

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139709.19 + phase: 0
(353 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
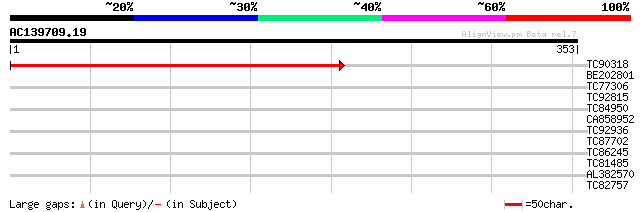
Score E
Sequences producing significant alignments: (bits) Value
TC90318 PIR|S11299|F2PMD1 photosystem II protein D1 precursor - ... 427 e-120
BE202801 homologue to GP|12322744|gb| hypothetical protein; 7880... 32 0.28
TC77306 similar to GP|17064820|gb|AAL32564.1 Unknown protein {Ar... 31 0.82
TC92815 weakly similar to GP|3413703|gb|AAC31226.1| unknown prot... 29 2.4
TC84950 29 2.4
CA858952 29 2.4
TC92936 similar to GP|15292741|gb|AAK92739.1 unknown protein {Ar... 29 3.1
TC87702 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 7... 29 3.1
TC86245 weakly similar to GP|9279671|dbj|BAB01228.1 transport in... 28 4.1
TC81485 weakly similar to PIR|T01135|T01135 probable GTP-binding... 28 7.0
AL382570 weakly similar to GP|10241621|em putative P-glycoprotei... 27 9.1
TC82757 similar to PIR|T08563|T08563 dnaJ-related protein T22F8.... 27 9.1
>TC90318 PIR|S11299|F2PMD1 photosystem II protein D1 precursor - garden pea
chloroplast, partial (58%)
Length = 697
Score = 427 bits (1097), Expect = e-120
Identities = 207/208 (99%), Positives = 207/208 (99%)
Frame = +3
Query: 1 MTAILERRDSENLWSRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI 60
MTAILERRDSENLWSRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI
Sbjct: 72 MTAILERRDSENLWSRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI 251
Query: 61 DGIREPVSGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELIVLHFL 120
DGIREPVSGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELIVLHFL
Sbjct: 252 DGIREPVSGSLLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELIVLHFL 431
Query: 121 LGVACYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTF 180
LGVACYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTF
Sbjct: 432 LGVACYMGREWELSFRLGMRPWIAVAYSAPVAAATAVFLIYPIGQGSFSDGMPLGISGTF 611
Query: 181 NFMIVFQAEHNILMHPFHMLGVAGVFGG 208
NFMIVFQAEHNILMHPFHMLGVAGVF G
Sbjct: 612 NFMIVFQAEHNILMHPFHMLGVAGVFXG 695
>BE202801 homologue to GP|12322744|gb| hypothetical protein; 78804-81924
{Arabidopsis thaliana}, partial (3%)
Length = 506
Score = 32.3 bits (72), Expect = 0.28
Identities = 18/51 (35%), Positives = 26/51 (50%)
Frame = +1
Query: 194 MHPFHMLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEGYRFGQEEE 244
MHP HM GVAG + G G S ++ N+ A GY +GQ+++
Sbjct: 82 MHPQHMQGVAGHYMGMASHQAQGGPAASMYPQQMYGNQFA--GYGYGQQQQ 228
>TC77306 similar to GP|17064820|gb|AAL32564.1 Unknown protein {Arabidopsis
thaliana}, partial (69%)
Length = 1709
Score = 30.8 bits (68), Expect = 0.82
Identities = 14/41 (34%), Positives = 23/41 (55%)
Frame = -2
Query: 20 WITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDI 60
WI T+ R+ + W G+L+ + L A + FI +PP+ I
Sbjct: 142 WIRRTDERILV-WVGLLLFFSFLQAKRDMCLLFITSPPIFI 23
>TC92815 weakly similar to GP|3413703|gb|AAC31226.1| unknown protein
{Arabidopsis thaliana}, partial (11%)
Length = 559
Score = 29.3 bits (64), Expect = 2.4
Identities = 13/47 (27%), Positives = 26/47 (54%)
Frame = -3
Query: 190 HNILMHPFHMLGVAGVFGGSLFSAMHGSLVTSSLIRETTENESANEG 236
H +L HP H+L + + S + + SL++SS + + +++ N G
Sbjct: 500 HQLLHHPLHLLLYSSLIHFSPLNTLSRSLISSSSVCYSPHHQNHNRG 360
>TC84950
Length = 826
Score = 29.3 bits (64), Expect = 2.4
Identities = 20/67 (29%), Positives = 33/67 (48%), Gaps = 2/67 (2%)
Frame = +3
Query: 28 LYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDIDGIREPV--SGSLLYGNNIISGAIIPT 85
LY GWFG++M S+ I F+ + + I V +G LY N I++ +
Sbjct: 642 LYFGWFGLVM--------SILFILFLYSDNTHLSYILYDVRYNGRFLYKNLILAYV*MYI 797
Query: 86 SAAIGLH 92
S+++ LH
Sbjct: 798 SSSVTLH 818
>CA858952
Length = 533
Score = 29.3 bits (64), Expect = 2.4
Identities = 26/86 (30%), Positives = 39/86 (45%), Gaps = 1/86 (1%)
Frame = +2
Query: 3 AILERRDSENLWSRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFIIAFIAAPPVDIDG 62
A +E R ++ L +RFC+W+ E + L P + +S FI+ F D +
Sbjct: 248 AAIETRSAQILLTRFCSWVFQFEVSPF------LQKPYIRKQSSQFILQFRPLILPDKET 409
Query: 63 IREPVSGSLLYGNN-IISGAIIPTSA 87
I P S L+ IIS + TSA
Sbjct: 410 IIHPQLESFLHNMP*IISSSTSTTSA 487
>TC92936 similar to GP|15292741|gb|AAK92739.1 unknown protein {Arabidopsis
thaliana}, partial (41%)
Length = 762
Score = 28.9 bits (63), Expect = 3.1
Identities = 13/42 (30%), Positives = 22/42 (51%)
Frame = +1
Query: 213 AMHGSLVTSSLIRETTENESANEGYRFGQEEETYNIVAAHGY 254
+++G VTS ++ + A + F +EE Y I+ HGY
Sbjct: 271 SINGPPVTSPRVKLSDGRHLAYREFGFSKEEARYKIIVIHGY 396
>TC87702 weakly similar to SP|O48923|C7DA_SOYBN Cytochrome P450 71D10 (EC
1.14.-.-). [Soybean] {Glycine max}, partial (51%)
Length = 1732
Score = 28.9 bits (63), Expect = 3.1
Identities = 16/37 (43%), Positives = 22/37 (59%)
Frame = +3
Query: 14 WSRFCNWITSTENRLYIGWFGVLMIPTLLTATSVFII 50
W +W+ EN +Y GWF +L+IP + TSV II
Sbjct: 1464 WMNPLDWLLE-ENMIY-GWFLLLIIPQVKIITSVKII 1568
>TC86245 weakly similar to GP|9279671|dbj|BAB01228.1 transport inhibitor
response-like protein {Arabidopsis thaliana}, partial
(80%)
Length = 2260
Score = 28.5 bits (62), Expect = 4.1
Identities = 16/48 (33%), Positives = 23/48 (47%)
Frame = +3
Query: 264 SFNNSRSLHFFLAAWPVVGIWFTALGISTMAFNLNGFNFNQSVVDSQG 311
+FNN RSLH W + L + NL NF+ + +DS+G
Sbjct: 1269 AFNNCRSLHTLSGLWVASAQYHQVL--YPVCTNLTFLNFSYAPLDSEG 1406
>TC81485 weakly similar to PIR|T01135|T01135 probable GTP-binding protein
(extra large) [imported] - Arabidopsis thaliana, partial
(21%)
Length = 1181
Score = 27.7 bits (60), Expect = 7.0
Identities = 21/89 (23%), Positives = 45/89 (49%), Gaps = 1/89 (1%)
Frame = -2
Query: 57 PVDIDGIREPVSGS-LLYGNNIISGAIIPTSAAIGLHFYPIWEAASVDEWLYNGGPYELI 115
P+ ++G V GS L YG I++ I+ + A + ++ I E ++ +W
Sbjct: 697 PLKLNGY---VMGSKLTYGGVIVNAVIVASGAIMANYWMKIREPVTLSQW---------N 554
Query: 116 VLHFLLGVACYMGREWELSFRLGMRPWIA 144
+L+FLL + ++ + E+ F ++ W++
Sbjct: 553 LLYFLLKLV*FVQNQQEVHF---LKSWMS 476
>AL382570 weakly similar to GP|10241621|em putative P-glycoprotein {Oryza
sativa}, partial (17%)
Length = 481
Score = 27.3 bits (59), Expect = 9.1
Identities = 18/51 (35%), Positives = 25/51 (48%), Gaps = 5/51 (9%)
Frame = -3
Query: 262 YASFNNSRSLHFFLAAWPVVGIW-----FTALGISTMAFNLNGFNFNQSVV 307
Y+ +SR L FLA V+ IW F +G S NL+ F F S++
Sbjct: 323 YSLLLSSRKLAAFLANNSVISIW*ITDKFMYIGFSC---NLHNFTFKSSII 180
>TC82757 similar to PIR|T08563|T08563 dnaJ-related protein T22F8.50 -
Arabidopsis thaliana, partial (95%)
Length = 1317
Score = 27.3 bits (59), Expect = 9.1
Identities = 17/56 (30%), Positives = 29/56 (51%), Gaps = 2/56 (3%)
Frame = -2
Query: 26 NRLYIGW--FGVLMIPTLLTATSVFIIAFIAAPPVDIDGIREPVSGSLLYGNNIIS 79
NRLY+ W L+ +L + I + + DG+ +P+S SLL+ + I+S
Sbjct: 653 NRLYVAWPLQKQLLKDYVLQNLPML*ILLLDHQQMVADGLSKPISASLLFPSAILS 486
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.324 0.139 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,820,408
Number of Sequences: 36976
Number of extensions: 169261
Number of successful extensions: 1094
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1091
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1093
length of query: 353
length of database: 9,014,727
effective HSP length: 97
effective length of query: 256
effective length of database: 5,428,055
effective search space: 1389582080
effective search space used: 1389582080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC139709.19


