

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
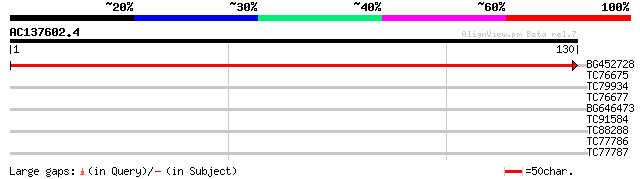
Query= AC137602.4 - phase: 0
(130 letters)
Database: MTGI
36,976 sequences; 27,044,181 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BG452728 weakly similar to PIR|T21367|T213 hypothetical protein ... 258 4e-70
TC76675 SP|P46259|TBA1_PEA Tubulin alpha-1 chain. [Garden pea] {... 29 0.46
TC79934 similar to GP|20259633|gb|AAM14173.1 putative GAR1 prote... 29 0.46
TC76677 homologue to SP|P33629|TBA_PRUDU Tubulin alpha chain. [A... 29 0.46
BG646473 similar to GP|15293255|gb| unknown protein {Arabidopsis... 27 1.7
TC91584 similar to GP|21592519|gb|AAM64469.1 unknown {Arabidopsi... 26 3.9
TC88288 weakly similar to GP|10172602|dbj|BAB01406. gene_id:MJG1... 26 3.9
TC77786 26 5.0
TC77787 25 8.6
>BG452728 weakly similar to PIR|T21367|T213 hypothetical protein F25H8.2 -
Caenorhabditis elegans, partial (5%)
Length = 538
Score = 258 bits (660), Expect = 4e-70
Identities = 127/130 (97%), Positives = 127/130 (97%)
Frame = +1
Query: 1 MEMIPLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVN 60
MEMIPLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVN
Sbjct: 40 MEMIPLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVN 219
Query: 61 YVMHPVSMMKRCPMKQNFWMMRKRLSTRDCKNIIKEVVTIKTPIERMENNIKQIPLKDGS 120
YVMHPVSMM RCPMK NFWMM KRLSTRDCKNIIKEVVTIKTPIERMENNIKQIPLKDGS
Sbjct: 220 YVMHPVSMMXRCPMKHNFWMMXKRLSTRDCKNIIKEVVTIKTPIERMENNIKQIPLKDGS 399
Query: 121 IPTMPVALGG 130
IPTMPVALGG
Sbjct: 400 IPTMPVALGG 429
>TC76675 SP|P46259|TBA1_PEA Tubulin alpha-1 chain. [Garden pea] {Pisum
sativum}, partial (98%)
Length = 1660
Score = 29.3 bits (64), Expect = 0.46
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 2/91 (2%)
Frame = +3
Query: 14 ITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCP 73
+TE QT+L I + + YA ++EK E VAE N P SMM +C
Sbjct: 861 VTEFQTNLVPYPRIHFMLSS--YAPVISAEKAYHEQLS---VAEITNSAFEPSSMMAKCD 1025
Query: 74 MKQNFWMMRKRLSTRDC--KNIIKEVVTIKT 102
+ +M + D K++ V TIKT
Sbjct: 1026PRHGKYMACCLMYRGDVVPKDVNAAVATIKT 1118
>TC79934 similar to GP|20259633|gb|AAM14173.1 putative GAR1 protein
{Arabidopsis thaliana}, partial (87%)
Length = 814
Score = 29.3 bits (64), Expect = 0.46
Identities = 13/38 (34%), Positives = 20/38 (52%)
Frame = +1
Query: 2 EMIPLIEGSILWITERQTSLGLIDEIFGQVKNPYYAVR 39
E IP I + T +G +DEIFG + Y++V+
Sbjct: 286 EKIPFFNAPIY--LQNMTQIGKVDEIFGPINESYFSVK 393
>TC76677 homologue to SP|P33629|TBA_PRUDU Tubulin alpha chain. [Almond
Prunus amygdalus] {Prunus dulcis}, partial (80%)
Length = 1392
Score = 29.3 bits (64), Expect = 0.46
Identities = 28/91 (30%), Positives = 40/91 (43%), Gaps = 2/91 (2%)
Frame = +1
Query: 14 ITERQTSLGLIDEIFGQVKNPYYAVRYNSEKGIREGTLISFVAEFVNYVMHPVSMMKRCP 73
+TE QT+L I + + YA ++EK E VAE N P SMM +C
Sbjct: 583 VTEFQTNLVPYPRIHFMLSS--YAPVISAEKAYHEQLS---VAEITNSAFEPSSMMAKCD 747
Query: 74 MKQNFWMMRKRLSTRDC--KNIIKEVVTIKT 102
+ +M + D K++ V TIKT
Sbjct: 748 PRHGKYMACCLMYRGDVVPKDVNAAVATIKT 840
>BG646473 similar to GP|15293255|gb| unknown protein {Arabidopsis thaliana},
partial (3%)
Length = 626
Score = 27.3 bits (59), Expect = 1.7
Identities = 16/83 (19%), Positives = 33/83 (39%), Gaps = 2/83 (2%)
Frame = +1
Query: 36 YAVRYNSEKGIR--EGTLISFVAEFVNYVMHPVSMMKRCPMKQNFWMMRKRLSTRDCKNI 93
+ +NS R E L+ + + ++HP M+ + M DCK +
Sbjct: 265 FMANHNSSGSKRPSEENLLQCLRYLYDSLLHPRMMLNYVDEETRTLAMSSAQKLLDCKQL 444
Query: 94 IKEVVTIKTPIERMENNIKQIPL 116
+K+ + P+ + N + + L
Sbjct: 445 VKDYCSEMLPVSDLYNYVSHVKL 513
>TC91584 similar to GP|21592519|gb|AAM64469.1 unknown {Arabidopsis
thaliana}, partial (71%)
Length = 590
Score = 26.2 bits (56), Expect = 3.9
Identities = 11/48 (22%), Positives = 21/48 (42%), Gaps = 6/48 (12%)
Frame = +2
Query: 63 MHPVSMMKRCPMKQN------FWMMRKRLSTRDCKNIIKEVVTIKTPI 104
M PV + K+ P+ FW+ + + CK ++ I+ P+
Sbjct: 425 MLPVDLQKQWPLSNKNLKDVRFWLSAMGIPCKSCKQFFMQLKNIRNPL 568
>TC88288 weakly similar to GP|10172602|dbj|BAB01406. gene_id:MJG19.10~similar
to probable membrane-associated salt-inducible protein,
partial (5%)
Length = 1668
Score = 26.2 bits (56), Expect = 3.9
Identities = 18/66 (27%), Positives = 36/66 (54%), Gaps = 6/66 (9%)
Frame = -2
Query: 51 LISFVAEFVNYVMHPVSMMKR-----CPMKQN-FWMMRKRLSTRDCKNIIKEVVTIKTPI 104
L+S VA V HP+S ++ C + + FW + +S +IIK++ ++ TPI
Sbjct: 1109 LVSIVA----VV*HPMSAPRQLVPLSCSLNNH*FWRGKSTMSPVILASIIKKLPSLTTPI 942
Query: 105 ERMENN 110
+++ ++
Sbjct: 941 QKVHHH 924
>TC77786
Length = 1094
Score = 25.8 bits (55), Expect = 5.0
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = -2
Query: 64 HPVSMMKRCPMKQNFWMMRKRLS 86
HP S K+ + N+WM+ KRL+
Sbjct: 424 HP*SCKKKKNKRYNYWMLLKRLN 356
>TC77787
Length = 436
Score = 25.0 bits (53), Expect = 8.6
Identities = 10/23 (43%), Positives = 15/23 (64%)
Frame = -1
Query: 64 HPVSMMKRCPMKQNFWMMRKRLS 86
HP S K+ + N+WM+ KRL+
Sbjct: 364 HP*SCKKKKNKRCNYWMLLKRLN 296
Database: MTGI
Posted date: Oct 22, 2004 3:39 PM
Number of letters in database: 27,044,181
Number of sequences in database: 36,976
Lambda K H
0.322 0.138 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,873,942
Number of Sequences: 36976
Number of extensions: 43029
Number of successful extensions: 193
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 192
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 193
length of query: 130
length of database: 9,014,727
effective HSP length: 85
effective length of query: 45
effective length of database: 5,871,767
effective search space: 264229515
effective search space used: 264229515
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC137602.4


