGVAPPPIHVIQP 263
P + HTL + P P + + TIP AP P P + ++PPP+H
Sbjct: 1112 PPPPLATDHTLLSPRPTPRATHPLSPSHTIPRLAPSPGLPAPLSRYYTSLSPPPLHTHDT 933
Query: 264 -PPPQAPAFQP-------------MQMPPP-GQQVWHQQQQGQPMMQQGMPP-------- 300
PP APA P + PP + H+ P ++ P
Sbjct: 932 LTPPSAPAPLPSISSRRLLRPTLHLSSPPARALPILHRPSAVNPPLRDPTPSLYAHALMS 753
Query: 301 ----PMQQFRPPHSMQM---PPPP--------PPQGMQG-------------PPRPLPPP 332
+ P+S ++ P PP PP ++G P PP
Sbjct: 752 ARIRDCETLHTPYS*RLTVHPAPPTSDHLWLIPPARVRGGFYWSCNTLLRFWEPAAPHPP 573
Query: 333 SVMAGQQQVWRPPPP-------PQQQGGHPMGYPQNS------------MPPPPPPNHHG 373
S Q PPPP P HP+ P ++ PPPP P HG
Sbjct: 572 STTTLTPQHPNPPPP*TDPPATPLPAPPHPVRPPTHARLRSPPLRPPTLRPPPPAPVAHG 393
Query: 374 MRP 376
RP
Sbjct: 392 HRP 384
Score = 36.2 bits (82), Expect = 0.021
Identities = 47/185 (25%), Positives = 67/185 (35%), Gaps = 21/185 (11%)
Frame = -1
Query: 216 KSRPHTLF---ASGPPSLPNAPQANGTIPAPV---PPRPFANGVAPPPIHVIQPPPPQAP 269
+S PHTL ++ PPS + P T P P ++ AP P+ P PP+A
Sbjct: 1152 RSTPHTLAPISSASPPSPRHRPYPPLTSPHTARYSPSLSLSHYPAPRPV----PRPPRA- 988
Query: 270 AFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPL 329
A + +P P H + P+ + P + P + PP + PR
Sbjct: 987 AVSILHLPLPTATT-HTRHTHTPLSPRSAP--LHLLAPTFTTNTPPLIATR-----PRST 832
Query: 330 PPPSVMAGQQQVWRPPPPPQQQGGHPMGYPQ------------NSMPPPP---PPNHHGM 374
+ GQ RP P P + H YP+ + P PP P H
Sbjct: 831 DITPTLRGQPPTPRPYPQPLRTRAHVGAYPRLRDAAHAVLVKAHRSPRPPHVGPSLAHPA 652
Query: 375 RPPSG 379
RP G
Sbjct: 651 RPRPG 637
Score = 35.8 bits (81), Expect = 0.028
Identities = 24/70 (34%), Positives = 27/70 (38%)
Frame = -1
Query: 200 GTPAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIH 259
G P V PT S P L + P P N P+P PP PPP
Sbjct: 327 GIPPPAVPPPPPPTPPPSSPPALVSPPPRPFLFPPPPNQPSPSPSPP-------PPPP-- 175
Query: 260 VIQPPPPQAP 269
+ PPPPQ P
Sbjct: 174 -LPPPPPQPP 148
>TC87476 homologue to PIR|T11622|T11622 extensin class 1 precursor - cowpea,
partial (48%)
Length = 939
Score = 70.1 bits (170), Expect = 1e-12
Identities = 51/163 (31%), Positives = 68/163 (41%), Gaps = 3/163 (1%)
Frame = +2
Query: 219 PHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPP 278
P ++ S PP P+ P +P PP P +PPP +V + PPP +P+ PP
Sbjct: 5 PPYIYKSPPPPSPSPPPPY-VYKSPPPPSP-----SPPPPYVYKSPPPPSPS------PP 148
Query: 279 PGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQ 338
P P + + PPP PP+ + PPPP P PPP
Sbjct: 149 P------------PYVYKSPPPPTPSPPPPYIYKSPPPPSPS---------PPP------ 247
Query: 339 QQVWRPPPPPQQQGGHPMGYPQNSMPPP---PPPNHHGMRPPS 378
V++ PPPP P Y S PPP PPP + PP+
Sbjct: 248 PYVYKSPPPPSP--SPPPPYVYKSPPPPSPSPPPPYIYKSPPT 370
Score = 69.7 bits (169), Expect = 2e-12
Identities = 45/139 (32%), Positives = 60/139 (42%), Gaps = 13/139 (9%)
Frame = +3
Query: 241 PAPVPPRPFAN------GVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMM 294
P+P PP P+ +PPP +V + PPP +P+ PPP P +
Sbjct: 372 PSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPS------PPP------------PYV 497
Query: 295 QQGMPPPMQQFRPPHSMQMPPPP-----PPQGMQGPP--RPLPPPSVMAGQQQVWRPPPP 347
+ PPP PP+ + PPPP PP G + PP P PPP +++ PPP
Sbjct: 498 YKSPPPPSASPPPPYYYKSPPPPSPSPPPPYGYKSPPPPSPSPPP------PYIYKSPPP 659
Query: 348 PQQQGGHPMGYPQNSMPPP 366
P Y NS PPP
Sbjct: 660 PSPSPPPYHPYLYNSPPPP 716
Score = 68.6 bits (166), Expect = 4e-12
Identities = 43/133 (32%), Positives = 56/133 (41%), Gaps = 5/133 (3%)
Frame = +3
Query: 245 PPRPFANGVAPPPIHVIQPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQ 304
PP P +PPP +V + PPP +P+ PPP P + + PPP
Sbjct: 369 PPSP-----SPPPPYVYKSPPPPSPS------PPP------------PYVYKSPPPPSPS 479
Query: 305 FRPPHSMQMPPPP-----PPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGYP 359
PP+ + PPPP PP + PP P P P G ++ PPPP P Y
Sbjct: 480 PPPPYVYKSPPPPSASPPPPYYYKSPPPPSPSPPPPYG----YKSPPPPSPSPPPPYIYK 647
Query: 360 QNSMPPPPPPNHH 372
P P PP +H
Sbjct: 648 SPPPPSPSPPPYH 686
Score = 58.5 bits (140), Expect(2) = 2e-14
Identities = 39/137 (28%), Positives = 57/137 (41%), Gaps = 8/137 (5%)
Frame = +2
Query: 202 PAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVI 261
P + + P + P ++ S PP P+ P +P PP P +PPP +V
Sbjct: 2 PPPYIYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPY-VYKSPPPPSP-----SPPPPYVY 163
Query: 262 QPPPPQAPAFQP---MQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPP- 317
+ PPP P+ P + PPP P + + PPP PP+ + PPPP
Sbjct: 164 KSPPPPTPSPPPPYIYKSPPPPSP-----SPPPPYVYKSPPPPSPSPPPPYVYKSPPPPS 328
Query: 318 ----PPQGMQGPPRPLP 330
PP + PP +P
Sbjct: 329 PSPPPPYIYKSPPTTIP 379
Score = 53.1 bits (126), Expect = 2e-07
Identities = 28/79 (35%), Positives = 32/79 (40%)
Frame = +3
Query: 299 PPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGY 358
PPP PP PPPP PP P PPP V++ PPPP P Y
Sbjct: 387 PPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPY------VYKSPPPPSASPPPPYYY 548
Query: 359 PQNSMPPPPPPNHHGMRPP 377
P P PP +G + P
Sbjct: 549 KSPPPPSPSPPPPYGYKSP 605
Score = 52.0 bits (123), Expect = 4e-07
Identities = 30/81 (37%), Positives = 33/81 (40%), Gaps = 2/81 (2%)
Frame = +2
Query: 299 PPPMQQFRPPHSMQMPPPPPPQGMQGPPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGY 358
PPP PP PPPP PP P PPP V++ PPPP P Y
Sbjct: 2 PPPYIYKSPPPPSPSPPPPYVYKSPPPPSPSPPP------PYVYKSPPPPSPSPPPPYVY 163
Query: 359 --PQNSMPPPPPPNHHGMRPP 377
P P PPPP + PP
Sbjct: 164 KSPPPPTPSPPPPYIYKSPPP 226
Score = 50.8 bits (120), Expect = 8e-07
Identities = 35/122 (28%), Positives = 45/122 (36%), Gaps = 4/122 (3%)
Frame = +3
Query: 212 PTAQKSRPHTLFASGPPSLPNAPQA----NGTIPAPVPPRPFANGVAPPPIHVIQPPPPQ 267
P + P ++ S PP P+ P + P+P PP P+ PPP PPPP
Sbjct: 369 PPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPPPS--ASPPPPY 542
Query: 268 APAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPMQQFRPPHSMQMPPPPPPQGMQGPPR 327
P P P ++ P PPP PP PPP P PP
Sbjct: 543 YYKSPPPPSPSPPPPYGYKSP---PPPSPSPPPPYIYKSPPPPSPSPPPYHPYLYNSPPP 713
Query: 328 PL 329
P+
Sbjct: 714 PI 719
Score = 41.6 bits (96), Expect = 5e-04
Identities = 29/101 (28%), Positives = 43/101 (41%)
Frame = +3
Query: 202 PAERVLAASNPTAQKSRPHTLFASGPPSLPNAPQANGTIPAPVPPRPFANGVAPPPIHVI 261
P V + P + P + S PP P+ P G +P PP P +PPP ++
Sbjct: 483 PPPYVYKSPPPPSASPPPPYYYKSPPPPSPSPPPPYG-YKSPPPPSP-----SPPPPYIY 644
Query: 262 QPPPPQAPAFQPMQMPPPGQQVWHQQQQGQPMMQQGMPPPM 302
+ PPP +P+ PPP +H P + PPP+
Sbjct: 645 KSPPPPSPS------PPP----YH------PYLYNSPPPPI 719
Score = 38.1 bits (87), Expect(2) = 2e-14
Identities = 21/55 (38%), Positives = 24/55 (43%), Gaps = 2/55 (3%)
Frame = +3
Query: 325 PPRPLPPPSVMAGQQQVWRPPPPPQQQGGHPMGY--PQNSMPPPPPPNHHGMRPP 377
PP P PPP V++ PPPP P Y P P PPPP + PP
Sbjct: 369 PPSPSPPPPY------VYKSPPPPSPSPPPPYVYKSPPPPSPSPPPPYVYKSPPP 515
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.315 0.136 0.426
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,723,760
Number of Sequences: 36976
Number of extensions: 474862
Number of successful extensions: 21706
Number of sequences better than 10.0: 1024
Number of HSP's better than 10.0 without gapping: 4206
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9606
length of query: 379
length of database: 9,014,727
effective HSP length: 98
effective length of query: 281
effective length of database: 5,391,079
effective search space: 1514893199
effective search space used: 1514893199
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC139882.5


