

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
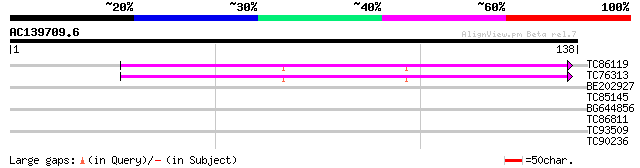
Query= AC139709.6 + phase: 0
(138 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC86119 homologue to SP|O22584|RS14_LUPLU 40S ribosomal protein ... 64 1e-11
TC76313 homologue to SP|O22584|RS14_LUPLU 40S ribosomal protein ... 64 2e-11
BE202927 homologue to SP|O22584|RS14 40S ribosomal protein S14. ... 35 0.012
TC85145 similar to GP|3288721|dbj|BAA31259.1 chalcone synthase {... 26 5.8
BG644856 similar to GP|18087544|gb| AT5g23980/MZF18_14 {Arabidop... 26 5.8
TC86811 similar to GP|9279617|dbj|BAB01075.1 gb|AAF35409.1~gene_... 26 5.8
TC93509 weakly similar to GP|3769330|dbj|BAA33879.1 alpha-amylas... 26 5.8
TC90236 similar to GP|4586308|dbj|BAA76348.1 protoporphyrinogen ... 25 9.9
>TC86119 homologue to SP|O22584|RS14_LUPLU 40S ribosomal protein S14.
[Yellow lupine] {Lupinus luteus}, complete
Length = 804
Score = 64.3 bits (155), Expect = 1e-11
Identities = 44/119 (36%), Positives = 59/119 (48%), Gaps = 9/119 (7%)
Frame = +3
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV + ASFN+T + VTD+ GR G K R +P+AA AA + +
Sbjct: 168 GVARIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVATRCKEL 347
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ + + K PG G AALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 348 GITALHIKLRATGGNKTKTPGPGAQAALRALARSGMKIGRIEDVTPIPSDSTRRKSGRR 524
>TC76313 homologue to SP|O22584|RS14_LUPLU 40S ribosomal protein S14.
[Yellow lupine] {Lupinus luteus}, complete
Length = 774
Score = 63.9 bits (154), Expect = 2e-11
Identities = 44/119 (36%), Positives = 59/119 (48%), Gaps = 9/119 (7%)
Frame = +3
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV + ASFN+T + VTD+ GR G K R +P+AA AA + +
Sbjct: 141 GVARIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVATRCKEL 320
Query: 87 GMQRAVVII--------KGPGLGRDAALRAIARSGILLRFIRDVTPIPHNGCRAPKKRR 137
G+ + + K PG G AALRA+ARSG+ + I DVTPIP + R RR
Sbjct: 321 GITALHIKLRATGGNKTKTPGPGAQAALRALARSGMKVGRIEDVTPIPSDSTRRKSGRR 497
>BE202927 homologue to SP|O22584|RS14 40S ribosomal protein S14. [Yellow
lupine] {Lupinus luteus}, partial (69%)
Length = 496
Score = 34.7 bits (78), Expect = 0.012
Identities = 22/73 (30%), Positives = 34/73 (46%), Gaps = 1/73 (1%)
Frame = +3
Query: 28 GVIYVQASFNNTIVTVTDVRGRVISWSSAGSCGFKGTR-RGTPFAAQTAAANAIRTVVDQ 86
GV + ASFN+T + VTD+ GR G K R +P+AA AA + +
Sbjct: 201 GVARIFASFNDTFIHVTDLSGRETLVRITGGMKVKADRDESSPYAAMLAAQDVATRCKEL 380
Query: 87 GMQRAVVIIKGPG 99
G+ + ++ G
Sbjct: 381 GITALHIKLRATG 419
>TC85145 similar to GP|3288721|dbj|BAA31259.1 chalcone synthase {Vitis
vinifera}, partial (98%)
Length = 1526
Score = 25.8 bits (55), Expect = 5.8
Identities = 11/22 (50%), Positives = 17/22 (77%)
Frame = +3
Query: 92 VVIIKGPGLGRDAALRAIARSG 113
+V+++ P LG+DAA +AIA G
Sbjct: 405 LVVVEVPKLGKDAAKKAIAEWG 470
>BG644856 similar to GP|18087544|gb| AT5g23980/MZF18_14 {Arabidopsis
thaliana}, partial (12%)
Length = 582
Score = 25.8 bits (55), Expect = 5.8
Identities = 12/36 (33%), Positives = 22/36 (60%)
Frame = -3
Query: 1 MAKSIPKIGSRKNGRIGSRKHPRKIPKGVIYVQASF 36
+A++IP +G +NGR+ KH + P + V+ +F
Sbjct: 301 IAETIPIVGITRNGRLRKHKHFFRAP---VLVEGNF 203
>TC86811 similar to GP|9279617|dbj|BAB01075.1
gb|AAF35409.1~gene_id:MPE11.29~similar to unknown protein
{Arabidopsis thaliana}, partial (94%)
Length = 2156
Score = 25.8 bits (55), Expect = 5.8
Identities = 14/36 (38%), Positives = 16/36 (43%), Gaps = 2/36 (5%)
Frame = +1
Query: 27 KGVIYVQASFNNTIVTVTDVRGRV--ISWSSAGSCG 60
K + VQA+ N V GRV SW S G G
Sbjct: 988 KNIFVVQAAIGNFFTAVLSREGRVYTFSWGSDGKLG 1095
>TC93509 weakly similar to GP|3769330|dbj|BAA33879.1 alpha-amylase
{Phaseolus vulgaris}, partial (20%)
Length = 474
Score = 25.8 bits (55), Expect = 5.8
Identities = 11/29 (37%), Positives = 15/29 (50%)
Frame = -2
Query: 53 WSSAGSCGFKGTRRGTPFAAQTAAANAIR 81
WS G +G R G+P A QT + A +
Sbjct: 392 WSHRACSGRRGAREGSPGAPQTGSVAAAK 306
>TC90236 similar to GP|4586308|dbj|BAA76348.1 protoporphyrinogen IX oxidase
{Glycine max}, partial (39%)
Length = 686
Score = 25.0 bits (53), Expect = 9.9
Identities = 21/68 (30%), Positives = 32/68 (46%), Gaps = 1/68 (1%)
Frame = -1
Query: 4 SIPKIGSRKNGRIGSRKHPRKIPKGVIYVQASFNNTIVT-VTDVRGRVISWSSAGSCGFK 62
S PK S+ + S R + +++ N + +T + D R + +SSA GF
Sbjct: 509 SFPKWRSKNLLTLSSEFSSRTADESLLF---HINGSKMT*IFDCAERSLLFSSAA--GFA 345
Query: 63 GTRRGTPF 70
GTR G PF
Sbjct: 344 GTRTGVPF 321
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.323 0.138 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,698,056
Number of Sequences: 36976
Number of extensions: 38466
Number of successful extensions: 205
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 202
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 203
length of query: 138
length of database: 9,014,727
effective HSP length: 86
effective length of query: 52
effective length of database: 5,834,791
effective search space: 303409132
effective search space used: 303409132
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC139709.6


