

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC137839.10 - phase: 0
(502 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
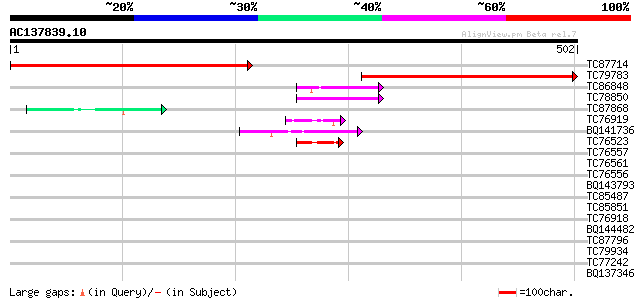
Score E
Sequences producing significant alignments: (bits) Value
TC87714 similar to PIR|S22697|S22697 extensin - Volvox carteri (... 431 e-121
TC79783 weakly similar to PIR|E96576|E96576 unknown protein 435... 387 e-108
TC86848 similar to GP|10177413|dbj|BAB10544. fibrillarin-like {A... 51 9e-07
TC78850 homologue to GP|10177413|dbj|BAB10544. fibrillarin-like ... 48 1e-05
TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR... 45 7e-05
TC76919 similar to GP|15724324|gb|AAL06555.1 At1g76010/T4O12_22 ... 44 2e-04
BQ141736 similar to PIR|T06291|T062 extensin homolog T9E8.80 - A... 44 2e-04
TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - ... 42 6e-04
TC76557 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 41 0.001
TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 41 0.001
TC76556 similar to PIR|S71779|S71779 glycine-rich RNA-binding pr... 41 0.001
BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich g... 41 0.001
TC85487 homologue to SP|P49688|RS2_ARATH 40S ribosomal protein S... 40 0.002
TC85851 similar to PIR|T12180|T12180 probable transcription fact... 40 0.002
TC76918 similar to GP|2231329|gb|AAB62000.1| bactinecin 11 {Ovis... 40 0.002
BQ144482 similar to GP|6523547|emb| hydroxyproline-rich glycopro... 40 0.002
TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear prot... 39 0.004
TC79934 similar to GP|20259633|gb|AAM14173.1 putative GAR1 prote... 39 0.004
TC77242 similar to GP|22655123|gb|AAM98152.1 putative protein {A... 39 0.005
BQ137346 homologue to GP|15027095|em hypothetical protein LT.09 ... 39 0.005
>TC87714 similar to PIR|S22697|S22697 extensin - Volvox carteri (fragment),
partial (4%)
Length = 810
Score = 431 bits (1109), Expect = e-121
Identities = 209/215 (97%), Positives = 210/215 (97%)
Frame = -2
Query: 1 MRGTIGVRLQNSTISNATRQTLVPFSTSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAA 60
MRGTIGVRLQNSTISNATRQTLVPFSTSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAA
Sbjct: 647 MRGTIGVRLQNSTISNATRQTLVPFSTSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAA 468
Query: 61 PGNPNPTPNVNESKPDATDSPIPPGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGR 120
PGNPNPTPNVNESKPDATDSPIPPGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGR
Sbjct: 467 PGNPNPTPNVNESKPDATDSPIPPGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGR 288
Query: 121 GRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSEDVRPVEPID 180
GRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSEDVRPVEPID
Sbjct: 287 GRGRGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSEDVRPVEPID 108
Query: 181 LSGDSESDNRFVMTVPKVLPGGGRGRGKPLEEAAQ 215
LSGDSESDNRFVMTVPK + GRGRGKPLE AAQ
Sbjct: 107 LSGDSESDNRFVMTVPKGVA*SGRGRGKPLERAAQ 3
Score = 36.6 bits (83), Expect = 0.023
Identities = 50/179 (27%), Positives = 70/179 (38%), Gaps = 29/179 (16%)
Frame = -2
Query: 115 GTGRGRGRGRGFD-PLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSEDV 173
G GRG GRGRG + F + P P NV+++ + P P R
Sbjct: 542 GDGRGGGRGRGGSGTVTFNFGEKAAPGNP---NPTPNVNESKPDATDSPIPPGAGRGHGR 372
Query: 174 RPVEPIDLSGDSESDNRFVMTVPKVLPGGGRGRGK-----PLEEAAQEAPQAPV-VNRHI 227
P S S + F+ ++ + G GRGRG+ P + P+ PV + R
Sbjct: 371 GGTVP---DFPSFSFSSFMSSIQQPGTGRGRGRGRGFDPLPPQFENDSVPKKPVFIKRED 201
Query: 228 RVRQT----------PADAESDNVPRRQPM---------NRFVRDDGDG---SGRGRGR 264
V QT P S++V +P+ NRFV G SGRGRG+
Sbjct: 200 NVSQTDANDFSPPKNPVFTRSEDVRPVEPIDLSGDSESDNRFVMTVPKGVA*SGRGRGK 24
Score = 28.5 bits (62), Expect = 6.3
Identities = 29/90 (32%), Positives = 34/90 (37%), Gaps = 21/90 (23%)
Frame = -2
Query: 201 GGGRGRGKP----LEEAAQEAPQAPVVNRHIRVRQTPADAESDNVP----RRQPMNRFVR 252
GGGRGRG + AP P N V ++ DA +P R V
Sbjct: 530 GGGRGRGGSGTVTFNFGEKAAPGNP--NPTPNVNESKPDATDSPIPPGAGRGHGRGGTVP 357
Query: 253 D-------------DGDGSGRGRGRGRGRD 269
D G+GRGRGRGRG D
Sbjct: 356 DFPSFSFSSFMSSIQQPGTGRGRGRGRGFD 267
>TC79783 weakly similar to PIR|E96576|E96576 unknown protein 43598-45751
[imported] - Arabidopsis thaliana, partial (29%)
Length = 1050
Score = 387 bits (993), Expect = e-108
Identities = 191/191 (100%), Positives = 191/191 (100%)
Frame = +2
Query: 312 DGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEAMDINCAIEFEP 371
DGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEAMDINCAIEFEP
Sbjct: 53 DGLYVGDNADGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEAMDINCAIEFEP 232
Query: 372 EYAVEFDNPDIDEKEPIALRDALEKMKPFLMTYEGIRSQEEWEEVIEELMQRVPLLKKIV 431
EYAVEFDNPDIDEKEPIALRDALEKMKPFLMTYEGIRSQEEWEEVIEELMQRVPLLKKIV
Sbjct: 233 EYAVEFDNPDIDEKEPIALRDALEKMKPFLMTYEGIRSQEEWEEVIEELMQRVPLLKKIV 412
Query: 432 DHYSGPDRVTAKKQQEELERVAKTLPTSAPSSVKEFTNRAVVSLQSNPGWGFDKKCQFMD 491
DHYSGPDRVTAKKQQEELERVAKTLPTSAPSSVKEFTNRAVVSLQSNPGWGFDKKCQFMD
Sbjct: 413 DHYSGPDRVTAKKQQEELERVAKTLPTSAPSSVKEFTNRAVVSLQSNPGWGFDKKCQFMD 592
Query: 492 KLVFEVSQHHK 502
KLVFEVSQHHK
Sbjct: 593 KLVFEVSQHHK 625
>TC86848 similar to GP|10177413|dbj|BAB10544. fibrillarin-like {Arabidopsis
thaliana}, partial (92%)
Length = 1250
Score = 51.2 bits (121), Expect = 9e-07
Identities = 33/85 (38%), Positives = 41/85 (47%), Gaps = 8/85 (9%)
Frame = +3
Query: 255 GDGSGRGRGRG--------RGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDRTSHQDI 306
GD GRG GRG RG ARG G RG GGRGGRG GRGG + G S +
Sbjct: 111 GDRGGRGGGRGGFGGRGGDRGTPFKARG-GGRGGGGRGGRGGGRGGGRGGGMKGGSKVIV 287
Query: 307 ARSNADGLYVGDNADGEKLAKKLGP 331
+G+++ + + K L P
Sbjct: 288 EPHRHEGIFIAKGKEDALVTKNLVP 362
Score = 34.3 bits (77), Expect = 0.11
Identities = 26/51 (50%), Positives = 26/51 (50%), Gaps = 1/51 (1%)
Frame = +3
Query: 251 VRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRG-GFKRYGDDR 300
VR G G RG GRG D RG GRGG GGRG RG FK G R
Sbjct: 69 VRGRGGGGFRG---GRGGD---RGGRGGGRGGFGGRGGDRGTPFKARGGGR 203
Score = 33.1 bits (74), Expect = 0.26
Identities = 31/94 (32%), Positives = 36/94 (37%), Gaps = 12/94 (12%)
Frame = +3
Query: 13 TISNATRQTLVPFSTSS-----------GFGGG-GGDGRGGGRGRGGSGTVTFNFGEKAA 60
T+ + T Q+L ST++ GF GG GGD G G GRGG G
Sbjct: 6 TLCSLTHQSLRNLSTTAMAPPVRGRGGGGFRGGRGGDRGGRGGGRGGFG----------G 155
Query: 61 PGNPNPTPNVNESKPDATDSPIPPGAGRGHGRGG 94
G TP G GRG GRGG
Sbjct: 156RGGDRGTPFKARGGGRGGGGRGGRGGGRGGGRGG 257
>TC78850 homologue to GP|10177413|dbj|BAB10544. fibrillarin-like
{Arabidopsis thaliana}, partial (87%)
Length = 1177
Score = 47.8 bits (112), Expect = 1e-05
Identities = 30/78 (38%), Positives = 37/78 (46%), Gaps = 1/78 (1%)
Frame = +3
Query: 255 GDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRG-GFKRYGDDRTSHQDIARSNADG 313
G G GRG G GRG D RG G GGRG RG GRG G R G S + +G
Sbjct: 108 GGGGGRGFGGGRGGDFKPRGGGRGFGGGRGARGGGRGRGGGRGGMKGGSKVVVEPHRHEG 287
Query: 314 LYVGDNADGEKLAKKLGP 331
+++ + + K L P
Sbjct: 288 IFIAKGKEDALVTKNLVP 341
Score = 36.6 bits (83), Expect = 0.023
Identities = 34/106 (32%), Positives = 36/106 (33%), Gaps = 2/106 (1%)
Frame = +3
Query: 22 LVPFSTSSGFGGGGGD--GRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATD 79
+ P GFGG GGD GRGGG GRG G +F
Sbjct: 48 MAPPRGRGGFGGRGGDRGGRGGGGGRGFGGGRGGDFK----------------------- 158
Query: 80 SPIPPGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRG 125
P G GRG G G G GRGRG GRG
Sbjct: 159 ---PRGGGRGFGGGRGA-----------------RGGGRGRGGGRG 236
Score = 32.0 bits (71), Expect = 0.57
Identities = 22/45 (48%), Positives = 24/45 (52%), Gaps = 3/45 (6%)
Frame = +3
Query: 259 GRGRGRGRGRDVYARGRGDRGRG---GRGGRGDGRGGFKRYGDDR 300
GRG GRG D RG G GRG GRGG RGG + +G R
Sbjct: 63 GRGGFGGRGGDRGGRGGGG-GRGFGGGRGGDFKPRGGGRGFGGGR 194
Score = 32.0 bits (71), Expect = 0.57
Identities = 27/70 (38%), Positives = 27/70 (38%), Gaps = 5/70 (7%)
Frame = +3
Query: 30 GFGGGGGDGRGGGRG-----RGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPP 84
G GGGGG G GGGRG RGG F G A G
Sbjct: 102 GRGGGGGRGFGGGRGGDFKPRGGGR--GFGGGRGARGG---------------------- 209
Query: 85 GAGRGHGRGG 94
G GRG GRGG
Sbjct: 210 GRGRGGGRGG 239
Score = 28.9 bits (63), Expect = 4.8
Identities = 12/15 (80%), Positives = 12/15 (80%)
Frame = +3
Query: 278 RGRGGRGGRGDGRGG 292
RGRGG GGRG RGG
Sbjct: 60 RGRGGFGGRGGDRGG 104
>TC87868 similar to PIR|T05112|T05112 splicing factor 9G8-like SR protein
RSZp22 [validated] - Arabidopsis thaliana, partial (91%)
Length = 860
Score = 45.1 bits (105), Expect = 7e-05
Identities = 42/129 (32%), Positives = 52/129 (39%), Gaps = 5/129 (3%)
Frame = +3
Query: 16 NATRQTLVPFSTSSGFGGGGGDGRG-GGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESK 74
N R L S S G GGGGG GRG GG G GGS + GE PG+
Sbjct: 225 NGWRVELSHNSRSGGGGGGGGGGRGRGGGGGGGSDLKCYECGE---PGH----------- 362
Query: 75 PDATDSPIPPGAGRGHGRGGTVPDF---PSFSFSSFMSSIQQPGTGRGRGRGRGFDPLPP 131
A + G G G R + P F PS+ S+ + P RGR + PP
Sbjct: 363 -FARECRNRGGGGAGRRRSRSPPRFRRSPSYGRRSYSPRGRSPRRRSVSPRGRSYSRSPP 539
Query: 132 QFE-NDSVP 139
+ + VP
Sbjct: 540 PYRGREEVP 566
>TC76919 similar to GP|15724324|gb|AAL06555.1 At1g76010/T4O12_22
{Arabidopsis thaliana}, partial (42%)
Length = 1207
Score = 43.5 bits (101), Expect = 2e-04
Identities = 29/56 (51%), Positives = 34/56 (59%), Gaps = 3/56 (5%)
Frame = +3
Query: 245 QPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGG---RGDGRGGFKRYG 297
+P+N + D+G+GS R RGRGRGR RGRG RGRG G GDG G YG
Sbjct: 522 KPLNEY-EDEGEGSPRMRGRGRGR---GRGRG-RGRGMYNGGMEYGDGWDGGPGYG 674
Score = 37.4 bits (85), Expect = 0.014
Identities = 21/41 (51%), Positives = 22/41 (53%)
Frame = +3
Query: 251 VRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRG 291
+R G G GRGRGRGRG GD GG G G GRG
Sbjct: 567 MRGRGRGRGRGRGRGRGMYNGGMEYGDGWDGGPGYGGRGRG 689
Score = 35.8 bits (81), Expect = 0.039
Identities = 23/54 (42%), Positives = 25/54 (45%)
Frame = +3
Query: 237 ESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGR 290
E + PR + R G G GRGRGRG G G G G GGRG GR
Sbjct: 546 EGEGSPRMRGRGR-----GRGRGRGRGRGMYNGGMEYGDGWDGGPGYGGRGRGR 692
Score = 35.4 bits (80), Expect = 0.052
Identities = 30/95 (31%), Positives = 38/95 (39%), Gaps = 1/95 (1%)
Frame = +3
Query: 32 GGGGGDGRGGGRGRGG-SGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRGH 90
G G G GRG GRGRG +G + + G PG + A A RG
Sbjct: 573 GRGRGRGRGRGRGRGMYNGGMEYGDGWDGGPGYGG------RGRGRAWGH-----AFRGR 719
Query: 91 GRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRG 125
GRG + + + + G GR RGR RG
Sbjct: 720 GRGYGAQPVGQYDYGEYDAPPAPRGRGRERGRARG 824
Score = 31.6 bits (70), Expect = 0.74
Identities = 23/59 (38%), Positives = 23/59 (38%), Gaps = 20/59 (33%)
Frame = +3
Query: 253 DDGDG-SGRGRGRGRGRDVYARGRG-------------------DRGRGGRGGRGDGRG 291
D G G GRGRGR G RGRG RGRG GR GRG
Sbjct: 654 DGGPGYGGRGRGRAWGHAFRGRGRGYGAQPVGQYDYGEYDAPPAPRGRGRERGRARGRG 830
Score = 28.1 bits (61), Expect = 8.2
Identities = 18/38 (47%), Positives = 22/38 (57%), Gaps = 1/38 (2%)
Frame = +3
Query: 245 QPMNRFVRDDGDGSGRGRGRGRGRDVYARGRG-DRGRG 281
QP+ ++ + D RGRGR R ARGRG D GRG
Sbjct: 738 QPVGQYDYGEYDAPPAPRGRGRERG-RARGRGRDAGRG 848
>BQ141736 similar to PIR|T06291|T062 extensin homolog T9E8.80 - Arabidopsis
thaliana, partial (4%)
Length = 1099
Score = 43.5 bits (101), Expect = 2e-04
Identities = 35/115 (30%), Positives = 51/115 (43%), Gaps = 6/115 (5%)
Frame = +1
Query: 204 RGRGKPLEEAAQEAPQAPVVNRHIRVR-----QTPADAESDNVPRRQPMNRFVRDDGDGS 258
R R +P Q P ++H + + +T + P+ +P R R+ G
Sbjct: 148 RTRPRPTPPRPTADLQQPQADKHTKPKHASKGETKGKEKKFTAPQTRPA-RPAREGGPRR 324
Query: 259 GRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDR-TSHQDIARSNAD 312
G GRGRGRG D + + GGRGG G +GG GD+ + +D R AD
Sbjct: 325 G-GRGRGRGPDDPQERQTETENGGRGGNGGRKGGSDGGGDENGGAEKDNRRRPAD 486
Score = 40.4 bits (93), Expect = 0.002
Identities = 27/82 (32%), Positives = 40/82 (47%), Gaps = 1/82 (1%)
Frame = +1
Query: 230 RQTPADAESDNVPRRQ-PMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGD 288
R+ PAD PR++ P N+ R G + GR + RGRGD R RG RG
Sbjct: 469 RRRPADPR----PRKEKPTNKRKRGTGTKKPKREGRRGEKKGRERGRGDGNREKRGERGG 636
Query: 289 GRGGFKRYGDDRTSHQDIARSN 310
G+G K+ + R + ++ R +
Sbjct: 637 GKGRTKKEKEKRPTKEEERREH 702
Score = 29.3 bits (64), Expect = 3.7
Identities = 26/110 (23%), Positives = 38/110 (33%), Gaps = 5/110 (4%)
Frame = +3
Query: 196 PKVLPGGGRGRGKPLEEAAQEAPQAPVVNRHIRVRQTPADAESDNVPRRQPM-----NRF 250
P PG G G G+P + + + P R RQ RQP
Sbjct: 318 PAGRPGEGEGTGRPARKTDGDRERRPGGERGPERRQRRGRGRKRGGGERQPAATGRPQAE 497
Query: 251 VRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDR 300
R + G G + + +G +R GGR G G +R G+ +
Sbjct: 498 KRKTNEQKKAGHGHQKTKKRGPQGGEEREGKRPGGRKQGEAGRERGGEGK 647
>TC76523 homologue to PIR|T09648|T09648 nucleolin homolog nuM1 - alfalfa,
partial (69%)
Length = 1615
Score = 42.0 bits (97), Expect = 6e-04
Identities = 24/41 (58%), Positives = 25/41 (60%)
Frame = +3
Query: 255 GDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKR 295
G GR GRG GR GR GRGGRGGRG GRGGF +
Sbjct: 1173 GGFGGRSGGRGGGR---FGGRDSGGRGGRGGRG-GRGGFNK 1283
Score = 33.5 bits (75), Expect = 0.20
Identities = 23/46 (50%), Positives = 24/46 (52%)
Frame = +3
Query: 254 DGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDD 299
D GSG GRG G GR R G GG GGR GRGG + G D
Sbjct: 1098 DSQGSG-GRGGG-GRSGGGRFGGGGRSGGFGGRSGGRGGGRFGGRD 1229
Score = 29.3 bits (64), Expect = 3.7
Identities = 19/47 (40%), Positives = 21/47 (44%), Gaps = 1/47 (2%)
Frame = +3
Query: 252 RDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRG-GRGDGRGGFKRYG 297
RD GRG G G + G G GGR GRG GR G + G
Sbjct: 1095 RDSQGSGGRGGGGRSGGGRFGGGGRSGGFGGRSGGRGGGRFGGRDSG 1235
Score = 29.3 bits (64), Expect = 3.7
Identities = 17/29 (58%), Positives = 17/29 (58%), Gaps = 7/29 (24%)
Frame = +3
Query: 28 SSGFGGGGGDGRGGGR-------GRGGSG 49
S GFGG G GRGGGR GRGG G
Sbjct: 1170 SGGFGGRSG-GRGGGRFGGRDSGGRGGRG 1253
Score = 28.1 bits (61), Expect = 8.2
Identities = 12/19 (63%), Positives = 12/19 (63%)
Frame = +3
Query: 31 FGGGGGDGRGGGRGRGGSG 49
FGG GRGG GRGG G
Sbjct: 1215 FGGRDSGGRGGRGGRGGRG 1271
>TC76557 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 912
Score = 41.2 bits (95), Expect = 0.001
Identities = 32/88 (36%), Positives = 37/88 (41%), Gaps = 5/88 (5%)
Frame = +1
Query: 235 DAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGD-----RGRGGRGGRGDG 289
D + N+ Q +R GSG G G GRG Y G G RG GG GG G G
Sbjct: 454 DLDGRNITVNQAQSR-------GSGGGGGGGRGGGGYGGGGGGYGGERRGYGGGGGYGGG 612
Query: 290 RGGFKRYGDDRTSHQDIARSNADGLYVG 317
GG YG+ R +R G Y G
Sbjct: 613 GGG--GYGERRGGGGGYSRGGGGGGYGG 690
Score = 34.7 bits (78), Expect = 0.088
Identities = 17/31 (54%), Positives = 18/31 (57%)
Frame = +1
Query: 19 RQTLVPFSTSSGFGGGGGDGRGGGRGRGGSG 49
R V + S G GGGGG GRGGG GG G
Sbjct: 466 RNITVNQAQSRGSGGGGGGGRGGGGYGGGGG 558
Score = 31.6 bits (70), Expect = 0.74
Identities = 15/24 (62%), Positives = 16/24 (66%), Gaps = 4/24 (16%)
Frame = +1
Query: 30 GFGGGGGDGRGGGRG----RGGSG 49
G+GGGGG G GGG G RGG G
Sbjct: 580 GYGGGGGYGGGGGGGYGERRGGGG 651
Score = 30.4 bits (67), Expect = 1.7
Identities = 16/31 (51%), Positives = 18/31 (57%), Gaps = 6/31 (19%)
Frame = +1
Query: 25 FSTSSGFGGGGGDG----RGGGRG--RGGSG 49
+ G+GGGGG G RGGG G RGG G
Sbjct: 583 YGGGGGYGGGGGGGYGERRGGGGGYSRGGGG 675
Score = 29.6 bits (65), Expect = 2.8
Identities = 17/41 (41%), Positives = 20/41 (48%), Gaps = 8/41 (19%)
Frame = +1
Query: 30 GFGGGGGD--------GRGGGRGRGGSGTVTFNFGEKAAPG 62
G+GGGGG G GGG G GG G +GE+ G
Sbjct: 538 GYGGGGGGYGGERRGYGGGGGYGGGGGG----GYGERRGGG 648
Score = 29.3 bits (64), Expect = 3.7
Identities = 15/24 (62%), Positives = 15/24 (62%)
Frame = +1
Query: 26 STSSGFGGGGGDGRGGGRGRGGSG 49
S SG GGGGG GGG G GG G
Sbjct: 493 SRGSG-GGGGGGRGGGGYGGGGGG 561
Score = 28.1 bits (61), Expect = 8.2
Identities = 12/18 (66%), Positives = 12/18 (66%)
Frame = +1
Query: 32 GGGGGDGRGGGRGRGGSG 49
GGGGG RGGG G G G
Sbjct: 640 GGGGGYSRGGGGGGYGGG 693
>TC76561 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 1054
Score = 41.2 bits (95), Expect = 0.001
Identities = 32/88 (36%), Positives = 37/88 (41%), Gaps = 5/88 (5%)
Frame = +1
Query: 235 DAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGD-----RGRGGRGGRGDG 289
D + N+ Q +R GSG G G GRG Y G G RG GG GG G G
Sbjct: 550 DLDGRNITVNQAQSR-------GSGGGGGGGRGGGGYGGGGGGYGGERRGYGGGGGYGGG 708
Query: 290 RGGFKRYGDDRTSHQDIARSNADGLYVG 317
GG YG+ R +R G Y G
Sbjct: 709 GGG--GYGERRGGGGGYSRGGGGGGYGG 786
Score = 34.7 bits (78), Expect = 0.088
Identities = 17/31 (54%), Positives = 18/31 (57%)
Frame = +1
Query: 19 RQTLVPFSTSSGFGGGGGDGRGGGRGRGGSG 49
R V + S G GGGGG GRGGG GG G
Sbjct: 562 RNITVNQAQSRGSGGGGGGGRGGGGYGGGGG 654
Score = 31.6 bits (70), Expect = 0.74
Identities = 15/24 (62%), Positives = 16/24 (66%), Gaps = 4/24 (16%)
Frame = +1
Query: 30 GFGGGGGDGRGGGRG----RGGSG 49
G+GGGGG G GGG G RGG G
Sbjct: 676 GYGGGGGYGGGGGGGYGERRGGGG 747
Score = 30.4 bits (67), Expect = 1.7
Identities = 16/31 (51%), Positives = 18/31 (57%), Gaps = 6/31 (19%)
Frame = +1
Query: 25 FSTSSGFGGGGGDG----RGGGRG--RGGSG 49
+ G+GGGGG G RGGG G RGG G
Sbjct: 679 YGGGGGYGGGGGGGYGERRGGGGGYSRGGGG 771
Score = 29.6 bits (65), Expect = 2.8
Identities = 17/41 (41%), Positives = 20/41 (48%), Gaps = 8/41 (19%)
Frame = +1
Query: 30 GFGGGGGD--------GRGGGRGRGGSGTVTFNFGEKAAPG 62
G+GGGGG G GGG G GG G +GE+ G
Sbjct: 634 GYGGGGGGYGGERRGYGGGGGYGGGGGG----GYGERRGGG 744
Score = 29.3 bits (64), Expect = 3.7
Identities = 15/24 (62%), Positives = 15/24 (62%)
Frame = +1
Query: 26 STSSGFGGGGGDGRGGGRGRGGSG 49
S SG GGGGG GGG G GG G
Sbjct: 589 SRGSG-GGGGGGRGGGGYGGGGGG 657
Score = 28.1 bits (61), Expect = 8.2
Identities = 12/18 (66%), Positives = 12/18 (66%)
Frame = +1
Query: 32 GGGGGDGRGGGRGRGGSG 49
GGGGG RGGG G G G
Sbjct: 736 GGGGGYSRGGGGGGYGGG 789
>TC76556 similar to PIR|S71779|S71779 glycine-rich RNA-binding protein GRP1
- wheat, partial (95%)
Length = 754
Score = 41.2 bits (95), Expect = 0.001
Identities = 32/88 (36%), Positives = 37/88 (41%), Gaps = 5/88 (5%)
Frame = +2
Query: 235 DAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGD-----RGRGGRGGRGDG 289
D + N+ Q +R GSG G G GRG Y G G RG GG GG G G
Sbjct: 296 DLDGRNITVNQAQSR-------GSGGGGGGGRGGGGYGGGGGGYGGERRGYGGGGGYGGG 454
Query: 290 RGGFKRYGDDRTSHQDIARSNADGLYVG 317
GG YG+ R +R G Y G
Sbjct: 455 GGG--GYGERRGGGGGYSRGGGGGGYGG 532
Score = 34.7 bits (78), Expect = 0.088
Identities = 17/31 (54%), Positives = 18/31 (57%)
Frame = +2
Query: 19 RQTLVPFSTSSGFGGGGGDGRGGGRGRGGSG 49
R V + S G GGGGG GRGGG GG G
Sbjct: 308 RNITVNQAQSRGSGGGGGGGRGGGGYGGGGG 400
Score = 31.6 bits (70), Expect = 0.74
Identities = 15/24 (62%), Positives = 16/24 (66%), Gaps = 4/24 (16%)
Frame = +2
Query: 30 GFGGGGGDGRGGGRG----RGGSG 49
G+GGGGG G GGG G RGG G
Sbjct: 422 GYGGGGGYGGGGGGGYGERRGGGG 493
Score = 30.4 bits (67), Expect = 1.7
Identities = 16/31 (51%), Positives = 18/31 (57%), Gaps = 6/31 (19%)
Frame = +2
Query: 25 FSTSSGFGGGGGDG----RGGGRG--RGGSG 49
+ G+GGGGG G RGGG G RGG G
Sbjct: 425 YGGGGGYGGGGGGGYGERRGGGGGYSRGGGG 517
Score = 29.6 bits (65), Expect = 2.8
Identities = 17/41 (41%), Positives = 20/41 (48%), Gaps = 8/41 (19%)
Frame = +2
Query: 30 GFGGGGGD--------GRGGGRGRGGSGTVTFNFGEKAAPG 62
G+GGGGG G GGG G GG G +GE+ G
Sbjct: 380 GYGGGGGGYGGERRGYGGGGGYGGGGGG----GYGERRGGG 490
Score = 29.3 bits (64), Expect = 3.7
Identities = 15/24 (62%), Positives = 15/24 (62%)
Frame = +2
Query: 26 STSSGFGGGGGDGRGGGRGRGGSG 49
S SG GGGGG GGG G GG G
Sbjct: 335 SRGSG-GGGGGGRGGGGYGGGGGG 403
Score = 28.1 bits (61), Expect = 8.2
Identities = 12/18 (66%), Positives = 12/18 (66%)
Frame = +2
Query: 32 GGGGGDGRGGGRGRGGSG 49
GGGGG RGGG G G G
Sbjct: 482 GGGGGYSRGGGGGGYGGG 535
>BQ143793 weakly similar to GP|6523547|emb| hydroxyproline-rich glycoprotein
DZ-HRGP {Volvox carteri f. nagariensis}, partial (19%)
Length = 1100
Score = 41.2 bits (95), Expect = 0.001
Identities = 29/74 (39%), Positives = 36/74 (48%), Gaps = 12/74 (16%)
Frame = -3
Query: 254 DGDGSG---RGRGRGRGRDVYARGRGDRGRGGRG-GRGDG-------RGGFKRYGD-DRT 301
+G G G G GRG G+ + G G R RGG G GRG G +G KR+G DR
Sbjct: 312 EGKGEG*NNIGEGRGDGKRKWGGGGGGRRRGGEGMGRGGGG*MIFKEKGKGKRWGG*DRV 133
Query: 302 SHQDIARSNADGLY 315
++ R DG Y
Sbjct: 132 GEREGGRGRGDGCY 91
Score = 34.3 bits (77), Expect = 0.11
Identities = 20/49 (40%), Positives = 24/49 (48%), Gaps = 13/49 (26%)
Frame = -1
Query: 255 GDGSGRGR-------GRGRGRDVY------ARGRGDRGRGGRGGRGDGR 290
G G G G+ GRG+G+D GRG+ G GG GG G GR
Sbjct: 359 GGGGGGGKRKKKREGGRGKGKDEII*EKGGGMGRGNGGEGGGGGEGGGR 213
Score = 33.1 bits (74), Expect = 0.26
Identities = 26/88 (29%), Positives = 41/88 (46%)
Frame = -2
Query: 241 VPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDR 300
V +R+ +++ + G+G G R +GRG G+G +G+ RG RG +GG +R
Sbjct: 550 VGKRKGESKWEQGQGEG-GDKRKKGRGE-----GKGGKGKTQRGKRGGKKGGRRR----- 404
Query: 301 TSHQDIARSNADGLYVGDNADGEKLAKK 328
++ G G GEK KK
Sbjct: 403 -GRREREMGYMKGSIRGGEGGGEKGKKK 323
Score = 32.3 bits (72), Expect = 0.44
Identities = 20/57 (35%), Positives = 27/57 (47%), Gaps = 17/57 (29%)
Frame = -2
Query: 256 DGSGRGRGRGRGRDVYARGRGDRGRGGRGGR-----------------GDGRGGFKR 295
+G GRG+GR RG+ + RG+G +G+ GR G G GG KR
Sbjct: 655 EGGGRGKGR-RGKRGWDRGKGSKGQETENGRRK*GKVGKRKGESKWEQGQGEGGDKR 488
Score = 32.3 bits (72), Expect = 0.44
Identities = 25/63 (39%), Positives = 28/63 (43%)
Frame = -1
Query: 32 GGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRGHG 91
G GGG G GGGR GG G G+K + E + D D I G GRG G
Sbjct: 248 GEGGGGGEGGGREWGGGG------GDK*SLKR-------KEKEKDGGDK-IG*GKGRGGG 111
Query: 92 RGG 94
GG
Sbjct: 110 EGG 102
Score = 30.4 bits (67), Expect = 1.7
Identities = 17/42 (40%), Positives = 24/42 (56%), Gaps = 3/42 (7%)
Frame = -2
Query: 259 GRGRGRGRGRD---VYARGRGDRGRGGRGGRGDGRGGFKRYG 297
GRG+G G G+ ++ G G G+GG+ RG +GG K G
Sbjct: 136 GRGKGGGEGKGGWMLWGMGGGGGGKGGKKRRGK-KGGKKERG 14
Score = 30.4 bits (67), Expect = 1.7
Identities = 35/135 (25%), Positives = 56/135 (40%), Gaps = 3/135 (2%)
Frame = -3
Query: 201 GGGRGRGKPLEEAAQEAPQAPVVNRHIRVRQTPADAESDNVPRRQPMNRFVRDDG---DG 257
GGGRG G +V R +RV++ D ++ R+ R+ G
Sbjct: 630 GGGRGDG--------------IVVREVRVKRRKMDGANEE--------RWGRERGRVNGN 517
Query: 258 SGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGDDRTSHQDIARSNADGLYVG 317
G+GRG R + + +RG+ G G+ +G + ++ + DI R G+ G
Sbjct: 516 RGKGRGGTSERKEEGKEKEERGKRKEGKGGERKG--EEGEEEGSGKWDI*R----GV*GG 355
Query: 318 DNADGEKLAKKLGPE 332
G+K KK G E
Sbjct: 354 GRGGGKKEKKKRGWE 310
>TC85487 homologue to SP|P49688|RS2_ARATH 40S ribosomal protein S2.
[Mouse-ear cress] {Arabidopsis thaliana}, partial (88%)
Length = 1114
Score = 40.4 bits (93), Expect = 0.002
Identities = 33/103 (32%), Positives = 47/103 (45%), Gaps = 3/103 (2%)
Frame = +1
Query: 264 RGRGRDVYARGRGDRG--RGGRGGRGD-GRGGFKRYGDDRTSHQDIARSNADGLYVGDNA 320
RG R + G G RG RGGRGGRGD GRGG +R G R + G V
Sbjct: 82 RGGDRGGFGSGFGGRGGDRGGRGGRGDRGRGGRRRAGGRREEEEKWVPVTKLGRLV---- 249
Query: 321 DGEKLAKKLGPEIMDQITEAYEEIIERVLPSPLQDEYVEAMDI 363
E + L + + +II+ ++ L+DE ++ M +
Sbjct: 250 -KEGKIRSLEQIYLHSLPIKEHQIIDTLVGPTLKDEVMKIMPV 375
Score = 35.8 bits (81), Expect = 0.039
Identities = 24/45 (53%), Positives = 25/45 (55%), Gaps = 5/45 (11%)
Frame = +1
Query: 247 MNRFVRDDGDGSGRGRG-RGRGRDVYARG-RGDRGRGGR---GGR 286
+N GD G G G GRG D RG RGDRGRGGR GGR
Sbjct: 64 INNMAERGGDRGGFGSGFGGRGGDRGGRGGRGDRGRGGRRRAGGR 198
Score = 30.8 bits (68), Expect = 1.3
Identities = 17/23 (73%), Positives = 17/23 (73%), Gaps = 4/23 (17%)
Frame = +1
Query: 29 SGFGGGGGD--GRG--GGRGRGG 47
SGFGG GGD GRG G RGRGG
Sbjct: 109 SGFGGRGGDRGGRGGRGDRGRGG 177
>TC85851 similar to PIR|T12180|T12180 probable transcription factor - fava
bean, partial (93%)
Length = 2007
Score = 40.4 bits (93), Expect = 0.002
Identities = 41/131 (31%), Positives = 52/131 (39%), Gaps = 9/131 (6%)
Frame = +3
Query: 178 PIDLSGDSESDN---------RFVMTVPKVLPGGGRGRGKPLEEAAQEAPQAPVVNRHIR 228
P DL GD D + P G + +GKP + A + P
Sbjct: 243 PFDLLGDDAEDPSQLIVSEQLKAAAAPPAAKKGAVKEQGKPNKPALLPSKPTP------- 401
Query: 229 VRQTPADAESDNVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGD 288
PA A ++ R++G GRG G GRG D R RG GRGG G G
Sbjct: 402 ----PAQAVRES-----------RNEGGRGGRGFG-GRGGD---RDRGFGGRGGDRGFGG 524
Query: 289 GRGGFKRYGDD 299
GRGG + +G D
Sbjct: 525 GRGG-RGFGRD 554
Score = 39.3 bits (90), Expect = 0.004
Identities = 71/286 (24%), Positives = 103/286 (35%), Gaps = 23/286 (8%)
Frame = +3
Query: 30 GFGGGGGD-GRGGGRGRGGSGTVTFN----FGEKAAPGNPNPTPNVNESKPDATDSPIPP 84
GFGG GGD G GGGRG G G N F AP N P + + P
Sbjct: 486 GFGGRGGDRGFGGGRGGRGFGRDYSNDDNSFPASGAPENQGPVEE-GDRNSERRSYGGPR 662
Query: 85 GAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGR-----GRGRGFDPLPPQFENDSVP 139
G RG RGG + + GTGRG G GRG E D +
Sbjct: 663 GPYRGGRRGGFNNGDAGEEGRPRRTFERHSGTGRGNDFKREGAGRGNWGT----ETDEIA 830
Query: 140 K--KPVFIKREDNVS-QTDANDFSPPKNPVFTRSEDVRPVEPIDLSGDSESDNRFVMTVP 196
+ + V + E N+ + A + + + + + EP D E + +
Sbjct: 831 QVTEEVMNEGEKNLGDEKPAVEADVAEGNKDSPANEAEEKEPEDKEMTLEEYEKVLEEKR 1010
Query: 197 KVLPGGGRGRGKPLEEAAQEAPQAPVVNRHI----------RVRQTPADAESDNVPRRQP 246
K L + + ++ A E QA + + ++ A + + +
Sbjct: 1011KALQTL-KTEERKVDTKAFETMQALSCKKDNTEIFAKLGADKDKRKEAYDKEEKAKKSVS 1187
Query: 247 MNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGG 292
+N F++ +G GRGR GRGGRG RG G G
Sbjct: 1188INEFLKP-AEGESHYNSGGRGR----------GRGGRGARGGGFRG 1292
Score = 37.4 bits (85), Expect = 0.014
Identities = 72/290 (24%), Positives = 106/290 (35%), Gaps = 35/290 (12%)
Frame = +3
Query: 30 GFGGGGGD------GRGG----GRGRGGSG------TVTFNFGEKAAPGNPNPTPNVNES 73
GFGG GGD GRGG G GRGG G +F AP N P +
Sbjct: 453 GFGGRGGDRDRGFGGRGGDRGFGGGRGGRGFGRDYSNDDNSFPASGAPENQGPVEE-GDR 629
Query: 74 KPDATDSPIPPGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRG-----RGRGRGFDP 128
+ P G RG RGG + + GTGRG G GRG
Sbjct: 630 NSERRSYGGPRGPYRGGRRGGFNNGDAGEEGRPRRTFERHSGTGRGNDFKREGAGRG--- 800
Query: 129 LPPQFENDSVPK--KPVFIKREDNV-SQTDANDFSPPKNPVFTRSEDVRPVEPIDLSGDS 185
E D + + + V + E N+ + A + + + + + EP D
Sbjct: 801 -NWGTETDEIAQVTEEVMNEGEKNLGDEKPAVEADVAEGNKDSPANEAEEKEPEDKEMTL 977
Query: 186 ESDNRFVMTVPKVLPGGGRGRGKPLEEAAQEAPQAPVVNRH----------IRVRQTPAD 235
E + + K L + + ++ A E QA + + ++ A
Sbjct: 978 EEYEKVLEEKRKALQ-TLKTEERKVDTKAFETMQALSCKKDNTEIFAKLGADKDKRKEAY 1154
Query: 236 AESDNVPRRQPMNRFVRD-DGDGSGRGRGRGRGRDVYARGRGDRGRGGRG 284
+ + + +N F++ +G+ GRGRGR GRG RG G RG
Sbjct: 1155DKEEKAKKSVSINEFLKPAEGESHYNSGGRGRGRG----GRGARGGGFRG 1292
>TC76918 similar to GP|2231329|gb|AAB62000.1| bactinecin 11 {Ovis aries},
partial (13%)
Length = 576
Score = 40.4 bits (93), Expect = 0.002
Identities = 29/64 (45%), Positives = 30/64 (46%), Gaps = 13/64 (20%)
Frame = +1
Query: 255 GDGSGRGRGRGRGR--------DVYARGRGDRGRG-----GRGGRGDGRGGFKRYGDDRT 301
G G GRGRGRGRG D Y GRG GRG GR RG GRG YG
Sbjct: 4 GRGRGRGRGRGRGMYNGGMEYGDGYDGGRGYGGRGRGRAWGRAFRGRGRG----YGAQPV 171
Query: 302 SHQD 305
+ D
Sbjct: 172 GYYD 183
Score = 35.0 bits (79), Expect = 0.067
Identities = 30/94 (31%), Positives = 36/94 (37%)
Frame = +1
Query: 34 GGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRGHGRG 93
G G GRG GRGRG +N G + G + A GRG G G
Sbjct: 4 GRGRGRGRGRGRG-----MYNGGMEYGDGYDGGRGYGGRGRGRAWGRAF---RGRGRGYG 159
Query: 94 GTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRGFD 127
+ + + G GRGRGRGRG D
Sbjct: 160 AQPVGYYDNGEYDAPPAPRGQGRGRGRGRGRGRD 261
Score = 33.1 bits (74), Expect = 0.26
Identities = 24/59 (40%), Positives = 25/59 (41%), Gaps = 20/59 (33%)
Frame = +1
Query: 253 DDGDG-SGRGRGRGRGRDVYARGRG-------------------DRGRGGRGGRGDGRG 291
D G G GRGRGR GR RGRG RG+G GRG GRG
Sbjct: 79 DGGRGYGGRGRGRAWGRAFRGRGRGYGAQPVGYYDNGEYDAPPAPRGQGRGRGRGRGRG 255
Score = 32.7 bits (73), Expect = 0.33
Identities = 29/94 (30%), Positives = 35/94 (36%)
Frame = +1
Query: 32 GGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRGHG 91
G G G GRG GRG +N G + G + A A RG G
Sbjct: 4 GRGRGRGRGRGRG-------MYNGGMEYGDGYDGGRGYGGRGRGRAWGR-----AFRGRG 147
Query: 92 RGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRG 125
RG + + + G GRGRGRGRG
Sbjct: 148 RGYGAQPVGYYDNGEYDAPPAPRGQGRGRGRGRG 249
Score = 32.3 bits (72), Expect = 0.44
Identities = 13/15 (86%), Positives = 13/15 (86%)
Frame = +1
Query: 255 GDGSGRGRGRGRGRD 269
G G GRGRGRGRGRD
Sbjct: 217 GQGRGRGRGRGRGRD 261
Score = 32.0 bits (71), Expect = 0.57
Identities = 32/102 (31%), Positives = 35/102 (33%), Gaps = 6/102 (5%)
Frame = +1
Query: 30 GFGGGG---GDGRGGGRGRGGSGTVTF---NFGEKAAPGNPNPTPNVNESKPDATDSPIP 83
G GG GDG GGRG GG G F + P + + DA P P
Sbjct: 40 GMYNGGMEYGDGYDGGRGYGGRGRGRAWGRAFRGRGRGYGAQPVGYYDNGEYDAP--PAP 213
Query: 84 PGAGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRG 125
G GRG GRG RGRGR G
Sbjct: 214 RGQGRGRGRG------------------------RGRGRDAG 267
Score = 28.1 bits (61), Expect = 8.2
Identities = 18/42 (42%), Positives = 22/42 (51%)
Frame = +1
Query: 245 QPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGR 286
QP+ + + D RG+GRG RGRG RGRG GR
Sbjct: 163 QPVGYYDNGEYDAPPAPRGQGRG-----RGRG-RGRGRDAGR 270
>BQ144482 similar to GP|6523547|emb| hydroxyproline-rich glycoprotein DZ-HRGP
{Volvox carteri f. nagariensis}, partial (16%)
Length = 993
Score = 40.0 bits (92), Expect = 0.002
Identities = 52/186 (27%), Positives = 65/186 (33%), Gaps = 9/186 (4%)
Frame = -1
Query: 124 RGFDPLPPQFENDSVPKKPVFIKREDNVSQTDANDFSPPKNPVFTRSEDVRPVEPIDLSG 183
RG P P Q + P F+++ PP +F S P L G
Sbjct: 726 RGALPFPSQGPGRTPPTLKCFLRKRGPGPHPGRGGERPPPEAIFWAS-------PPLLKG 568
Query: 184 DSESDNRFVMTVPKVLPGGGRGRGKPLEEA----AQEAPQAPVVNRHIRVRQTPADAESD 239
F T+ P GG R PL++ + P+A R+ R P
Sbjct: 567 GGGG---FFKTLSPPSPPGGAPRS-PLKKKKGGRSLPPPRASRGRRYAAARCAPVTGP-- 406
Query: 240 NVPRRQPMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRG-----GRGGRGDGRGGFK 294
P G G G G GRG G A GRG GRG G GG G GRG +
Sbjct: 405 ------PGGAGQHSGGTGPGGGPGRGGGARG-AGGRGGEGRGQGEAEGGGGGGRGRGAKE 247
Query: 295 RYGDDR 300
G +R
Sbjct: 246 GEGRER 229
Score = 39.7 bits (91), Expect = 0.003
Identities = 39/118 (33%), Positives = 44/118 (37%), Gaps = 3/118 (2%)
Frame = -1
Query: 28 SSGFGGGGGDGRGGG-RGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGA 86
S G G GGG GRGGG RG GG G GE G E G
Sbjct: 384 SGGTGPGGGPGRGGGARGAGGRGGEGRGQGEAEGGGGGGRGRGAKE------------GE 241
Query: 87 GRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGR--GRGRGRGFDPLPPQFENDSVPKKP 142
GR GRGG ++ G GR G G GRG P P ++P +P
Sbjct: 240 GRERGRGGG---------DRRGGGGEKVGAGRLWGAGAGRGAAP-PRPAPGRALPARP 97
Score = 38.1 bits (87), Expect = 0.008
Identities = 24/56 (42%), Positives = 26/56 (45%)
Frame = +2
Query: 29 SGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPP 84
SG GGG G RGG GR G G A P P P +S+P T SP PP
Sbjct: 62 SGAGGGAGGRRGGRAGRARPGA-----GRGGAAPRPAPAP---QSRPAPTFSPPPP 205
Score = 33.1 bits (74), Expect = 0.26
Identities = 23/71 (32%), Positives = 27/71 (37%)
Frame = -2
Query: 24 PFSTSSGFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIP 83
P +T +G G GG G GGGRG G+G E+ G
Sbjct: 395 PANTRAGRGRAGGRGGGGGRGGPGAGGAKAGAKERLRGGGGG-----------------G 267
Query: 84 PGAGRGHGRGG 94
G GR GRGG
Sbjct: 266 GGGGRKRGRGG 234
Score = 33.1 bits (74), Expect = 0.26
Identities = 19/39 (48%), Positives = 21/39 (53%), Gaps = 1/39 (2%)
Frame = -3
Query: 255 GDGSGRGRGRGRGRDVYARGRGDRGRGG-RGGRGDGRGG 292
G G G G RG G GR +GRGG +GGRG GG
Sbjct: 277 GGGEGEGGERGGGEGEGEGGRR*KGRGGGKGGRGATLGG 161
Score = 32.7 bits (73), Expect = 0.33
Identities = 29/100 (29%), Positives = 40/100 (40%), Gaps = 6/100 (6%)
Frame = -3
Query: 32 GGGGGDGRGGGRG------RGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPG 85
GGG RG G RGG G +F K G P +P + P ++ P PG
Sbjct: 628 GGGKAPPRGHFLGFPPPFERGGGGIFQNSFPAKPTGGGPPVSP*KKKRGPFSSSPPGEPG 449
Query: 86 AGRGHGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRG 125
+ RGG + + ++ + G GRG G G G
Sbjct: 448 SAL---RGGAMRASDGAARRG-RPTLGRDGAGRGAGAGGG 341
Score = 31.6 bits (70), Expect = 0.74
Identities = 25/66 (37%), Positives = 31/66 (46%), Gaps = 8/66 (12%)
Frame = -2
Query: 30 GFGGGGGDGR---GGGRGRGGSGTVTFNFGEKAAPGN-----PNPTPNVNESKPDATDSP 81
G GGGGG GR GGRG GG+ G++ A G+ P P ++P A SP
Sbjct: 281 GGGGGGGGGRKRGRGGRGGGGAEIEGEGGGKRWARGDFGGRGPVAGPR-RRARPPAGPSP 105
Query: 82 IPPGAG 87
P G
Sbjct: 104 RGPPGG 87
Score = 30.8 bits (68), Expect = 1.3
Identities = 20/69 (28%), Positives = 24/69 (33%), Gaps = 19/69 (27%)
Frame = -3
Query: 32 GGGGGDGRGG------------------GRGRGGSGTVTFNFG-EKAAPGNPNPTPNVNE 72
GGGGG+G GG G G+GG G G + P P P
Sbjct: 283 GGGGGEGEGGERGGGEGEGEGGRR*KGRGGGKGGRGATLGGGGRSRGRAAAPGPRPGPPR 104
Query: 73 SKPDATDSP 81
+ P A P
Sbjct: 103 AAPPAASGP 77
Score = 30.8 bits (68), Expect = 1.3
Identities = 18/37 (48%), Positives = 20/37 (53%)
Frame = -2
Query: 260 RGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRY 296
RG G G G RGRG RGG G +G GG KR+
Sbjct: 287 RGGGGGGGGGGRKRGRGG--RGGGGAEIEGEGGGKRW 183
Score = 30.0 bits (66), Expect = 2.2
Identities = 19/47 (40%), Positives = 21/47 (44%)
Frame = -3
Query: 252 RDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYGD 298
R G G G GRG G RG G GG G G RGG + G+
Sbjct: 364 RGAGAGGGGAGGRGPGGRRPGPRRG*GGGGGGEGEGGERGGGEGEGE 224
Score = 28.9 bits (63), Expect = 4.8
Identities = 19/42 (45%), Positives = 21/42 (49%), Gaps = 3/42 (7%)
Frame = -3
Query: 252 RDDGDGSGRGRGRGRGRDVYARGRGDRGRG---GRGGRGDGR 290
R G+G G G R +G RG G GRG G GGR GR
Sbjct: 250 RGGGEGEGEGGRR*KG-----RGGGKGGRGATLGGGGRSRGR 140
>TC87796 weakly similar to SP|P03211|EBN1_EBV EBNA-1 nuclear protein.
[strain B95-8 Human herpesvirus 4] {Epstein-barr
virus}, partial (23%)
Length = 431
Score = 39.3 bits (90), Expect = 0.004
Identities = 30/99 (30%), Positives = 39/99 (39%)
Frame = +1
Query: 30 GFGGGGGDGRGGGRGRGGSGTVTFNFGEKAAPGNPNPTPNVNESKPDATDSPIPPGAGRG 89
G+GGGGG G GGG G GG+ V + G G +P G G G
Sbjct: 19 GYGGGGGSGGGGGGGAGGAHGVGYGSGGGTGGGYGGGSPGGG-------------GGGGG 159
Query: 90 HGRGGTVPDFPSFSFSSFMSSIQQPGTGRGRGRGRGFDP 128
G GG ++ + G G G G G G++P
Sbjct: 160 SGGGGGGGGAHGGAYGGGI------GGGEGGGHG-GYEP 255
Score = 32.3 bits (72), Expect = 0.44
Identities = 13/23 (56%), Positives = 14/23 (60%)
Frame = +1
Query: 27 TSSGFGGGGGDGRGGGRGRGGSG 49
T G+GGG G GGG G GG G
Sbjct: 106 TGGGYGGGSPGGGGGGGGSGGGG 174
Score = 29.3 bits (64), Expect = 3.7
Identities = 13/22 (59%), Positives = 13/22 (59%)
Frame = +1
Query: 28 SSGFGGGGGDGRGGGRGRGGSG 49
S G GGGGG GGG G G G
Sbjct: 130 SPGGGGGGGGSGGGGGGGGAHG 195
>TC79934 similar to GP|20259633|gb|AAM14173.1 putative GAR1 protein
{Arabidopsis thaliana}, partial (87%)
Length = 814
Score = 39.3 bits (90), Expect = 0.004
Identities = 22/47 (46%), Positives = 26/47 (54%)
Frame = +1
Query: 246 PMNRFVRDDGDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGG 292
P+ RF+ + GRG GRG + RG G RG GG GRG RGG
Sbjct: 466 PLARFLPQPKGQASGGRGGGRGGGGFGRG-GGRGGGGFRGRGPPRGG 603
Score = 36.6 bits (83), Expect = 0.023
Identities = 22/39 (56%), Positives = 22/39 (56%)
Frame = +1
Query: 255 GDGSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGF 293
G G GRG G RGR GRG RGGRGG GRG F
Sbjct: 544 GRGGGRGGGGFRGRGPPRGGRGP-PRGGRGGGFRGRGRF 657
Score = 35.8 bits (81), Expect = 0.039
Identities = 26/58 (44%), Positives = 31/58 (52%), Gaps = 7/58 (12%)
Frame = +1
Query: 247 MNRFVRDDGDGSGRGRGRGRGRDVYARGRGD----RGRGG---RGGRGDGRGGFKRYG 297
+++ +R G GRG G GRD RG D RGRGG GGRG G GGF+ G
Sbjct: 40 LSQTMRPPARGGGRGGGFRGGRDGGFRGGRDGGGFRGRGGGRFGGGRGGG-GGFRDEG 210
Score = 35.8 bits (81), Expect = 0.039
Identities = 22/37 (59%), Positives = 22/37 (59%), Gaps = 1/37 (2%)
Frame = +1
Query: 257 GSGRGRGRGRGRDVYARGRGDRGRG-GRGGRGDGRGG 292
G G GRG GRG G G RGRG RGGRG RGG
Sbjct: 532 GGGFGRGGGRG------GGGFRGRGPPRGGRGPPRGG 624
Score = 32.3 bits (72), Expect = 0.44
Identities = 14/18 (77%), Positives = 14/18 (77%)
Frame = +1
Query: 30 GFGGGGGDGRGGGRGRGG 47
G GGGG GRGGGRG GG
Sbjct: 520 GGRGGGGFGRGGGRGGGG 573
Score = 30.4 bits (67), Expect = 1.7
Identities = 13/17 (76%), Positives = 13/17 (76%)
Frame = +1
Query: 30 GFGGGGGDGRGGGRGRG 46
GFG GGG G GG RGRG
Sbjct: 538 GFGRGGGRGGGGFRGRG 588
Score = 29.3 bits (64), Expect = 3.7
Identities = 15/23 (65%), Positives = 15/23 (65%), Gaps = 3/23 (13%)
Frame = +1
Query: 30 GFGGGGGDGRGGGR---GRGGSG 49
G GGG GRGGGR GRGG G
Sbjct: 124 GRDGGGFRGRGGGRFGGGRGGGG 192
Score = 28.9 bits (63), Expect = 4.8
Identities = 21/42 (50%), Positives = 23/42 (54%), Gaps = 1/42 (2%)
Frame = +1
Query: 260 RGRGRGRGRDVYARGRGDRGRGGRGGRG-DGRGGFKRYGDDR 300
RG GRG G + GR RGGR G G GRGG R+G R
Sbjct: 67 RGGGRGGG---FRGGRDGGFRGGRDGGGFRGRGG-GRFGGGR 180
Score = 28.5 bits (62), Expect = 6.3
Identities = 17/30 (56%), Positives = 17/30 (56%), Gaps = 5/30 (16%)
Frame = +1
Query: 25 FSTSSGFGGGGGDGRG---GGRG--RGGSG 49
F G GGGG GRG GGRG RGG G
Sbjct: 541 FGRGGGRGGGGFRGRGPPRGGRGPPRGGRG 630
>TC77242 similar to GP|22655123|gb|AAM98152.1 putative protein {Arabidopsis
thaliana}, partial (43%)
Length = 1244
Score = 38.9 bits (89), Expect = 0.005
Identities = 26/51 (50%), Positives = 32/51 (61%), Gaps = 1/51 (1%)
Frame = +2
Query: 257 GSGRGRGRGRGRDVYARGRGDRGRGGRGGRGDGRGGFKRYG-DDRTSHQDI 306
G RG G GRG+ + R RG RGRG G RG GRGG + G DD+ S +D+
Sbjct: 716 GEFRGPG-GRGQGI-RRNRG-RGRGSGGPRGGGRGGGRGRGRDDKVSAEDL 859
Score = 31.2 bits (69), Expect = 0.97
Identities = 14/23 (60%), Positives = 15/23 (64%)
Frame = +2
Query: 30 GFGGGGGDGRGGGRGRGGSGTVT 52
G GG G GRGGGRGRG V+
Sbjct: 779 GSGGPRGGGRGGGRGRGRDDKVS 847
>BQ137346 homologue to GP|15027095|em hypothetical protein LT.09 {Leishmania
major}, partial (0%)
Length = 1101
Score = 38.9 bits (89), Expect = 0.005
Identities = 34/114 (29%), Positives = 50/114 (43%), Gaps = 5/114 (4%)
Frame = +3
Query: 204 RGRGKPLEEAAQEAP--QAPVVNRHIR--VRQTPADAESDNVPRRQPMNRFVRDDGDGSG 259
R KP E A++A QAP N+ + PA +N P+ Q R
Sbjct: 216 RAGPKPKRENAKDAHKRQAPTTNKFSAGATQHPPARHTKNNTPKTQKQTTHDRQTTQNKS 395
Query: 260 RGRGRGRGRDVYARGRGDRGRGGR-GGRGDGRGGFKRYGDDRTSHQDIARSNAD 312
RG+ R ++ +GR DRG R G+G + G + G+DR Q +R A+
Sbjct: 396 RGK---RAKN---QGRPDRGTERRKAGQGGEKQGPAQQGNDRKKKQPASRGRAE 539
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.313 0.137 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,619,496
Number of Sequences: 36976
Number of extensions: 181309
Number of successful extensions: 5215
Number of sequences better than 10.0: 264
Number of HSP's better than 10.0 without gapping: 1968
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4008
length of query: 502
length of database: 9,014,727
effective HSP length: 100
effective length of query: 402
effective length of database: 5,317,127
effective search space: 2137485054
effective search space used: 2137485054
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC137839.10


